Difference between revisions of "Part:BBa K1159000"
IngmarPolte (Talk | contribs) |
IngmarPolte (Talk | contribs) |
||
| Line 8: | Line 8: | ||
===Protein Data Table for the erythromycin esterase (EreB) <partinfo>BBa_K1159000</partinfo>=== | ===Protein Data Table for the erythromycin esterase (EreB) <partinfo>BBa_K1159000</partinfo>=== | ||
<html><style type="text/css">table#AutoAnnotator {border:1px solid black; width:100%; border-collapse:collapse;} th#AutoAnnotatorHeader { border:1px solid black; width:100%; background-color: rgb(221, 221, 221);} td.AutoAnnotator1col { width:100%; border:1px solid black; } span.AutoAnnotatorSequence { font-family:'Courier New', Arial; } td.AutoAnnotatorSeqNum { text-align:right; width:2%; } td.AutoAnnotatorSeqSeq { width:98% } td.AutoAnnotatorSeqFeat1 { width:3% } td.AutoAnnotatorSeqFeat2a { width:27% } td.AutoAnnotatorSeqFeat2b { width:97% } td.AutoAnnotatorSeqFeat3 { width:70% } table.AutoAnnotatorNoBorder { border:0px; width:100%; border-collapse:collapse; } table.AutoAnnotatorWithBorder { border:1px solid black; width:100%; border-collapse:collapse; } td.AutoAnnotatorOuterAmino { border:0px solid black; width:20% } td.AutoAnnotatorInnerAmino { border:1px solid black; width:50% } td.AutoAnnotatorAminoCountingOuter { border:1px solid black; width:40%; } td.AutoAnnotatorBiochemParOuter { border:1px solid black; width:60%; } td.AutoAnnotatorAminoCountingInner1 { width: 7.5% } td.AutoAnnotatorAminoCountingInner2 { width:62.5% } td.AutoAnnotatorAminoCountingInner3 { width:30% } td.AutoAnnotatorBiochemParInner1 { width: 5% } td.AutoAnnotatorBiochemParInner2 { width:55% } td.AutoAnnotatorBiochemParInner3 { width:40% } td.AutoAnnotatorCodonUsage1 { width: 3% } td.AutoAnnotatorCodonUsage2 { width:14.2% } td.AutoAnnotatorCodonUsage3 { width:13.8% } td.AutoAnnotatorAlignment1 { width: 3% } td.AutoAnnotatorAlignment2 { width: 10% } td.AutoAnnotatorAlignment3 { width: 87% } td.AutoAnnotatorLocalizationOuter {border:1px solid black; width:40%} td.AutoAnnotatorGOOuter {border:1px solid black; width:60%} td.AutoAnnotatorLocalization1 { width: 7.5% } td.AutoAnnotatorLocalization2 { width: 22.5% } td.AutoAnnotatorLocalization3 { width: 70% } td.AutoAnnotatorGO1 { width: 5% } td.AutoAnnotatorGO2 { width: 35% } td.AutoAnnotatorGO3 { width: 60% } td.AutoAnnotatorPredFeat1 { width:3% } td.AutoAnnotatorPredFeat2a { width:27% } td.AutoAnnotatorPredFeat3 { width:70% } div.AutoAnnotator_trans { position:absolute; background:rgb(11,140,143); background-color:rgba(11,140,143, 0.8); height:5px; top:100px; } div.AutoAnnotator_sec_helix { position:absolute; background:rgb(102,0,102); background-color:rgba(102,0,102, 0.8); height:5px; top:110px; } div.AutoAnnotator_sec_strand { position:absolute; background:rgb(245,170,26); background-color:rgba(245,170,26, 1); height:5px; top:110px; } div.AutoAnnotator_acc_buried { position:absolute; background:rgb(89,168,15); background-color:rgba(89,168,15, 0.8); height:5px; top:120px; } div.AutoAnnotator_acc_exposed { position:absolute; background:rgb(0, 0, 255); background-color:rgba(0, 0, 255, 0.8); height:5px; top:120px; } div.AutoAnnotator_dis { position:absolute; text-align:center; font-family:Arial,Helvetica,sans-serif; background:rgb(255, 200, 0); background-color:rgba(255, 200, 0, 1); height:16px; width:16px; top:80px; border-radius:50%; } </style><div id='AutoAnnotator_container_1380912466041'><table id="AutoAnnotator"><tr><!-- Time stamp in ms since 1/1/1970 1380912466041 --><th id="AutoAnnotatorHeader" colspan="2">Protein data table for BioBrick <a href="https://parts.igem.org/wiki/index.php?title=Part:BBa_BBa_K1159000">BBa_BBa_K1159000</a> automatically created by the <a href="http://2013.igem.org/Team:TU-Munich/Results/AutoAnnotator">BioBrick-AutoAnnotator</a> version 1.0</th></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Nucleotide sequence</strong> in <strong>RFC 25</strong>, so ATGGCCGGC and ACCGGT were added (in italics) to the 5' and 3' ends: (underlined part encodes the protein)<br><span class="AutoAnnotatorSequence"> <u><i>ATGGCCGGC</i>AGGTTCGAA ... GTTTATGAA<i>ACCGGT</i></u></span><br> <strong>ORF</strong> from nucleotide position -8 to 1260 (excluding stop-codon)</td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Amino acid sequence:</strong> (RFC 25 scars in shown in bold, other sequence features underlined; both given below)<br><span class="AutoAnnotatorSequence"><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorSeqNum">1 <br>101 <br>201 <br>301 <br>401 </td><td class="AutoAnnotatorSeqSeq">MAGRFEEWVKDKHIPFKLNHPDDNYDDFKPLRKIIGDTRVVALGENSHFIKEFFLLRHTLLRFFIEDLGFTTFAFEFGFAEGQIINNWIHGQGTDDEIGR<br>FLKHFYYPEELKTTFLWLREYNKAAKEKITFLGIDIPRNGGSYLPNMEIVHDFFRTADKEALHIIDDAFNIAKKIDYFSTSQAALNLHELTDSEKCRLTS<br>QLARVKVRLEAMAPIHIEKYGIDKYETILHYANGMIYLDYNIQAMSGFISGGGMQGDMGAKDKYMADSVLWHLKNPQSEQKVIVVAHNAHIQKTPILYDG<br>FLSCLPMGQRLKNAIGDDYMSLGITSYSGHTAALYPEVDTKYGFRVDNFQLQEPNEGSVEKAISGCGVTNSFVFFRNIPEDLQSIPNMIRFDSIYMKAEL<br>EKAFDGIFQIEKSSVSEVVYETG*</td></tr></table></span></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Sequence features:</strong> (with their position in the amino acid sequence, see the <a href="http://2013.igem.org/Team:TU-Munich/Results/Software/FeatureList">list of supported features</a>)<table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorSeqFeat1"></td><td class="AutoAnnotatorSeqFeat2b">None of the supported features appeared in the sequence</td></tr></table></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Amino acid composition:</strong><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorOuterAmino"><table class="AutoAnnotatorWithBorder"><tr><td class="AutoAnnotatorInnerAmino">Ala (A)</td><td class="AutoAnnotatorInnerAmino">27 (6.4%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Arg (R)</td><td class="AutoAnnotatorInnerAmino">16 (3.8%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Asn (N)</td><td class="AutoAnnotatorInnerAmino">20 (4.7%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Asp (D)</td><td class="AutoAnnotatorInnerAmino">29 (6.9%)</td></tr></table></td><td class="AutoAnnotatorOuterAmino"><table class="AutoAnnotatorWithBorder"><tr><td class="AutoAnnotatorInnerAmino">Cys (C)</td><td class="AutoAnnotatorInnerAmino">3 (0.7%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Gln (Q)</td><td class="AutoAnnotatorInnerAmino">14 (3.3%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Glu (E)</td><td class="AutoAnnotatorInnerAmino">30 (7.1%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Gly (G)</td><td class="AutoAnnotatorInnerAmino">31 (7.3%)</td></tr></table></td><td class="AutoAnnotatorOuterAmino"><table class="AutoAnnotatorWithBorder"><tr><td class="AutoAnnotatorInnerAmino">His (H)</td><td class="AutoAnnotatorInnerAmino">15 (3.5%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Ile (I)</td><td class="AutoAnnotatorInnerAmino">36 (8.5%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Leu (L)</td><td class="AutoAnnotatorInnerAmino">34 (8.0%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Lys (K)</td><td class="AutoAnnotatorInnerAmino">29 (6.9%)</td></tr></table></td><td class="AutoAnnotatorOuterAmino"><table class="AutoAnnotatorWithBorder"><tr><td class="AutoAnnotatorInnerAmino">Met (M)</td><td class="AutoAnnotatorInnerAmino">12 (2.8%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Phe (F)</td><td class="AutoAnnotatorInnerAmino">31 (7.3%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Pro (P)</td><td class="AutoAnnotatorInnerAmino">14 (3.3%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Ser (S)</td><td class="AutoAnnotatorInnerAmino">22 (5.2%)</td></tr></table></td><td class="AutoAnnotatorOuterAmino"><table class="AutoAnnotatorWithBorder"><tr><td class="AutoAnnotatorInnerAmino">Thr (T)</td><td class="AutoAnnotatorInnerAmino">19 (4.5%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Trp (W)</td><td class="AutoAnnotatorInnerAmino">4 (0.9%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Tyr (Y)</td><td class="AutoAnnotatorInnerAmino">19 (4.5%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Val (V)</td><td class="AutoAnnotatorInnerAmino">18 (4.3%)</td></tr></table></td></tr></table></td></tr><tr><td class="AutoAnnotatorAminoCountingOuter"><strong>Amino acid counting</strong><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorAminoCountingInner1"></td><td class="AutoAnnotatorAminoCountingInner2">Total number:</td><td class="AutoAnnotatorAminoCountingInner3">423</td></tr><tr><td class="AutoAnnotatorAminoCountingInner1"></td><td class="AutoAnnotatorAminoCountingInner2">Positively charged (Arg+Lys):</td><td class="AutoAnnotatorAminoCountingInner3">45 (10.6%)</td></tr><tr><td class="AutoAnnotatorAminoCountingInner1"></td><td class="AutoAnnotatorAminoCountingInner2">Negatively charged (Asp+Glu):</td><td class="AutoAnnotatorAminoCountingInner3">59 (13.9%)</td></tr><tr><td class="AutoAnnotatorAminoCountingInner1"></td><td class="AutoAnnotatorAminoCountingInner2">Aromatic (Phe+His+Try+Tyr):</td><td class="AutoAnnotatorAminoCountingInner3">69 (16.3%)</td></tr></table></td><td class="AutoAnnotatorBiochemParOuter"><strong>Biochemical parameters</strong><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorBiochemParInner1"></td><td class="AutoAnnotatorBiochemParInner2">Atomic composition:</td><td class="AutoAnnotatorBiochemParInner3">C<sub>2204</sub>H<sub>3348</sub>N<sub>568</sub>O<sub>636</sub>S<sub>15</sub></td></tr><tr><td class="AutoAnnotatorBiochemParInner1"></td><td class="AutoAnnotatorBiochemParInner2">Molecular mass [Da]:</td><td class="AutoAnnotatorBiochemParInner3">48459.2</td></tr><tr><td class="AutoAnnotatorBiochemParInner1"></td><td class="AutoAnnotatorBiochemParInner2">Theoretical pI:</td><td class="AutoAnnotatorBiochemParInner3">5.55</td></tr><tr><td class="AutoAnnotatorBiochemParInner1"></td><td class="AutoAnnotatorBiochemParInner2">Extinction coefficient at 280 nm [M<sup>-1</sup> cm<sup>-1</sup>]:</td><td class="AutoAnnotatorBiochemParInner3">50310 / 50498 (all Cys red/ox)</td></tr></table></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Plot for hydrophobicity, charge, predicted secondary structure, solvent accessability, transmembrane helices and disulfid bridges</strong> <input type='button' id='hydrophobicity_charge_button' onclick='show_or_hide_plot_1380912466041()' value='Show'><span id="hydrophobicity_charge_explanation"></span><div id="hydrophobicity_charge_container" style='display:none'><div id="hydrophobicity_charge_placeholder0" style="width:100%;height:150px"></div><div id="hydrophobicity_charge_placeholder1" style="width:100%;height:150px"></div><div id="hydrophobicity_charge_placeholder2" style="width:100%;height:150px"></div></div></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Codon usage</strong><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorCodonUsage1"></td><td class="AutoAnnotatorCodonUsage2">Organism:</td><td class="AutoAnnotatorCodonUsage3"><i>E. coli</i></td><td class="AutoAnnotatorCodonUsage3"><i>B. subtilis</i></td><td class="AutoAnnotatorCodonUsage3"><i>S. cerevisiae</i></td><td class="AutoAnnotatorCodonUsage3"><i>A. thaliana</i></td><td class="AutoAnnotatorCodonUsage3"><i>P. patens</i></td><td class="AutoAnnotatorCodonUsage3">Mammals</td></tr><tr><td class="AutoAnnotatorCodonUsage1"></td><td class="AutoAnnotatorCodonUsage2">Codon quality (<a href="http://en.wikipedia.org/wiki/Codon_Adaptation_Index">CAI</a>):</td><td class="AutoAnnotatorCodonUsage3">good (0.74)</td><td class="AutoAnnotatorCodonUsage3">good (0.77)</td><td class="AutoAnnotatorCodonUsage3">good (0.75)</td><td class="AutoAnnotatorCodonUsage3">good (0.80)</td><td class="AutoAnnotatorCodonUsage3">good (0.76)</td><td class="AutoAnnotatorCodonUsage3">good (0.66)</td></tr></table></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Alignments</strong> (obtained from <a href='http://predictprotein.org'>PredictProtein.org</a>)<table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorAlignment1"></td><td class="AutoAnnotatorAlignment2">SwissProt:</td><td class="AutoAnnotatorAlignment3"><a href='http://www.uniprot.org/uniprot/P05789'>P05789</a> (100% identity on 418 AAs), <a href='http://www.uniprot.org/uniprot/P07684'>P07684</a> (24% identity on 381 AAs)</td></tr><tr><td class="AutoAnnotatorAlignment1"></td><td class="AutoAnnotatorAlignment2">TrEML:</td><td class="AutoAnnotatorAlignment3"><a href='http://www.uniprot.org/uniprot/Q2V0Y9'>Q2V0Y9</a> (100% identity on 418 AAs), <a href='http://www.uniprot.org/uniprot/D7J5T5'>D7J5T5</a> (44% identity on 402 AAs)</td></tr><tr><td class="AutoAnnotatorAlignment1"></td><td class="AutoAnnotatorAlignment2">PDB:</td><td class="AutoAnnotatorAlignment3"><a href='http://www.rcsb.org/pdb/explore/explore.do?structureId=2qgm'>2qgm</a> (20% identity on 373 AAs), <a href='http://www.rcsb.org/pdb/explore/explore.do?structureId=2rad'>2rad</a> (19% identity on 386 AAs)</td></tr></table></td></tr><tr><th id='AutoAnnotatorHeader' colspan="2"><strong>Predictions</strong> (obtained from <a href='http://predictprotein.org'>PredictProtein.org</a>)</th></tr><tr><td class="AutoAnnotatorLocalizationOuter"><strong>Subcellular Localization</strong> (reliability in brackets)<table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorLocalization1"></td><td class="AutoAnnotatorLocalization2">Archaea:</td><td class="AutoAnnotatorLocalization3">cytosol (100%)</td></tr><tr><td class="AutoAnnotatorLocalization1"></td><td class="AutoAnnotatorLocalization2">Bacteria:</td><td class="AutoAnnotatorLocalization3">cytosol (100%)</td></tr><tr><td class="AutoAnnotatorLocalization1"></td><td class="AutoAnnotatorLocalization2">Eukarya:</td><td class="AutoAnnotatorLocalization3">cytosol (58%)</td></tr></table></td><td class="AutoAnnotatorGOOuter"><strong>Gene Ontology</strong> (reliability in brackets)<br><table class="AutoAnnotatorNoBorder"><tr><td class='AutoAnnotatorGO1'></td><td class='AutoAnnotatorGO2'>Molecular Function Ontology:</td><td class='AutoAnnotatorGO3'> - </td></tr><tr><td class='AutoAnnotatorGO1'></td><td class='AutoAnnotatorGO2'>Biological Process Ontology:</td><td class='AutoAnnotatorGO3'><a href='http://amigo.geneontology.org/cgi-bin/amigo/term_details?term=GO:0046677'>GO:0046677</a> (51%)</td></tr><tr><td class='AutoAnnotatorGO1'> </td><td class='AutoAnnotatorGO2'> </td><td class='AutoAnnotatorGO3'> </td></tr></table></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Predicted features:</strong><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorPredFeat1"></td><td class="AutoAnnotatorPredFeat2a">Disulfid bridges:</td><td class="AutoAnnotatorPredFeat3"> - </td></tr><tr><td class="AutoAnnotatorPredFeat1"></td><td class="AutoAnnotatorPredFeat2a">Transmembrane helices:</td><td class="AutoAnnotatorPredFeat3"> - </td></tr></table></td></tr><tr><td class="AutoAnnotator1col" colspan="2"> The BioBrick-AutoAnnotator was created by <a href="http://2013.igem.org/Team:TU-Munich">TU-Munich 2013</a> iGEM team. For more information please see the <a href="http://2013.igem.org/Team:TU-Munich/Results/Software">documentation</a>.<br>If you have any questions, comments or suggestions, please leave us a <a href="http://2013.igem.org/Team:TU-Munich/Results/AutoAnnotator">comment</a>.</td></tr></table></div><br><!-- IMPORTANT: DON'T REMOVE THIS LINE, OTHERWISE NOT SUPPORTED FOR IE BEFORE 9 --><!--[if lte IE 8]><script language="javascript" type="text/javascript" src="excanvas.min.js"></script><![endif]--><script type='text/javascript' src='http://code.jquery.com/jquery-1.10.0.min.js'></script><script type='text/javascript' src='http://2013.igem.org/Team:TU-Munich/Flot.js?action=raw&ctype=text/js'></script><script>function show_or_hide_plot_1380912466041(){hydrophobicity_datapoints = [[2.5,0.32],[3.5,-0.76],[4.5,-1.82],[5.5,-1.92],[6.5,-0.18],[7.5,-1.52],[8.5,-1.52],[9.5,-1.60],[10.5,-2.06],[11.5,-2.00],[12.5,-1.54],[13.5,-0.28],[14.5,-0.28],[15.5,1.12],[16.5,-0.48],[17.5,-0.80],[18.5,-1.68],[19.5,-1.60],[20.5,-3.06],[21.5,-3.06],[22.5,-2.68],[23.5,-3.06],[24.5,-3.06],[25.5,-1.80],[26.5,-1.88],[27.5,-1.94],[28.5,-0.48],[29.5,-0.68],[30.5,-2.02],[31.5,-0.34],[32.5,0.88],[33.5,0.04],[34.5,0.24],[35.5,0.88],[36.5,-0.92],[37.5,-0.98],[38.5,-0.06],[39.5,1.00],[40.5,1.90],[41.5,2.72],[42.5,1.18],[43.5,-0.36],[44.5,-0.88],[45.5,-2.28],[46.5,-1.64],[47.5,-0.04],[48.5,-0.12],[49.5,-0.66],[50.5,0.54],[51.5,0.54],[52.5,0.40],[53.5,1.94],[54.5,1.74],[55.5,0.54],[56.5,-0.16],[57.5,-0.16],[58.5,-0.16],[59.5,-0.16],[60.5,1.04],[61.5,1.74],[62.5,1.88],[63.5,0.42],[64.5,0.62],[65.5,0.82],[66.5,0.18],[67.5,-0.16],[68.5,0.40],[69.5,0.96],[70.5,0.76],[71.5,1.20],[72.5,1.20],[73.5,0.64],[74.5,1.34],[75.5,0.70],[76.5,0.90],[77.5,0.70],[78.5,0.70],[79.5,0.06],[80.5,-0.56],[81.5,-0.22],[82.5,0.32],[83.5,0.32],[84.5,-0.30],[85.5,0.22],[86.5,0.22],[87.5,-1.32],[88.5,-0.70],[89.5,-0.70],[90.5,-0.60],[91.5,-1.64],[92.5,-1.70],[93.5,-2.32],[94.5,-2.32],[95.5,-1.34],[96.5,-1.28],[97.5,-1.48],[98.5,-0.22],[99.5,1.24],[100.5,-0.44],[101.5,-1.00],[102.5,0.46],[103.5,-0.36],[104.5,-1.38],[105.5,-0.92],[106.5,-0.98],[107.5,-2.24],[108.5,-1.22],[109.5,-1.74],[110.5,-1.56],[111.5,-1.00],[112.5,0.26],[113.5,0.26],[114.5,0.86],[115.5,1.76],[116.5,1.00],[117.5,-0.26],[118.5,-1.28],[119.5,-1.80],[120.5,-3.34],[121.5,-2.08],[122.5,-1.02],[123.5,-1.54],[124.5,-1.54],[125.5,-1.54],[126.5,-1.00],[127.5,-1.50],[128.5,-0.16],[129.5,1.30],[130.5,2.00],[131.5,2.00],[132.5,1.44],[133.5,1.78],[134.5,0.70],[135.5,-0.12],[136.5,-1.72],[137.5,-1.10],[138.5,-2.08],[139.5,-1.92],[140.5,-1.28],[141.5,0.18],[142.5,-0.06],[143.5,-0.68],[144.5,-0.14],[145.5,-0.58],[146.5,-0.44],[147.5,0.72],[148.5,0.78],[149.5,-0.30],[150.5,0.96],[151.5,0.62],[152.5,-1.12],[153.5,-0.62],[154.5,0.44],[155.5,-0.82],[156.5,-2.16],[157.5,-1.96],[158.5,-1.46],[159.5,-1.06],[160.5,-1.00],[161.5,0.68],[162.5,2.28],[163.5,1.22],[164.5,-0.24],[165.5,0.76],[166.5,0.42],[167.5,-1.18],[168.5,0.42],[169.5,1.48],[170.5,0.34],[171.5,-1.00],[172.5,0.60],[173.5,-1.00],[174.5,-1.62],[175.5,-0.28],[176.5,0.34],[177.5,-0.70],[178.5,-0.16],[179.5,-0.60],[180.5,-0.80],[181.5,-0.28],[182.5,0.62],[183.5,0.08],[184.5,1.54],[185.5,0.54],[186.5,-0.52],[187.5,-0.52],[188.5,0.04],[189.5,-1.42],[190.5,-0.94],[191.5,-0.94],[192.5,-2.48],[193.5,-1.84],[194.5,-2.04],[195.5,-1.12],[196.5,-0.56],[197.5,0.06],[198.5,-1.14],[199.5,0.52],[200.5,0.12],[201.5,-0.64],[202.5,0.36],[203.5,0.28],[204.5,0.36],[205.5,-0.90],[206.5,0.76],[207.5,-0.78],[208.5,0.36],[209.5,-0.10],[210.5,1.16],[211.5,0.08],[212.5,1.68],[213.5,0.68],[214.5,1.20],[215.5,0.14],[216.5,-0.32],[217.5,-1.48],[218.5,-0.92],[219.5,-0.92],[220.5,-0.92],[221.5,-0.92],[222.5,-0.92],[223.5,-1.54],[224.5,-2.58],[225.5,-0.98],[226.5,0.56],[227.5,0.18],[228.5,0.62],[229.5,1.12],[230.5,-0.48],[231.5,-1.32],[232.5,-0.30],[233.5,0.86],[234.5,0.24],[235.5,1.70],[236.5,1.08],[237.5,0.44],[238.5,-1.16],[239.5,-0.00],[240.5,-1.46],[241.5,-0.40],[242.5,0.24],[243.5,0.78],[244.5,-0.20],[245.5,1.06],[246.5,1.60],[247.5,1.06],[248.5,1.14],[249.5,1.14],[250.5,0.50],[251.5,-0.02],[252.5,-0.56],[253.5,-0.56],[254.5,-1.18],[255.5,-0.72],[256.5,-1.18],[257.5,-0.12],[258.5,-0.82],[259.5,-0.82],[260.5,-1.98],[261.5,-2.16],[262.5,-2.14],[263.5,-1.00],[264.5,-1.00],[265.5,-0.38],[266.5,0.72],[267.5,1.10],[268.5,0.56],[269.5,0.62],[270.5,1.54],[271.5,-0.08],[272.5,-1.54],[273.5,-1.68],[274.5,-1.74],[275.5,-2.66],[276.5,-2.58],[277.5,-2.58],[278.5,-3.04],[279.5,-1.50],[280.5,-0.44],[281.5,1.10],[282.5,2.64],[283.5,3.78],[284.5,2.30],[285.5,0.70],[286.5,0.22],[287.5,-1.26],[288.5,-0.72],[289.5,-0.78],[290.5,-0.86],[291.5,-1.36],[292.5,-1.04],[293.5,-1.04],[294.5,0.42],[295.5,0.94],[296.5,0.38],[297.5,0.62],[298.5,0.28],[299.5,0.28],[300.5,0.38],[301.5,1.58],[302.5,2.42],[303.5,1.54],[304.5,1.16],[305.5,1.24],[306.5,0.04],[307.5,-1.62],[308.5,-0.54],[309.5,-1.70],[310.5,-2.32],[311.5,-1.26],[312.5,0.54],[313.5,-0.30],[314.5,-0.22],[315.5,-0.22],[316.5,-0.84],[317.5,-1.36],[318.5,-1.44],[319.5,0.02],[320.5,0.64],[321.5,1.80],[322.5,1.28],[323.5,1.28],[324.5,0.26],[325.5,0.18],[326.5,-0.80],[327.5,-1.30],[328.5,-1.28],[329.5,-0.66],[330.5,-0.14],[331.5,0.70],[332.5,1.08],[333.5,0.90],[334.5,-0.16],[335.5,0.32],[336.5,-1.14],[337.5,-1.02],[338.5,-1.48],[339.5,-1.04],[340.5,-1.96],[341.5,-0.70],[342.5,-1.46],[343.5,0.16],[344.5,-0.28],[345.5,-0.90],[346.5,-0.90],[347.5,-0.70],[348.5,-0.78],[349.5,-0.78],[350.5,-0.78],[351.5,-1.66],[352.5,-1.66],[353.5,-3.12],[354.5,-2.50],[355.5,-1.96],[356.5,-0.80],[357.5,-0.80],[358.5,-0.88],[359.5,-0.44],[360.5,0.62],[361.5,-0.38],[362.5,0.24],[363.5,1.52],[364.5,1.08],[365.5,1.02],[366.5,1.04],[367.5,0.42],[368.5,-0.24],[369.5,0.40],[370.5,0.40],[371.5,1.10],[372.5,2.36],[373.5,1.62],[374.5,0.36],[375.5,0.42],[376.5,-0.46],[377.5,-1.72],[378.5,-1.52],[379.5,-0.06],[380.5,-1.66],[381.5,-1.50],[382.5,0.10],[383.5,0.48],[384.5,-0.98],[385.5,0.10],[386.5,1.16],[387.5,-0.64],[388.5,0.24],[389.5,0.24],[390.5,-0.30],[391.5,-0.30],[392.5,0.34],[393.5,0.16],[394.5,0.08],[395.5,0.60],[396.5,-1.00],[397.5,0.02],[398.5,-1.06],[399.5,-1.06],[400.5,-1.06],[401.5,0.20],[402.5,-1.26],[403.5,-0.64],[404.5,1.04],[405.5,1.24],[406.5,-0.02],[407.5,1.58],[408.5,0.96],[409.5,-0.72],[410.5,-1.44],[411.5,-0.90],[412.5,-0.96],[413.5,-0.42],[414.5,-0.34],[415.5,0.66],[416.5,1.66],[417.5,0.56],[418.5,0.02],[419.5,0.58],[420.5,-0.34]];charge_datapoints = [[2.5,0.20],[3.5,0.00],[4.5,-0.20],[5.5,-0.20],[6.5,-0.40],[7.5,-0.20],[8.5,-0.20],[9.5,0.20],[10.5,0.30],[11.5,0.30],[12.5,0.10],[13.5,0.30],[14.5,0.30],[15.5,0.20],[16.5,0.20],[17.5,0.30],[18.5,0.30],[19.5,-0.10],[20.5,-0.30],[21.5,-0.30],[22.5,-0.40],[23.5,-0.60],[24.5,-0.60],[25.5,-0.40],[26.5,-0.20],[27.5,-0.20],[28.5,-0.00],[29.5,0.40],[30.5,0.60],[31.5,0.40],[32.5,0.40],[33.5,0.40],[34.5,-0.00],[35.5,-0.20],[36.5,-0.00],[37.5,-0.00],[38.5,-0.00],[39.5,0.20],[40.5,0.20],[41.5,-0.00],[42.5,-0.20],[43.5,-0.20],[44.5,-0.20],[45.5,-0.10],[46.5,-0.10],[47.5,0.10],[48.5,0.30],[49.5,0.10],[50.5,-0.00],[51.5,-0.00],[52.5,-0.00],[53.5,-0.20],[54.5,0.20],[55.5,0.30],[56.5,0.30],[57.5,0.30],[58.5,0.30],[59.5,0.30],[60.5,0.20],[61.5,0.20],[62.5,0.20],[63.5,-0.00],[64.5,-0.40],[65.5,-0.40],[66.5,-0.40],[67.5,-0.40],[68.5,-0.20],[69.5,-0.00],[70.5,-0.00],[71.5,-0.00],[72.5,-0.00],[73.5,-0.20],[74.5,-0.20],[75.5,-0.20],[76.5,-0.20],[77.5,-0.20],[78.5,-0.20],[79.5,-0.20],[80.5,-0.20],[81.5,-0.20],[82.5,-0.20],[83.5,-0.00],[84.5,-0.00],[85.5,-0.00],[86.5,-0.00],[87.5,0.10],[88.5,0.10],[89.5,0.10],[90.5,0.10],[91.5,0.10],[92.5,-0.20],[93.5,-0.40],[94.5,-0.60],[95.5,-0.60],[96.5,-0.60],[97.5,-0.20],[98.5,-0.00],[99.5,0.20],[100.5,0.40],[101.5,0.50],[102.5,0.30],[103.5,0.30],[104.5,0.30],[105.5,0.10],[106.5,-0.20],[107.5,-0.40],[108.5,-0.40],[109.5,-0.20],[110.5,-0.20],[111.5,0.00],[112.5,0.20],[113.5,0.20],[114.5,0.00],[115.5,0.00],[116.5,0.20],[117.5,0.00],[118.5,0.00],[119.5,0.00],[120.5,0.20],[121.5,0.00],[122.5,0.20],[123.5,0.40],[124.5,0.20],[125.5,0.20],[126.5,0.20],[127.5,0.20],[128.5,0.00],[129.5,0.20],[130.5,0.00],[131.5,0.00],[132.5,-0.20],[133.5,-0.20],[134.5,-0.20],[135.5,0.00],[136.5,0.00],[137.5,0.20],[138.5,0.20],[139.5,0.20],[140.5,0.00],[141.5,0.00],[142.5,0.00],[143.5,0.00],[144.5,0.00],[145.5,-0.20],[146.5,-0.20],[147.5,-0.20],[148.5,-0.10],[149.5,-0.30],[150.5,-0.10],[151.5,-0.10],[152.5,0.10],[153.5,-0.00],[154.5,0.20],[155.5,-0.00],[156.5,0.20],[157.5,-0.20],[158.5,-0.20],[159.5,-0.20],[160.5,0.10],[161.5,-0.10],[162.5,0.10],[163.5,-0.10],[164.5,-0.30],[165.5,-0.40],[166.5,-0.40],[167.5,-0.40],[168.5,-0.20],[169.5,0.00],[170.5,0.20],[171.5,0.40],[172.5,0.40],[173.5,0.20],[174.5,0.20],[175.5,0.00],[176.5,-0.20],[177.5,-0.20],[178.5,0.00],[179.5,0.00],[180.5,0.00],[181.5,0.00],[182.5,0.00],[183.5,0.00],[184.5,0.00],[185.5,0.10],[186.5,-0.10],[187.5,-0.10],[188.5,-0.10],[189.5,-0.30],[190.5,-0.40],[191.5,-0.40],[192.5,-0.20],[193.5,-0.20],[194.5,0.20],[195.5,0.20],[196.5,0.40],[197.5,0.20],[198.5,0.20],[199.5,0.00],[200.5,0.00],[201.5,0.20],[202.5,0.20],[203.5,0.40],[204.5,0.40],[205.5,0.60],[206.5,0.40],[207.5,0.20],[208.5,0.00],[209.5,0.00],[210.5,-0.20],[211.5,-0.20],[212.5,0.00],[213.5,0.10],[214.5,0.10],[215.5,-0.10],[216.5,0.10],[217.5,0.10],[218.5,0.00],[219.5,0.00],[220.5,0.00],[221.5,0.00],[222.5,0.00],[223.5,-0.20],[224.5,-0.20],[225.5,0.00],[226.5,-0.20],[227.5,-0.10],[228.5,0.10],[229.5,0.10],[230.5,0.10],[231.5,0.10],[232.5,0.00],[233.5,0.00],[234.5,0.00],[235.5,0.00],[236.5,-0.20],[237.5,-0.20],[238.5,-0.20],[239.5,-0.20],[240.5,-0.20],[241.5,0.00],[242.5,0.00],[243.5,0.00],[244.5,0.00],[245.5,0.00],[246.5,0.00],[247.5,0.00],[248.5,0.00],[249.5,0.00],[250.5,0.00],[251.5,0.00],[252.5,0.00],[253.5,0.00],[254.5,-0.20],[255.5,-0.20],[256.5,-0.20],[257.5,-0.20],[258.5,0.00],[259.5,0.00],[260.5,0.20],[261.5,0.20],[262.5,0.20],[263.5,0.00],[264.5,0.00],[265.5,-0.20],[266.5,-0.20],[267.5,-0.20],[268.5,-0.20],[269.5,0.10],[270.5,0.10],[271.5,0.30],[272.5,0.30],[273.5,0.30],[274.5,0.20],[275.5,0.20],[276.5,-0.20],[277.5,-0.20],[278.5,0.00],[279.5,0.00],[280.5,0.00],[281.5,0.20],[282.5,0.20],[283.5,0.00],[284.5,0.10],[285.5,0.10],[286.5,0.10],[287.5,0.20],[288.5,0.20],[289.5,0.10],[290.5,0.30],[291.5,0.30],[292.5,0.20],[293.5,0.20],[294.5,0.20],[295.5,0.00],[296.5,-0.20],[297.5,-0.20],[298.5,-0.20],[299.5,-0.20],[300.5,-0.20],[301.5,0.00],[302.5,0.00],[303.5,0.00],[304.5,0.00],[305.5,0.00],[306.5,0.00],[307.5,0.20],[308.5,0.20],[309.5,0.40],[310.5,0.40],[311.5,0.40],[312.5,0.20],[313.5,0.20],[314.5,-0.20],[315.5,-0.40],[316.5,-0.40],[317.5,-0.40],[318.5,-0.40],[319.5,-0.20],[320.5,0.00],[321.5,0.00],[322.5,0.00],[323.5,0.00],[324.5,0.00],[325.5,0.00],[326.5,0.00],[327.5,0.10],[328.5,0.10],[329.5,0.10],[330.5,0.10],[331.5,0.10],[332.5,0.00],[333.5,0.00],[334.5,-0.20],[335.5,-0.20],[336.5,-0.40],[337.5,-0.40],[338.5,-0.20],[339.5,0.00],[340.5,0.00],[341.5,0.20],[342.5,0.40],[343.5,0.20],[344.5,0.00],[345.5,0.00],[346.5,0.00],[347.5,-0.20],[348.5,-0.20],[349.5,0.00],[350.5,-0.20],[351.5,-0.20],[352.5,-0.20],[353.5,-0.40],[354.5,-0.40],[355.5,-0.20],[356.5,-0.20],[357.5,-0.40],[358.5,0.00],[359.5,0.00],[360.5,0.00],[361.5,0.00],[362.5,0.20],[363.5,0.00],[364.5,0.00],[365.5,0.00],[366.5,0.00],[367.5,0.00],[368.5,0.00],[369.5,0.00],[370.5,0.00],[371.5,0.00],[372.5,0.00],[373.5,0.20],[374.5,0.20],[375.5,0.20],[376.5,0.20],[377.5,0.00],[378.5,-0.40],[379.5,-0.40],[380.5,-0.40],[381.5,-0.40],[382.5,-0.20],[383.5,0.00],[384.5,0.00],[385.5,0.00],[386.5,0.00],[387.5,0.20],[388.5,0.20],[389.5,0.00],[390.5,0.00],[391.5,0.00],[392.5,-0.20],[393.5,-0.20],[394.5,0.20],[395.5,0.20],[396.5,0.00],[397.5,0.00],[398.5,-0.20],[399.5,-0.20],[400.5,-0.20],[401.5,0.00],[402.5,-0.20],[403.5,0.00],[404.5,-0.20],[405.5,-0.20],[406.5,-0.20],[407.5,0.00],[408.5,-0.20],[409.5,0.00],[410.5,0.00],[411.5,0.00],[412.5,0.00],[413.5,0.20],[414.5,-0.20],[415.5,-0.20],[416.5,-0.20],[417.5,-0.20],[418.5,-0.40],[419.5,-0.20],[420.5,-0.20]];dis_datapoints = [];trans_datapoints = [];sec_helix_datapoints = [[3,12],[28,34],[50,67],[81,90],[95,105],[109,121],[144,156],[159,172],[181,188],[193,200],[201,212],[215,218],[224,249],[261,276],[305,314],[359,365],[380,382],[389,391],[401,403]];sec_strand_datapoints = [[39,43],[72,76],[128,134],[282,286],[319,325],[330,334],[371,374],[406,411]];acc_exposed_datapoints = [[1,4],[7,7],[10,11],[15,15],[17,17],[19,19],[21,24],[26,26],[29,29],[33,33],[36,37],[66,67],[90,90],[92,92],[95,96],[99,100],[109,109],[120,120],[123,127],[141,141],[155,156],[158,160],[163,163],[167,167],[170,170],[173,174],[176,177],[179,179],[189,189],[191,193],[197,197],[200,200],[204,204],[207,207],[210,211],[214,214],[218,219],[221,221],[223,224],[251,253],[274,274],[277,279],[313,313],[316,317],[337,341],[343,343],[345,345],[347,347],[350,350],[353,353],[356,356],[364,365],[367,367],[369,369],[380,381],[384,384],[387,387],[397,397],[399,399],[402,402],[412,412],[414,414],[417,417],[420,423]];acc_buried_datapoints = [[5,5],[9,9],[13,13],[16,16],[28,28],[31,31],[35,35],[38,38],[40,50],[53,54],[56,61],[64,65],[69,79],[82,82],[84,86],[88,89],[91,91],[98,98],[101,102],[105,108],[111,115],[117,118],[122,122],[129,137],[142,143],[146,147],[149,151],[153,154],[161,162],[165,165],[168,169],[172,172],[183,187],[198,199],[201,202],[205,205],[209,209],[212,213],[216,217],[225,225],[228,232],[234,238],[240,242],[244,249],[258,258],[260,260],[262,262],[264,266],[268,270],[272,273],[280,280],[282,292],[294,294],[296,296],[298,298],[301,301],[304,311],[315,315],[319,327],[329,334],[346,346],[351,351],[358,359],[362,363],[366,366],[371,375],[378,379],[382,382],[388,388],[391,396],[400,400],[404,410],[415,415]];flot_plot_options = []; flot_plot_options[0] = {grid: {borderWidth: {top: 0,right: 0,bottom: 0,left: 0}},legend: {show: false},xaxes: [{show: true,min: 0,max: 200,ticks: [[0.5, '1'], [24.5, '25'], [49.5, '50'], [74.5, '75'], [99.5, '100'], [124.5, '125'], [149.5, '150'], [174.5, '175'], [199.5, '200']],tickLength: -5}],yaxes: [{show: true,ticks: [[0, '0'], [4.5,'hydro-<br>phobic '], [-4.5,'hydro-<br>philic ']],min: -4.5,max: +4.5,font: {size: 12,lineHeight: 14,style: 'italic',weight: 'bold',family: 'sans-serif',variant: 'small-caps',color: 'rgba(100,149,237,1)'}},{show: true,ticks: [[0, ''], [1,'positive<br> charge'], [-1,'negative<br> charge']],position: 'right',min: -1,max: 1,font: {size: 12,lineHeight: 14,style: 'italic',weight: 'bold',family: 'sans-serif',variant: 'small-caps',color: 'rgba(255,99,71,1)'}}]};number_of_plots = 3;for ( plot_num = 1 ; plot_num < number_of_plots ; plot_num ++){flot_plot_options[plot_num] = $.extend(true, {} ,flot_plot_options[0]);flot_plot_options[plot_num].xaxes = [{min: plot_num*200,max: (plot_num + 1)*200,ticks: [ [plot_num*200 + 0.5, (plot_num*200 + 1).toString()], [plot_num*200 + 24.5, (plot_num*200 + 25).toString()], [plot_num*200 + 49.5, (plot_num*200 + 50).toString()], [plot_num*200 + 74.5, (plot_num*200 + 75).toString()], [plot_num*200 + 99.5, (plot_num*200 + 100).toString()], [plot_num*200 + 124.5, (plot_num*200 + 125).toString()], [plot_num*200 + 149.5, (plot_num*200 + 150).toString()], [plot_num*200 + 174.5, (plot_num*200 + 175).toString()], [plot_num*200 + 199.5, (plot_num*200 + 200).toString()] ],tickLength: -5}];};try {if( $('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_button').val() =='Show' ){$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_container').css('display','block');$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_button').val('Hide');var description_html = '<div id=\'AutoAnnotator_plot_selectors\'>';description_html = description_html + '<br> <input type=\'checkbox\' id=\'hydrophobicity_checkbox\' checked=\'checked\'> Moving average over 5 amino acids for hydrophobicity (<img src=\'https://static.igem.org/mediawiki/2013/e/e9/TUM13_hydrophobicity_icon.png\' alt=\'blue graph\' height=\'10\'></img>)';description_html = description_html + '<br> <input type=\'checkbox\' id=\'charge_checkbox\' checked=\'checked\'> Moving average over 5 amino acids for charge (<img src=\'https://static.igem.org/mediawiki/2013/3/3e/TUM13_charge_icon.png\' alt=\'red graph\' height=\'10\'></img>)';description_html = description_html + '<br> <input type=\'checkbox\' id=\'dis_checkbox\' checked=\'checked\'> Predicted disulfid bridges (<img src=\'https://static.igem.org/mediawiki/2013/2/28/TUM13_dis_icon.png\' alt=\'yellow circle\' height=\'10\'></img>) with the number of the bridge in the center';description_html = description_html + '<br> <input type=\'checkbox\' id=\'trans_checkbox\' checked=\'checked\'> Predicted transmembrane helices (<img src=\'https://static.igem.org/mediawiki/2013/7/78/TUM13_trans_icon.png\' alt=\'turquois bars\' height=\'10\'></img>)';description_html = description_html + '<br> <input type=\'checkbox\' id=\'sec_checkbox\' checked=\'checked\'> Predicted secondary structure: Helices (<img src=\'https://static.igem.org/mediawiki/2013/b/bf/TUM13_helix_icon.png\' alt=\'violet bars\' height=\'10\'></img>) and beta-strands (<img src=\'https://static.igem.org/mediawiki/2013/b/bf/TUM13_strand_icon.png\' alt=\'yellow bars\' height=\'10\'></img>)';description_html = description_html + '<br> <input type=\'checkbox\' id=\'acc_checkbox\' checked=\'checked\'> Predicted solvent accessability: Exposed (<img src=\'https://static.igem.org/mediawiki/2013/1/16/TUM13_exposed_icon.png\' alt=\'blue bars\' height=\'10\'></img>) and buried (<img src=\'https://static.igem.org/mediawiki/2013/0/0b/TUM13_buried_icon.png\' alt=\'green bars\' height=\'10\'></img>) residues';description_html = description_html + '<br></div>';$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_explanation').html(description_html);plot_according_to_selectors_1380912466041();$('#AutoAnnotator_container_1380912466041 #AutoAnnotator_plot_selectors').find('input').click(plot_according_to_selectors_1380912466041);}else{$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_container').css('display','none');$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_button').val('Show');$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_explanation').html('');}}catch(err){txt='There was an error with the button controlling the visibility of the plot.\n';txt=txt+'The originating error is:\n' + err + '\n\n';alert(txt);}};function plot_according_to_selectors_1380912466041(){try{var plot_datasets = [[],[]];if($('#AutoAnnotator_container_1380912466041 #hydrophobicity_checkbox').prop('checked') == true){plot_datasets[0] = { color: 'rgba(100,149,237,1)',data: hydrophobicity_datapoints,label: 'Hydrophobicity',lines: { show: true, fill: true, fillColor: 'rgba(100,149,237,0.1)' },yaxis: 1};}if($('#AutoAnnotator_container_1380912466041 #charge_checkbox').prop('checked') == true){plot_datasets[1] = {color: 'rgba(255,99,71,1)',data: charge_datapoints,label: 'Charge',lines: { show: true, fill: true, fillColor: 'rgba(255,99,71,0.1)' },yaxis: 2};}for (plot_num = 0 ; plot_num < number_of_plots ; plot_num ++){$.plot('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder'+ plot_num.toString(), plot_datasets, flot_plot_options[plot_num] );}var screen_width = $('canvas.flot-base').width(); var pos_of_first_tick = 46;var pos_of_last_tick = screen_width - 51;var tick_diff = (screen_width - 97)/199;if($('#AutoAnnotator_container_1380912466041 #dis_checkbox').prop('checked') == true){for ( j = 0 ; j < dis_datapoints.length ; j++ ){$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder' + Math.floor((dis_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_dis\' style=\'left:' + ((pos_of_first_tick - 8 + (dis_datapoints[j][0] - 1)*tick_diff - Math.floor((dis_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px;\'><b>' + (j+1) + '</b></div>');$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder' + Math.floor((dis_datapoints[j][1] - 1)/200) ).append('<div class=\'AutoAnnotator_dis\' style=\'left:' + ((pos_of_first_tick - 8 + (dis_datapoints[j][1] - 1)*tick_diff - Math.floor((dis_datapoints[j][1] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px;\'><b>' + (j+1) + '</b></div>');}}if($('#AutoAnnotator_container_1380912466041 #trans_checkbox').prop('checked') == true){for ( j = 0 ; j < trans_datapoints.length ; j++ ){$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder' + Math.floor((trans_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_trans\' style=\'width:' + (((trans_datapoints[j][1] - trans_datapoints[j][0] + 1)*tick_diff).toFixed(0)).toString() + 'px ;left:' + ((pos_of_first_tick + (trans_datapoints[j][0] - 1.5)*tick_diff - Math.floor((trans_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px\'></div>');}}if($('#AutoAnnotator_container_1380912466041 #sec_checkbox').prop('checked') == true){for ( j = 0 ; j < sec_helix_datapoints.length ; j++ ){$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder' + Math.floor((sec_helix_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_sec_helix\' style=\'width:' + (((sec_helix_datapoints[j][1] - sec_helix_datapoints[j][0] + 1)*tick_diff).toFixed(0)).toString() + 'px; left:' + ((pos_of_first_tick + (sec_helix_datapoints[j][0] - 1.5)*tick_diff - Math.floor((sec_helix_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px\'></div>');}for ( j = 0 ; j < sec_strand_datapoints.length ; j++ ){$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder' + Math.floor((sec_strand_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_sec_strand\' style=\'width:' + (((sec_strand_datapoints[j][1] - sec_strand_datapoints[j][0] + 1)*tick_diff).toFixed(0)).toString() + 'px; left:' + ((pos_of_first_tick + (sec_strand_datapoints[j][0] - 1.5)*tick_diff - Math.floor((sec_strand_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px\'></div>');}}if($('#AutoAnnotator_container_1380912466041 #acc_checkbox').prop('checked') == true){for ( j = 0 ; j < acc_buried_datapoints.length ; j++ ){$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder' + Math.floor((acc_buried_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_acc_buried\' style=\'width:' + (((acc_buried_datapoints[j][1] - acc_buried_datapoints[j][0] + 1)*tick_diff).toFixed(0)).toString() + 'px; left:' + ((pos_of_first_tick + (acc_buried_datapoints[j][0] - 1.5)*tick_diff - Math.floor((acc_buried_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px\'></div>');}for ( j = 0 ; j < acc_exposed_datapoints.length ; j++ ){$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder' + Math.floor((acc_exposed_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_acc_exposed\' style=\'width:' + (((acc_exposed_datapoints[j][1] - acc_exposed_datapoints[j][0] + 1)*tick_diff).toFixed(0)).toString() + 'px; left:' + ((pos_of_first_tick + (acc_exposed_datapoints[j][0] - 1.5)*tick_diff - Math.floor((acc_exposed_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px\'></div>');}}}catch(err){txt='There was an error while drawing the selected elements for the plot.\n';txt=txt+'The originating error is:\n' + err + '\n\n';throw(txt);}}</script></html> | <html><style type="text/css">table#AutoAnnotator {border:1px solid black; width:100%; border-collapse:collapse;} th#AutoAnnotatorHeader { border:1px solid black; width:100%; background-color: rgb(221, 221, 221);} td.AutoAnnotator1col { width:100%; border:1px solid black; } span.AutoAnnotatorSequence { font-family:'Courier New', Arial; } td.AutoAnnotatorSeqNum { text-align:right; width:2%; } td.AutoAnnotatorSeqSeq { width:98% } td.AutoAnnotatorSeqFeat1 { width:3% } td.AutoAnnotatorSeqFeat2a { width:27% } td.AutoAnnotatorSeqFeat2b { width:97% } td.AutoAnnotatorSeqFeat3 { width:70% } table.AutoAnnotatorNoBorder { border:0px; width:100%; border-collapse:collapse; } table.AutoAnnotatorWithBorder { border:1px solid black; width:100%; border-collapse:collapse; } td.AutoAnnotatorOuterAmino { border:0px solid black; width:20% } td.AutoAnnotatorInnerAmino { border:1px solid black; width:50% } td.AutoAnnotatorAminoCountingOuter { border:1px solid black; width:40%; } td.AutoAnnotatorBiochemParOuter { border:1px solid black; width:60%; } td.AutoAnnotatorAminoCountingInner1 { width: 7.5% } td.AutoAnnotatorAminoCountingInner2 { width:62.5% } td.AutoAnnotatorAminoCountingInner3 { width:30% } td.AutoAnnotatorBiochemParInner1 { width: 5% } td.AutoAnnotatorBiochemParInner2 { width:55% } td.AutoAnnotatorBiochemParInner3 { width:40% } td.AutoAnnotatorCodonUsage1 { width: 3% } td.AutoAnnotatorCodonUsage2 { width:14.2% } td.AutoAnnotatorCodonUsage3 { width:13.8% } td.AutoAnnotatorAlignment1 { width: 3% } td.AutoAnnotatorAlignment2 { width: 10% } td.AutoAnnotatorAlignment3 { width: 87% } td.AutoAnnotatorLocalizationOuter {border:1px solid black; width:40%} td.AutoAnnotatorGOOuter {border:1px solid black; width:60%} td.AutoAnnotatorLocalization1 { width: 7.5% } td.AutoAnnotatorLocalization2 { width: 22.5% } td.AutoAnnotatorLocalization3 { width: 70% } td.AutoAnnotatorGO1 { width: 5% } td.AutoAnnotatorGO2 { width: 35% } td.AutoAnnotatorGO3 { width: 60% } td.AutoAnnotatorPredFeat1 { width:3% } td.AutoAnnotatorPredFeat2a { width:27% } td.AutoAnnotatorPredFeat3 { width:70% } div.AutoAnnotator_trans { position:absolute; background:rgb(11,140,143); background-color:rgba(11,140,143, 0.8); height:5px; top:100px; } div.AutoAnnotator_sec_helix { position:absolute; background:rgb(102,0,102); background-color:rgba(102,0,102, 0.8); height:5px; top:110px; } div.AutoAnnotator_sec_strand { position:absolute; background:rgb(245,170,26); background-color:rgba(245,170,26, 1); height:5px; top:110px; } div.AutoAnnotator_acc_buried { position:absolute; background:rgb(89,168,15); background-color:rgba(89,168,15, 0.8); height:5px; top:120px; } div.AutoAnnotator_acc_exposed { position:absolute; background:rgb(0, 0, 255); background-color:rgba(0, 0, 255, 0.8); height:5px; top:120px; } div.AutoAnnotator_dis { position:absolute; text-align:center; font-family:Arial,Helvetica,sans-serif; background:rgb(255, 200, 0); background-color:rgba(255, 200, 0, 1); height:16px; width:16px; top:80px; border-radius:50%; } </style><div id='AutoAnnotator_container_1380912466041'><table id="AutoAnnotator"><tr><!-- Time stamp in ms since 1/1/1970 1380912466041 --><th id="AutoAnnotatorHeader" colspan="2">Protein data table for BioBrick <a href="https://parts.igem.org/wiki/index.php?title=Part:BBa_BBa_K1159000">BBa_BBa_K1159000</a> automatically created by the <a href="http://2013.igem.org/Team:TU-Munich/Results/AutoAnnotator">BioBrick-AutoAnnotator</a> version 1.0</th></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Nucleotide sequence</strong> in <strong>RFC 25</strong>, so ATGGCCGGC and ACCGGT were added (in italics) to the 5' and 3' ends: (underlined part encodes the protein)<br><span class="AutoAnnotatorSequence"> <u><i>ATGGCCGGC</i>AGGTTCGAA ... GTTTATGAA<i>ACCGGT</i></u></span><br> <strong>ORF</strong> from nucleotide position -8 to 1260 (excluding stop-codon)</td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Amino acid sequence:</strong> (RFC 25 scars in shown in bold, other sequence features underlined; both given below)<br><span class="AutoAnnotatorSequence"><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorSeqNum">1 <br>101 <br>201 <br>301 <br>401 </td><td class="AutoAnnotatorSeqSeq">MAGRFEEWVKDKHIPFKLNHPDDNYDDFKPLRKIIGDTRVVALGENSHFIKEFFLLRHTLLRFFIEDLGFTTFAFEFGFAEGQIINNWIHGQGTDDEIGR<br>FLKHFYYPEELKTTFLWLREYNKAAKEKITFLGIDIPRNGGSYLPNMEIVHDFFRTADKEALHIIDDAFNIAKKIDYFSTSQAALNLHELTDSEKCRLTS<br>QLARVKVRLEAMAPIHIEKYGIDKYETILHYANGMIYLDYNIQAMSGFISGGGMQGDMGAKDKYMADSVLWHLKNPQSEQKVIVVAHNAHIQKTPILYDG<br>FLSCLPMGQRLKNAIGDDYMSLGITSYSGHTAALYPEVDTKYGFRVDNFQLQEPNEGSVEKAISGCGVTNSFVFFRNIPEDLQSIPNMIRFDSIYMKAEL<br>EKAFDGIFQIEKSSVSEVVYETG*</td></tr></table></span></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Sequence features:</strong> (with their position in the amino acid sequence, see the <a href="http://2013.igem.org/Team:TU-Munich/Results/Software/FeatureList">list of supported features</a>)<table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorSeqFeat1"></td><td class="AutoAnnotatorSeqFeat2b">None of the supported features appeared in the sequence</td></tr></table></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Amino acid composition:</strong><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorOuterAmino"><table class="AutoAnnotatorWithBorder"><tr><td class="AutoAnnotatorInnerAmino">Ala (A)</td><td class="AutoAnnotatorInnerAmino">27 (6.4%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Arg (R)</td><td class="AutoAnnotatorInnerAmino">16 (3.8%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Asn (N)</td><td class="AutoAnnotatorInnerAmino">20 (4.7%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Asp (D)</td><td class="AutoAnnotatorInnerAmino">29 (6.9%)</td></tr></table></td><td class="AutoAnnotatorOuterAmino"><table class="AutoAnnotatorWithBorder"><tr><td class="AutoAnnotatorInnerAmino">Cys (C)</td><td class="AutoAnnotatorInnerAmino">3 (0.7%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Gln (Q)</td><td class="AutoAnnotatorInnerAmino">14 (3.3%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Glu (E)</td><td class="AutoAnnotatorInnerAmino">30 (7.1%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Gly (G)</td><td class="AutoAnnotatorInnerAmino">31 (7.3%)</td></tr></table></td><td class="AutoAnnotatorOuterAmino"><table class="AutoAnnotatorWithBorder"><tr><td class="AutoAnnotatorInnerAmino">His (H)</td><td class="AutoAnnotatorInnerAmino">15 (3.5%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Ile (I)</td><td class="AutoAnnotatorInnerAmino">36 (8.5%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Leu (L)</td><td class="AutoAnnotatorInnerAmino">34 (8.0%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Lys (K)</td><td class="AutoAnnotatorInnerAmino">29 (6.9%)</td></tr></table></td><td class="AutoAnnotatorOuterAmino"><table class="AutoAnnotatorWithBorder"><tr><td class="AutoAnnotatorInnerAmino">Met (M)</td><td class="AutoAnnotatorInnerAmino">12 (2.8%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Phe (F)</td><td class="AutoAnnotatorInnerAmino">31 (7.3%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Pro (P)</td><td class="AutoAnnotatorInnerAmino">14 (3.3%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Ser (S)</td><td class="AutoAnnotatorInnerAmino">22 (5.2%)</td></tr></table></td><td class="AutoAnnotatorOuterAmino"><table class="AutoAnnotatorWithBorder"><tr><td class="AutoAnnotatorInnerAmino">Thr (T)</td><td class="AutoAnnotatorInnerAmino">19 (4.5%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Trp (W)</td><td class="AutoAnnotatorInnerAmino">4 (0.9%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Tyr (Y)</td><td class="AutoAnnotatorInnerAmino">19 (4.5%)</td></tr><tr><td class="AutoAnnotatorInnerAmino">Val (V)</td><td class="AutoAnnotatorInnerAmino">18 (4.3%)</td></tr></table></td></tr></table></td></tr><tr><td class="AutoAnnotatorAminoCountingOuter"><strong>Amino acid counting</strong><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorAminoCountingInner1"></td><td class="AutoAnnotatorAminoCountingInner2">Total number:</td><td class="AutoAnnotatorAminoCountingInner3">423</td></tr><tr><td class="AutoAnnotatorAminoCountingInner1"></td><td class="AutoAnnotatorAminoCountingInner2">Positively charged (Arg+Lys):</td><td class="AutoAnnotatorAminoCountingInner3">45 (10.6%)</td></tr><tr><td class="AutoAnnotatorAminoCountingInner1"></td><td class="AutoAnnotatorAminoCountingInner2">Negatively charged (Asp+Glu):</td><td class="AutoAnnotatorAminoCountingInner3">59 (13.9%)</td></tr><tr><td class="AutoAnnotatorAminoCountingInner1"></td><td class="AutoAnnotatorAminoCountingInner2">Aromatic (Phe+His+Try+Tyr):</td><td class="AutoAnnotatorAminoCountingInner3">69 (16.3%)</td></tr></table></td><td class="AutoAnnotatorBiochemParOuter"><strong>Biochemical parameters</strong><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorBiochemParInner1"></td><td class="AutoAnnotatorBiochemParInner2">Atomic composition:</td><td class="AutoAnnotatorBiochemParInner3">C<sub>2204</sub>H<sub>3348</sub>N<sub>568</sub>O<sub>636</sub>S<sub>15</sub></td></tr><tr><td class="AutoAnnotatorBiochemParInner1"></td><td class="AutoAnnotatorBiochemParInner2">Molecular mass [Da]:</td><td class="AutoAnnotatorBiochemParInner3">48459.2</td></tr><tr><td class="AutoAnnotatorBiochemParInner1"></td><td class="AutoAnnotatorBiochemParInner2">Theoretical pI:</td><td class="AutoAnnotatorBiochemParInner3">5.55</td></tr><tr><td class="AutoAnnotatorBiochemParInner1"></td><td class="AutoAnnotatorBiochemParInner2">Extinction coefficient at 280 nm [M<sup>-1</sup> cm<sup>-1</sup>]:</td><td class="AutoAnnotatorBiochemParInner3">50310 / 50498 (all Cys red/ox)</td></tr></table></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Plot for hydrophobicity, charge, predicted secondary structure, solvent accessability, transmembrane helices and disulfid bridges</strong> <input type='button' id='hydrophobicity_charge_button' onclick='show_or_hide_plot_1380912466041()' value='Show'><span id="hydrophobicity_charge_explanation"></span><div id="hydrophobicity_charge_container" style='display:none'><div id="hydrophobicity_charge_placeholder0" style="width:100%;height:150px"></div><div id="hydrophobicity_charge_placeholder1" style="width:100%;height:150px"></div><div id="hydrophobicity_charge_placeholder2" style="width:100%;height:150px"></div></div></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Codon usage</strong><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorCodonUsage1"></td><td class="AutoAnnotatorCodonUsage2">Organism:</td><td class="AutoAnnotatorCodonUsage3"><i>E. coli</i></td><td class="AutoAnnotatorCodonUsage3"><i>B. subtilis</i></td><td class="AutoAnnotatorCodonUsage3"><i>S. cerevisiae</i></td><td class="AutoAnnotatorCodonUsage3"><i>A. thaliana</i></td><td class="AutoAnnotatorCodonUsage3"><i>P. patens</i></td><td class="AutoAnnotatorCodonUsage3">Mammals</td></tr><tr><td class="AutoAnnotatorCodonUsage1"></td><td class="AutoAnnotatorCodonUsage2">Codon quality (<a href="http://en.wikipedia.org/wiki/Codon_Adaptation_Index">CAI</a>):</td><td class="AutoAnnotatorCodonUsage3">good (0.74)</td><td class="AutoAnnotatorCodonUsage3">good (0.77)</td><td class="AutoAnnotatorCodonUsage3">good (0.75)</td><td class="AutoAnnotatorCodonUsage3">good (0.80)</td><td class="AutoAnnotatorCodonUsage3">good (0.76)</td><td class="AutoAnnotatorCodonUsage3">good (0.66)</td></tr></table></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Alignments</strong> (obtained from <a href='http://predictprotein.org'>PredictProtein.org</a>)<table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorAlignment1"></td><td class="AutoAnnotatorAlignment2">SwissProt:</td><td class="AutoAnnotatorAlignment3"><a href='http://www.uniprot.org/uniprot/P05789'>P05789</a> (100% identity on 418 AAs), <a href='http://www.uniprot.org/uniprot/P07684'>P07684</a> (24% identity on 381 AAs)</td></tr><tr><td class="AutoAnnotatorAlignment1"></td><td class="AutoAnnotatorAlignment2">TrEML:</td><td class="AutoAnnotatorAlignment3"><a href='http://www.uniprot.org/uniprot/Q2V0Y9'>Q2V0Y9</a> (100% identity on 418 AAs), <a href='http://www.uniprot.org/uniprot/D7J5T5'>D7J5T5</a> (44% identity on 402 AAs)</td></tr><tr><td class="AutoAnnotatorAlignment1"></td><td class="AutoAnnotatorAlignment2">PDB:</td><td class="AutoAnnotatorAlignment3"><a href='http://www.rcsb.org/pdb/explore/explore.do?structureId=2qgm'>2qgm</a> (20% identity on 373 AAs), <a href='http://www.rcsb.org/pdb/explore/explore.do?structureId=2rad'>2rad</a> (19% identity on 386 AAs)</td></tr></table></td></tr><tr><th id='AutoAnnotatorHeader' colspan="2"><strong>Predictions</strong> (obtained from <a href='http://predictprotein.org'>PredictProtein.org</a>)</th></tr><tr><td class="AutoAnnotatorLocalizationOuter"><strong>Subcellular Localization</strong> (reliability in brackets)<table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorLocalization1"></td><td class="AutoAnnotatorLocalization2">Archaea:</td><td class="AutoAnnotatorLocalization3">cytosol (100%)</td></tr><tr><td class="AutoAnnotatorLocalization1"></td><td class="AutoAnnotatorLocalization2">Bacteria:</td><td class="AutoAnnotatorLocalization3">cytosol (100%)</td></tr><tr><td class="AutoAnnotatorLocalization1"></td><td class="AutoAnnotatorLocalization2">Eukarya:</td><td class="AutoAnnotatorLocalization3">cytosol (58%)</td></tr></table></td><td class="AutoAnnotatorGOOuter"><strong>Gene Ontology</strong> (reliability in brackets)<br><table class="AutoAnnotatorNoBorder"><tr><td class='AutoAnnotatorGO1'></td><td class='AutoAnnotatorGO2'>Molecular Function Ontology:</td><td class='AutoAnnotatorGO3'> - </td></tr><tr><td class='AutoAnnotatorGO1'></td><td class='AutoAnnotatorGO2'>Biological Process Ontology:</td><td class='AutoAnnotatorGO3'><a href='http://amigo.geneontology.org/cgi-bin/amigo/term_details?term=GO:0046677'>GO:0046677</a> (51%)</td></tr><tr><td class='AutoAnnotatorGO1'> </td><td class='AutoAnnotatorGO2'> </td><td class='AutoAnnotatorGO3'> </td></tr></table></td></tr><tr><td class="AutoAnnotator1col" colspan="2"><strong>Predicted features:</strong><table class="AutoAnnotatorNoBorder"><tr><td class="AutoAnnotatorPredFeat1"></td><td class="AutoAnnotatorPredFeat2a">Disulfid bridges:</td><td class="AutoAnnotatorPredFeat3"> - </td></tr><tr><td class="AutoAnnotatorPredFeat1"></td><td class="AutoAnnotatorPredFeat2a">Transmembrane helices:</td><td class="AutoAnnotatorPredFeat3"> - </td></tr></table></td></tr><tr><td class="AutoAnnotator1col" colspan="2"> The BioBrick-AutoAnnotator was created by <a href="http://2013.igem.org/Team:TU-Munich">TU-Munich 2013</a> iGEM team. For more information please see the <a href="http://2013.igem.org/Team:TU-Munich/Results/Software">documentation</a>.<br>If you have any questions, comments or suggestions, please leave us a <a href="http://2013.igem.org/Team:TU-Munich/Results/AutoAnnotator">comment</a>.</td></tr></table></div><br><!-- IMPORTANT: DON'T REMOVE THIS LINE, OTHERWISE NOT SUPPORTED FOR IE BEFORE 9 --><!--[if lte IE 8]><script language="javascript" type="text/javascript" src="excanvas.min.js"></script><![endif]--><script type='text/javascript' src='http://code.jquery.com/jquery-1.10.0.min.js'></script><script type='text/javascript' src='http://2013.igem.org/Team:TU-Munich/Flot.js?action=raw&ctype=text/js'></script><script>function show_or_hide_plot_1380912466041(){hydrophobicity_datapoints = [[2.5,0.32],[3.5,-0.76],[4.5,-1.82],[5.5,-1.92],[6.5,-0.18],[7.5,-1.52],[8.5,-1.52],[9.5,-1.60],[10.5,-2.06],[11.5,-2.00],[12.5,-1.54],[13.5,-0.28],[14.5,-0.28],[15.5,1.12],[16.5,-0.48],[17.5,-0.80],[18.5,-1.68],[19.5,-1.60],[20.5,-3.06],[21.5,-3.06],[22.5,-2.68],[23.5,-3.06],[24.5,-3.06],[25.5,-1.80],[26.5,-1.88],[27.5,-1.94],[28.5,-0.48],[29.5,-0.68],[30.5,-2.02],[31.5,-0.34],[32.5,0.88],[33.5,0.04],[34.5,0.24],[35.5,0.88],[36.5,-0.92],[37.5,-0.98],[38.5,-0.06],[39.5,1.00],[40.5,1.90],[41.5,2.72],[42.5,1.18],[43.5,-0.36],[44.5,-0.88],[45.5,-2.28],[46.5,-1.64],[47.5,-0.04],[48.5,-0.12],[49.5,-0.66],[50.5,0.54],[51.5,0.54],[52.5,0.40],[53.5,1.94],[54.5,1.74],[55.5,0.54],[56.5,-0.16],[57.5,-0.16],[58.5,-0.16],[59.5,-0.16],[60.5,1.04],[61.5,1.74],[62.5,1.88],[63.5,0.42],[64.5,0.62],[65.5,0.82],[66.5,0.18],[67.5,-0.16],[68.5,0.40],[69.5,0.96],[70.5,0.76],[71.5,1.20],[72.5,1.20],[73.5,0.64],[74.5,1.34],[75.5,0.70],[76.5,0.90],[77.5,0.70],[78.5,0.70],[79.5,0.06],[80.5,-0.56],[81.5,-0.22],[82.5,0.32],[83.5,0.32],[84.5,-0.30],[85.5,0.22],[86.5,0.22],[87.5,-1.32],[88.5,-0.70],[89.5,-0.70],[90.5,-0.60],[91.5,-1.64],[92.5,-1.70],[93.5,-2.32],[94.5,-2.32],[95.5,-1.34],[96.5,-1.28],[97.5,-1.48],[98.5,-0.22],[99.5,1.24],[100.5,-0.44],[101.5,-1.00],[102.5,0.46],[103.5,-0.36],[104.5,-1.38],[105.5,-0.92],[106.5,-0.98],[107.5,-2.24],[108.5,-1.22],[109.5,-1.74],[110.5,-1.56],[111.5,-1.00],[112.5,0.26],[113.5,0.26],[114.5,0.86],[115.5,1.76],[116.5,1.00],[117.5,-0.26],[118.5,-1.28],[119.5,-1.80],[120.5,-3.34],[121.5,-2.08],[122.5,-1.02],[123.5,-1.54],[124.5,-1.54],[125.5,-1.54],[126.5,-1.00],[127.5,-1.50],[128.5,-0.16],[129.5,1.30],[130.5,2.00],[131.5,2.00],[132.5,1.44],[133.5,1.78],[134.5,0.70],[135.5,-0.12],[136.5,-1.72],[137.5,-1.10],[138.5,-2.08],[139.5,-1.92],[140.5,-1.28],[141.5,0.18],[142.5,-0.06],[143.5,-0.68],[144.5,-0.14],[145.5,-0.58],[146.5,-0.44],[147.5,0.72],[148.5,0.78],[149.5,-0.30],[150.5,0.96],[151.5,0.62],[152.5,-1.12],[153.5,-0.62],[154.5,0.44],[155.5,-0.82],[156.5,-2.16],[157.5,-1.96],[158.5,-1.46],[159.5,-1.06],[160.5,-1.00],[161.5,0.68],[162.5,2.28],[163.5,1.22],[164.5,-0.24],[165.5,0.76],[166.5,0.42],[167.5,-1.18],[168.5,0.42],[169.5,1.48],[170.5,0.34],[171.5,-1.00],[172.5,0.60],[173.5,-1.00],[174.5,-1.62],[175.5,-0.28],[176.5,0.34],[177.5,-0.70],[178.5,-0.16],[179.5,-0.60],[180.5,-0.80],[181.5,-0.28],[182.5,0.62],[183.5,0.08],[184.5,1.54],[185.5,0.54],[186.5,-0.52],[187.5,-0.52],[188.5,0.04],[189.5,-1.42],[190.5,-0.94],[191.5,-0.94],[192.5,-2.48],[193.5,-1.84],[194.5,-2.04],[195.5,-1.12],[196.5,-0.56],[197.5,0.06],[198.5,-1.14],[199.5,0.52],[200.5,0.12],[201.5,-0.64],[202.5,0.36],[203.5,0.28],[204.5,0.36],[205.5,-0.90],[206.5,0.76],[207.5,-0.78],[208.5,0.36],[209.5,-0.10],[210.5,1.16],[211.5,0.08],[212.5,1.68],[213.5,0.68],[214.5,1.20],[215.5,0.14],[216.5,-0.32],[217.5,-1.48],[218.5,-0.92],[219.5,-0.92],[220.5,-0.92],[221.5,-0.92],[222.5,-0.92],[223.5,-1.54],[224.5,-2.58],[225.5,-0.98],[226.5,0.56],[227.5,0.18],[228.5,0.62],[229.5,1.12],[230.5,-0.48],[231.5,-1.32],[232.5,-0.30],[233.5,0.86],[234.5,0.24],[235.5,1.70],[236.5,1.08],[237.5,0.44],[238.5,-1.16],[239.5,-0.00],[240.5,-1.46],[241.5,-0.40],[242.5,0.24],[243.5,0.78],[244.5,-0.20],[245.5,1.06],[246.5,1.60],[247.5,1.06],[248.5,1.14],[249.5,1.14],[250.5,0.50],[251.5,-0.02],[252.5,-0.56],[253.5,-0.56],[254.5,-1.18],[255.5,-0.72],[256.5,-1.18],[257.5,-0.12],[258.5,-0.82],[259.5,-0.82],[260.5,-1.98],[261.5,-2.16],[262.5,-2.14],[263.5,-1.00],[264.5,-1.00],[265.5,-0.38],[266.5,0.72],[267.5,1.10],[268.5,0.56],[269.5,0.62],[270.5,1.54],[271.5,-0.08],[272.5,-1.54],[273.5,-1.68],[274.5,-1.74],[275.5,-2.66],[276.5,-2.58],[277.5,-2.58],[278.5,-3.04],[279.5,-1.50],[280.5,-0.44],[281.5,1.10],[282.5,2.64],[283.5,3.78],[284.5,2.30],[285.5,0.70],[286.5,0.22],[287.5,-1.26],[288.5,-0.72],[289.5,-0.78],[290.5,-0.86],[291.5,-1.36],[292.5,-1.04],[293.5,-1.04],[294.5,0.42],[295.5,0.94],[296.5,0.38],[297.5,0.62],[298.5,0.28],[299.5,0.28],[300.5,0.38],[301.5,1.58],[302.5,2.42],[303.5,1.54],[304.5,1.16],[305.5,1.24],[306.5,0.04],[307.5,-1.62],[308.5,-0.54],[309.5,-1.70],[310.5,-2.32],[311.5,-1.26],[312.5,0.54],[313.5,-0.30],[314.5,-0.22],[315.5,-0.22],[316.5,-0.84],[317.5,-1.36],[318.5,-1.44],[319.5,0.02],[320.5,0.64],[321.5,1.80],[322.5,1.28],[323.5,1.28],[324.5,0.26],[325.5,0.18],[326.5,-0.80],[327.5,-1.30],[328.5,-1.28],[329.5,-0.66],[330.5,-0.14],[331.5,0.70],[332.5,1.08],[333.5,0.90],[334.5,-0.16],[335.5,0.32],[336.5,-1.14],[337.5,-1.02],[338.5,-1.48],[339.5,-1.04],[340.5,-1.96],[341.5,-0.70],[342.5,-1.46],[343.5,0.16],[344.5,-0.28],[345.5,-0.90],[346.5,-0.90],[347.5,-0.70],[348.5,-0.78],[349.5,-0.78],[350.5,-0.78],[351.5,-1.66],[352.5,-1.66],[353.5,-3.12],[354.5,-2.50],[355.5,-1.96],[356.5,-0.80],[357.5,-0.80],[358.5,-0.88],[359.5,-0.44],[360.5,0.62],[361.5,-0.38],[362.5,0.24],[363.5,1.52],[364.5,1.08],[365.5,1.02],[366.5,1.04],[367.5,0.42],[368.5,-0.24],[369.5,0.40],[370.5,0.40],[371.5,1.10],[372.5,2.36],[373.5,1.62],[374.5,0.36],[375.5,0.42],[376.5,-0.46],[377.5,-1.72],[378.5,-1.52],[379.5,-0.06],[380.5,-1.66],[381.5,-1.50],[382.5,0.10],[383.5,0.48],[384.5,-0.98],[385.5,0.10],[386.5,1.16],[387.5,-0.64],[388.5,0.24],[389.5,0.24],[390.5,-0.30],[391.5,-0.30],[392.5,0.34],[393.5,0.16],[394.5,0.08],[395.5,0.60],[396.5,-1.00],[397.5,0.02],[398.5,-1.06],[399.5,-1.06],[400.5,-1.06],[401.5,0.20],[402.5,-1.26],[403.5,-0.64],[404.5,1.04],[405.5,1.24],[406.5,-0.02],[407.5,1.58],[408.5,0.96],[409.5,-0.72],[410.5,-1.44],[411.5,-0.90],[412.5,-0.96],[413.5,-0.42],[414.5,-0.34],[415.5,0.66],[416.5,1.66],[417.5,0.56],[418.5,0.02],[419.5,0.58],[420.5,-0.34]];charge_datapoints = [[2.5,0.20],[3.5,0.00],[4.5,-0.20],[5.5,-0.20],[6.5,-0.40],[7.5,-0.20],[8.5,-0.20],[9.5,0.20],[10.5,0.30],[11.5,0.30],[12.5,0.10],[13.5,0.30],[14.5,0.30],[15.5,0.20],[16.5,0.20],[17.5,0.30],[18.5,0.30],[19.5,-0.10],[20.5,-0.30],[21.5,-0.30],[22.5,-0.40],[23.5,-0.60],[24.5,-0.60],[25.5,-0.40],[26.5,-0.20],[27.5,-0.20],[28.5,-0.00],[29.5,0.40],[30.5,0.60],[31.5,0.40],[32.5,0.40],[33.5,0.40],[34.5,-0.00],[35.5,-0.20],[36.5,-0.00],[37.5,-0.00],[38.5,-0.00],[39.5,0.20],[40.5,0.20],[41.5,-0.00],[42.5,-0.20],[43.5,-0.20],[44.5,-0.20],[45.5,-0.10],[46.5,-0.10],[47.5,0.10],[48.5,0.30],[49.5,0.10],[50.5,-0.00],[51.5,-0.00],[52.5,-0.00],[53.5,-0.20],[54.5,0.20],[55.5,0.30],[56.5,0.30],[57.5,0.30],[58.5,0.30],[59.5,0.30],[60.5,0.20],[61.5,0.20],[62.5,0.20],[63.5,-0.00],[64.5,-0.40],[65.5,-0.40],[66.5,-0.40],[67.5,-0.40],[68.5,-0.20],[69.5,-0.00],[70.5,-0.00],[71.5,-0.00],[72.5,-0.00],[73.5,-0.20],[74.5,-0.20],[75.5,-0.20],[76.5,-0.20],[77.5,-0.20],[78.5,-0.20],[79.5,-0.20],[80.5,-0.20],[81.5,-0.20],[82.5,-0.20],[83.5,-0.00],[84.5,-0.00],[85.5,-0.00],[86.5,-0.00],[87.5,0.10],[88.5,0.10],[89.5,0.10],[90.5,0.10],[91.5,0.10],[92.5,-0.20],[93.5,-0.40],[94.5,-0.60],[95.5,-0.60],[96.5,-0.60],[97.5,-0.20],[98.5,-0.00],[99.5,0.20],[100.5,0.40],[101.5,0.50],[102.5,0.30],[103.5,0.30],[104.5,0.30],[105.5,0.10],[106.5,-0.20],[107.5,-0.40],[108.5,-0.40],[109.5,-0.20],[110.5,-0.20],[111.5,0.00],[112.5,0.20],[113.5,0.20],[114.5,0.00],[115.5,0.00],[116.5,0.20],[117.5,0.00],[118.5,0.00],[119.5,0.00],[120.5,0.20],[121.5,0.00],[122.5,0.20],[123.5,0.40],[124.5,0.20],[125.5,0.20],[126.5,0.20],[127.5,0.20],[128.5,0.00],[129.5,0.20],[130.5,0.00],[131.5,0.00],[132.5,-0.20],[133.5,-0.20],[134.5,-0.20],[135.5,0.00],[136.5,0.00],[137.5,0.20],[138.5,0.20],[139.5,0.20],[140.5,0.00],[141.5,0.00],[142.5,0.00],[143.5,0.00],[144.5,0.00],[145.5,-0.20],[146.5,-0.20],[147.5,-0.20],[148.5,-0.10],[149.5,-0.30],[150.5,-0.10],[151.5,-0.10],[152.5,0.10],[153.5,-0.00],[154.5,0.20],[155.5,-0.00],[156.5,0.20],[157.5,-0.20],[158.5,-0.20],[159.5,-0.20],[160.5,0.10],[161.5,-0.10],[162.5,0.10],[163.5,-0.10],[164.5,-0.30],[165.5,-0.40],[166.5,-0.40],[167.5,-0.40],[168.5,-0.20],[169.5,0.00],[170.5,0.20],[171.5,0.40],[172.5,0.40],[173.5,0.20],[174.5,0.20],[175.5,0.00],[176.5,-0.20],[177.5,-0.20],[178.5,0.00],[179.5,0.00],[180.5,0.00],[181.5,0.00],[182.5,0.00],[183.5,0.00],[184.5,0.00],[185.5,0.10],[186.5,-0.10],[187.5,-0.10],[188.5,-0.10],[189.5,-0.30],[190.5,-0.40],[191.5,-0.40],[192.5,-0.20],[193.5,-0.20],[194.5,0.20],[195.5,0.20],[196.5,0.40],[197.5,0.20],[198.5,0.20],[199.5,0.00],[200.5,0.00],[201.5,0.20],[202.5,0.20],[203.5,0.40],[204.5,0.40],[205.5,0.60],[206.5,0.40],[207.5,0.20],[208.5,0.00],[209.5,0.00],[210.5,-0.20],[211.5,-0.20],[212.5,0.00],[213.5,0.10],[214.5,0.10],[215.5,-0.10],[216.5,0.10],[217.5,0.10],[218.5,0.00],[219.5,0.00],[220.5,0.00],[221.5,0.00],[222.5,0.00],[223.5,-0.20],[224.5,-0.20],[225.5,0.00],[226.5,-0.20],[227.5,-0.10],[228.5,0.10],[229.5,0.10],[230.5,0.10],[231.5,0.10],[232.5,0.00],[233.5,0.00],[234.5,0.00],[235.5,0.00],[236.5,-0.20],[237.5,-0.20],[238.5,-0.20],[239.5,-0.20],[240.5,-0.20],[241.5,0.00],[242.5,0.00],[243.5,0.00],[244.5,0.00],[245.5,0.00],[246.5,0.00],[247.5,0.00],[248.5,0.00],[249.5,0.00],[250.5,0.00],[251.5,0.00],[252.5,0.00],[253.5,0.00],[254.5,-0.20],[255.5,-0.20],[256.5,-0.20],[257.5,-0.20],[258.5,0.00],[259.5,0.00],[260.5,0.20],[261.5,0.20],[262.5,0.20],[263.5,0.00],[264.5,0.00],[265.5,-0.20],[266.5,-0.20],[267.5,-0.20],[268.5,-0.20],[269.5,0.10],[270.5,0.10],[271.5,0.30],[272.5,0.30],[273.5,0.30],[274.5,0.20],[275.5,0.20],[276.5,-0.20],[277.5,-0.20],[278.5,0.00],[279.5,0.00],[280.5,0.00],[281.5,0.20],[282.5,0.20],[283.5,0.00],[284.5,0.10],[285.5,0.10],[286.5,0.10],[287.5,0.20],[288.5,0.20],[289.5,0.10],[290.5,0.30],[291.5,0.30],[292.5,0.20],[293.5,0.20],[294.5,0.20],[295.5,0.00],[296.5,-0.20],[297.5,-0.20],[298.5,-0.20],[299.5,-0.20],[300.5,-0.20],[301.5,0.00],[302.5,0.00],[303.5,0.00],[304.5,0.00],[305.5,0.00],[306.5,0.00],[307.5,0.20],[308.5,0.20],[309.5,0.40],[310.5,0.40],[311.5,0.40],[312.5,0.20],[313.5,0.20],[314.5,-0.20],[315.5,-0.40],[316.5,-0.40],[317.5,-0.40],[318.5,-0.40],[319.5,-0.20],[320.5,0.00],[321.5,0.00],[322.5,0.00],[323.5,0.00],[324.5,0.00],[325.5,0.00],[326.5,0.00],[327.5,0.10],[328.5,0.10],[329.5,0.10],[330.5,0.10],[331.5,0.10],[332.5,0.00],[333.5,0.00],[334.5,-0.20],[335.5,-0.20],[336.5,-0.40],[337.5,-0.40],[338.5,-0.20],[339.5,0.00],[340.5,0.00],[341.5,0.20],[342.5,0.40],[343.5,0.20],[344.5,0.00],[345.5,0.00],[346.5,0.00],[347.5,-0.20],[348.5,-0.20],[349.5,0.00],[350.5,-0.20],[351.5,-0.20],[352.5,-0.20],[353.5,-0.40],[354.5,-0.40],[355.5,-0.20],[356.5,-0.20],[357.5,-0.40],[358.5,0.00],[359.5,0.00],[360.5,0.00],[361.5,0.00],[362.5,0.20],[363.5,0.00],[364.5,0.00],[365.5,0.00],[366.5,0.00],[367.5,0.00],[368.5,0.00],[369.5,0.00],[370.5,0.00],[371.5,0.00],[372.5,0.00],[373.5,0.20],[374.5,0.20],[375.5,0.20],[376.5,0.20],[377.5,0.00],[378.5,-0.40],[379.5,-0.40],[380.5,-0.40],[381.5,-0.40],[382.5,-0.20],[383.5,0.00],[384.5,0.00],[385.5,0.00],[386.5,0.00],[387.5,0.20],[388.5,0.20],[389.5,0.00],[390.5,0.00],[391.5,0.00],[392.5,-0.20],[393.5,-0.20],[394.5,0.20],[395.5,0.20],[396.5,0.00],[397.5,0.00],[398.5,-0.20],[399.5,-0.20],[400.5,-0.20],[401.5,0.00],[402.5,-0.20],[403.5,0.00],[404.5,-0.20],[405.5,-0.20],[406.5,-0.20],[407.5,0.00],[408.5,-0.20],[409.5,0.00],[410.5,0.00],[411.5,0.00],[412.5,0.00],[413.5,0.20],[414.5,-0.20],[415.5,-0.20],[416.5,-0.20],[417.5,-0.20],[418.5,-0.40],[419.5,-0.20],[420.5,-0.20]];dis_datapoints = [];trans_datapoints = [];sec_helix_datapoints = [[3,12],[28,34],[50,67],[81,90],[95,105],[109,121],[144,156],[159,172],[181,188],[193,200],[201,212],[215,218],[224,249],[261,276],[305,314],[359,365],[380,382],[389,391],[401,403]];sec_strand_datapoints = [[39,43],[72,76],[128,134],[282,286],[319,325],[330,334],[371,374],[406,411]];acc_exposed_datapoints = [[1,4],[7,7],[10,11],[15,15],[17,17],[19,19],[21,24],[26,26],[29,29],[33,33],[36,37],[66,67],[90,90],[92,92],[95,96],[99,100],[109,109],[120,120],[123,127],[141,141],[155,156],[158,160],[163,163],[167,167],[170,170],[173,174],[176,177],[179,179],[189,189],[191,193],[197,197],[200,200],[204,204],[207,207],[210,211],[214,214],[218,219],[221,221],[223,224],[251,253],[274,274],[277,279],[313,313],[316,317],[337,341],[343,343],[345,345],[347,347],[350,350],[353,353],[356,356],[364,365],[367,367],[369,369],[380,381],[384,384],[387,387],[397,397],[399,399],[402,402],[412,412],[414,414],[417,417],[420,423]];acc_buried_datapoints = [[5,5],[9,9],[13,13],[16,16],[28,28],[31,31],[35,35],[38,38],[40,50],[53,54],[56,61],[64,65],[69,79],[82,82],[84,86],[88,89],[91,91],[98,98],[101,102],[105,108],[111,115],[117,118],[122,122],[129,137],[142,143],[146,147],[149,151],[153,154],[161,162],[165,165],[168,169],[172,172],[183,187],[198,199],[201,202],[205,205],[209,209],[212,213],[216,217],[225,225],[228,232],[234,238],[240,242],[244,249],[258,258],[260,260],[262,262],[264,266],[268,270],[272,273],[280,280],[282,292],[294,294],[296,296],[298,298],[301,301],[304,311],[315,315],[319,327],[329,334],[346,346],[351,351],[358,359],[362,363],[366,366],[371,375],[378,379],[382,382],[388,388],[391,396],[400,400],[404,410],[415,415]];flot_plot_options = []; flot_plot_options[0] = {grid: {borderWidth: {top: 0,right: 0,bottom: 0,left: 0}},legend: {show: false},xaxes: [{show: true,min: 0,max: 200,ticks: [[0.5, '1'], [24.5, '25'], [49.5, '50'], [74.5, '75'], [99.5, '100'], [124.5, '125'], [149.5, '150'], [174.5, '175'], [199.5, '200']],tickLength: -5}],yaxes: [{show: true,ticks: [[0, '0'], [4.5,'hydro-<br>phobic '], [-4.5,'hydro-<br>philic ']],min: -4.5,max: +4.5,font: {size: 12,lineHeight: 14,style: 'italic',weight: 'bold',family: 'sans-serif',variant: 'small-caps',color: 'rgba(100,149,237,1)'}},{show: true,ticks: [[0, ''], [1,'positive<br> charge'], [-1,'negative<br> charge']],position: 'right',min: -1,max: 1,font: {size: 12,lineHeight: 14,style: 'italic',weight: 'bold',family: 'sans-serif',variant: 'small-caps',color: 'rgba(255,99,71,1)'}}]};number_of_plots = 3;for ( plot_num = 1 ; plot_num < number_of_plots ; plot_num ++){flot_plot_options[plot_num] = $.extend(true, {} ,flot_plot_options[0]);flot_plot_options[plot_num].xaxes = [{min: plot_num*200,max: (plot_num + 1)*200,ticks: [ [plot_num*200 + 0.5, (plot_num*200 + 1).toString()], [plot_num*200 + 24.5, (plot_num*200 + 25).toString()], [plot_num*200 + 49.5, (plot_num*200 + 50).toString()], [plot_num*200 + 74.5, (plot_num*200 + 75).toString()], [plot_num*200 + 99.5, (plot_num*200 + 100).toString()], [plot_num*200 + 124.5, (plot_num*200 + 125).toString()], [plot_num*200 + 149.5, (plot_num*200 + 150).toString()], [plot_num*200 + 174.5, (plot_num*200 + 175).toString()], [plot_num*200 + 199.5, (plot_num*200 + 200).toString()] ],tickLength: -5}];};try {if( $('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_button').val() =='Show' ){$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_container').css('display','block');$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_button').val('Hide');var description_html = '<div id=\'AutoAnnotator_plot_selectors\'>';description_html = description_html + '<br> <input type=\'checkbox\' id=\'hydrophobicity_checkbox\' checked=\'checked\'> Moving average over 5 amino acids for hydrophobicity (<img src=\'https://static.igem.org/mediawiki/2013/e/e9/TUM13_hydrophobicity_icon.png\' alt=\'blue graph\' height=\'10\'></img>)';description_html = description_html + '<br> <input type=\'checkbox\' id=\'charge_checkbox\' checked=\'checked\'> Moving average over 5 amino acids for charge (<img src=\'https://static.igem.org/mediawiki/2013/3/3e/TUM13_charge_icon.png\' alt=\'red graph\' height=\'10\'></img>)';description_html = description_html + '<br> <input type=\'checkbox\' id=\'dis_checkbox\' checked=\'checked\'> Predicted disulfid bridges (<img src=\'https://static.igem.org/mediawiki/2013/2/28/TUM13_dis_icon.png\' alt=\'yellow circle\' height=\'10\'></img>) with the number of the bridge in the center';description_html = description_html + '<br> <input type=\'checkbox\' id=\'trans_checkbox\' checked=\'checked\'> Predicted transmembrane helices (<img src=\'https://static.igem.org/mediawiki/2013/7/78/TUM13_trans_icon.png\' alt=\'turquois bars\' height=\'10\'></img>)';description_html = description_html + '<br> <input type=\'checkbox\' id=\'sec_checkbox\' checked=\'checked\'> Predicted secondary structure: Helices (<img src=\'https://static.igem.org/mediawiki/2013/b/bf/TUM13_helix_icon.png\' alt=\'violet bars\' height=\'10\'></img>) and beta-strands (<img src=\'https://static.igem.org/mediawiki/2013/b/bf/TUM13_strand_icon.png\' alt=\'yellow bars\' height=\'10\'></img>)';description_html = description_html + '<br> <input type=\'checkbox\' id=\'acc_checkbox\' checked=\'checked\'> Predicted solvent accessability: Exposed (<img src=\'https://static.igem.org/mediawiki/2013/1/16/TUM13_exposed_icon.png\' alt=\'blue bars\' height=\'10\'></img>) and buried (<img src=\'https://static.igem.org/mediawiki/2013/0/0b/TUM13_buried_icon.png\' alt=\'green bars\' height=\'10\'></img>) residues';description_html = description_html + '<br></div>';$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_explanation').html(description_html);plot_according_to_selectors_1380912466041();$('#AutoAnnotator_container_1380912466041 #AutoAnnotator_plot_selectors').find('input').click(plot_according_to_selectors_1380912466041);}else{$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_container').css('display','none');$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_button').val('Show');$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_explanation').html('');}}catch(err){txt='There was an error with the button controlling the visibility of the plot.\n';txt=txt+'The originating error is:\n' + err + '\n\n';alert(txt);}};function plot_according_to_selectors_1380912466041(){try{var plot_datasets = [[],[]];if($('#AutoAnnotator_container_1380912466041 #hydrophobicity_checkbox').prop('checked') == true){plot_datasets[0] = { color: 'rgba(100,149,237,1)',data: hydrophobicity_datapoints,label: 'Hydrophobicity',lines: { show: true, fill: true, fillColor: 'rgba(100,149,237,0.1)' },yaxis: 1};}if($('#AutoAnnotator_container_1380912466041 #charge_checkbox').prop('checked') == true){plot_datasets[1] = {color: 'rgba(255,99,71,1)',data: charge_datapoints,label: 'Charge',lines: { show: true, fill: true, fillColor: 'rgba(255,99,71,0.1)' },yaxis: 2};}for (plot_num = 0 ; plot_num < number_of_plots ; plot_num ++){$.plot('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder'+ plot_num.toString(), plot_datasets, flot_plot_options[plot_num] );}var screen_width = $('canvas.flot-base').width(); var pos_of_first_tick = 46;var pos_of_last_tick = screen_width - 51;var tick_diff = (screen_width - 97)/199;if($('#AutoAnnotator_container_1380912466041 #dis_checkbox').prop('checked') == true){for ( j = 0 ; j < dis_datapoints.length ; j++ ){$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder' + Math.floor((dis_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_dis\' style=\'left:' + ((pos_of_first_tick - 8 + (dis_datapoints[j][0] - 1)*tick_diff - Math.floor((dis_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px;\'><b>' + (j+1) + '</b></div>');$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder' + Math.floor((dis_datapoints[j][1] - 1)/200) ).append('<div class=\'AutoAnnotator_dis\' style=\'left:' + ((pos_of_first_tick - 8 + (dis_datapoints[j][1] - 1)*tick_diff - Math.floor((dis_datapoints[j][1] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px;\'><b>' + (j+1) + '</b></div>');}}if($('#AutoAnnotator_container_1380912466041 #trans_checkbox').prop('checked') == true){for ( j = 0 ; j < trans_datapoints.length ; j++ ){$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder' + Math.floor((trans_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_trans\' style=\'width:' + (((trans_datapoints[j][1] - trans_datapoints[j][0] + 1)*tick_diff).toFixed(0)).toString() + 'px ;left:' + ((pos_of_first_tick + (trans_datapoints[j][0] - 1.5)*tick_diff - Math.floor((trans_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px\'></div>');}}if($('#AutoAnnotator_container_1380912466041 #sec_checkbox').prop('checked') == true){for ( j = 0 ; j < sec_helix_datapoints.length ; j++ ){$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder' + Math.floor((sec_helix_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_sec_helix\' style=\'width:' + (((sec_helix_datapoints[j][1] - sec_helix_datapoints[j][0] + 1)*tick_diff).toFixed(0)).toString() + 'px; left:' + ((pos_of_first_tick + (sec_helix_datapoints[j][0] - 1.5)*tick_diff - Math.floor((sec_helix_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px\'></div>');}for ( j = 0 ; j < sec_strand_datapoints.length ; j++ ){$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder' + Math.floor((sec_strand_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_sec_strand\' style=\'width:' + (((sec_strand_datapoints[j][1] - sec_strand_datapoints[j][0] + 1)*tick_diff).toFixed(0)).toString() + 'px; left:' + ((pos_of_first_tick + (sec_strand_datapoints[j][0] - 1.5)*tick_diff - Math.floor((sec_strand_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px\'></div>');}}if($('#AutoAnnotator_container_1380912466041 #acc_checkbox').prop('checked') == true){for ( j = 0 ; j < acc_buried_datapoints.length ; j++ ){$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder' + Math.floor((acc_buried_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_acc_buried\' style=\'width:' + (((acc_buried_datapoints[j][1] - acc_buried_datapoints[j][0] + 1)*tick_diff).toFixed(0)).toString() + 'px; left:' + ((pos_of_first_tick + (acc_buried_datapoints[j][0] - 1.5)*tick_diff - Math.floor((acc_buried_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px\'></div>');}for ( j = 0 ; j < acc_exposed_datapoints.length ; j++ ){$('#AutoAnnotator_container_1380912466041 #hydrophobicity_charge_placeholder' + Math.floor((acc_exposed_datapoints[j][0] - 1)/200) ).append('<div class=\'AutoAnnotator_acc_exposed\' style=\'width:' + (((acc_exposed_datapoints[j][1] - acc_exposed_datapoints[j][0] + 1)*tick_diff).toFixed(0)).toString() + 'px; left:' + ((pos_of_first_tick + (acc_exposed_datapoints[j][0] - 1.5)*tick_diff - Math.floor((acc_exposed_datapoints[j][0] - 1)/200)*200*tick_diff).toFixed(0)).toString() + 'px\'></div>');}}}catch(err){txt='There was an error while drawing the selected elements for the plot.\n';txt=txt+'The originating error is:\n' + err + '\n\n';throw(txt);}}</script></html> | ||
| + | |||
| + | ===Production and purification of recombinant EreB=== | ||
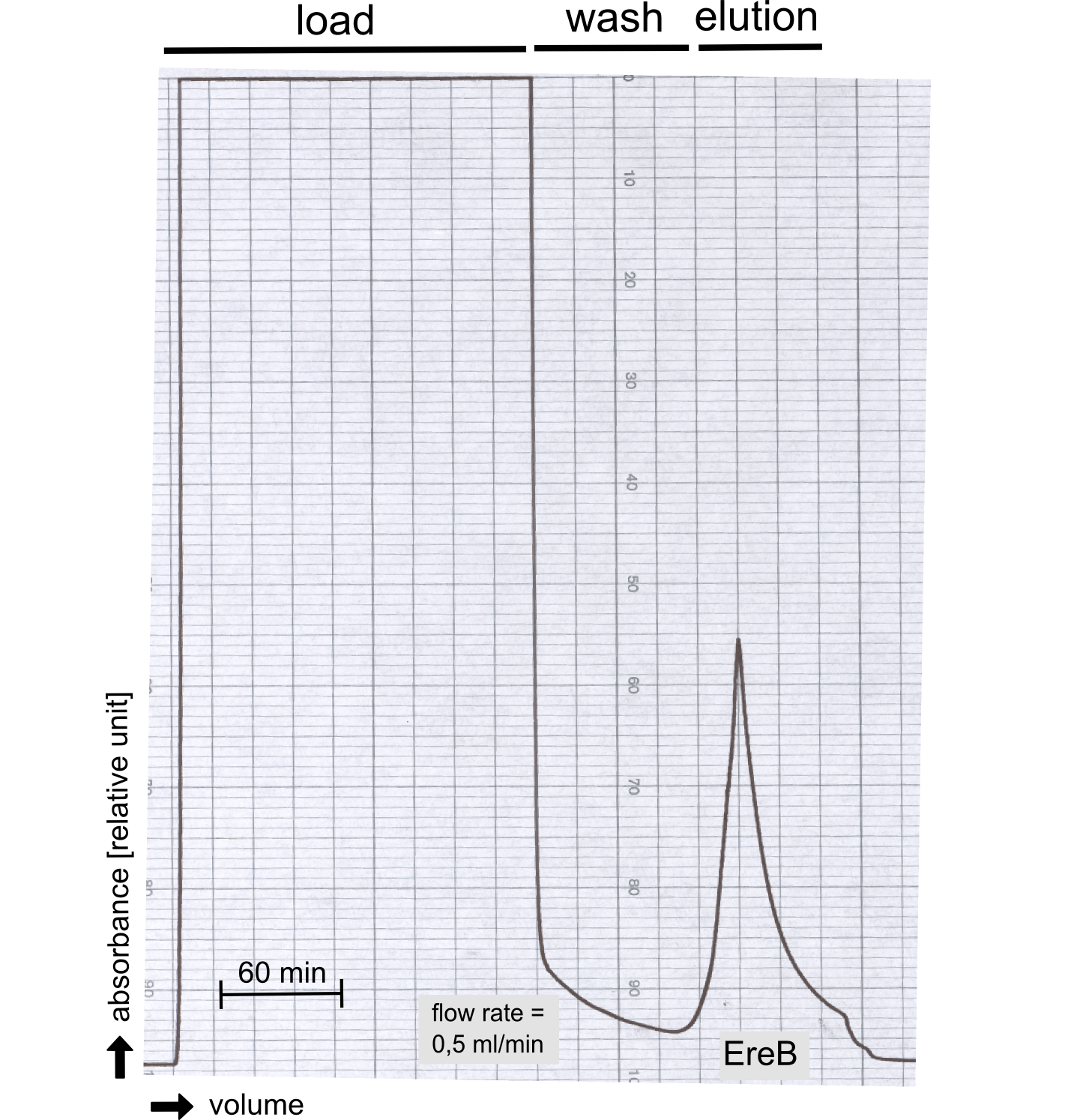
| + | [[File:TUM13_P917.png|thumb|left|320px| '''Figure A:''' Streptavidin affinity chromatography for the erythromycin esterase]] | ||
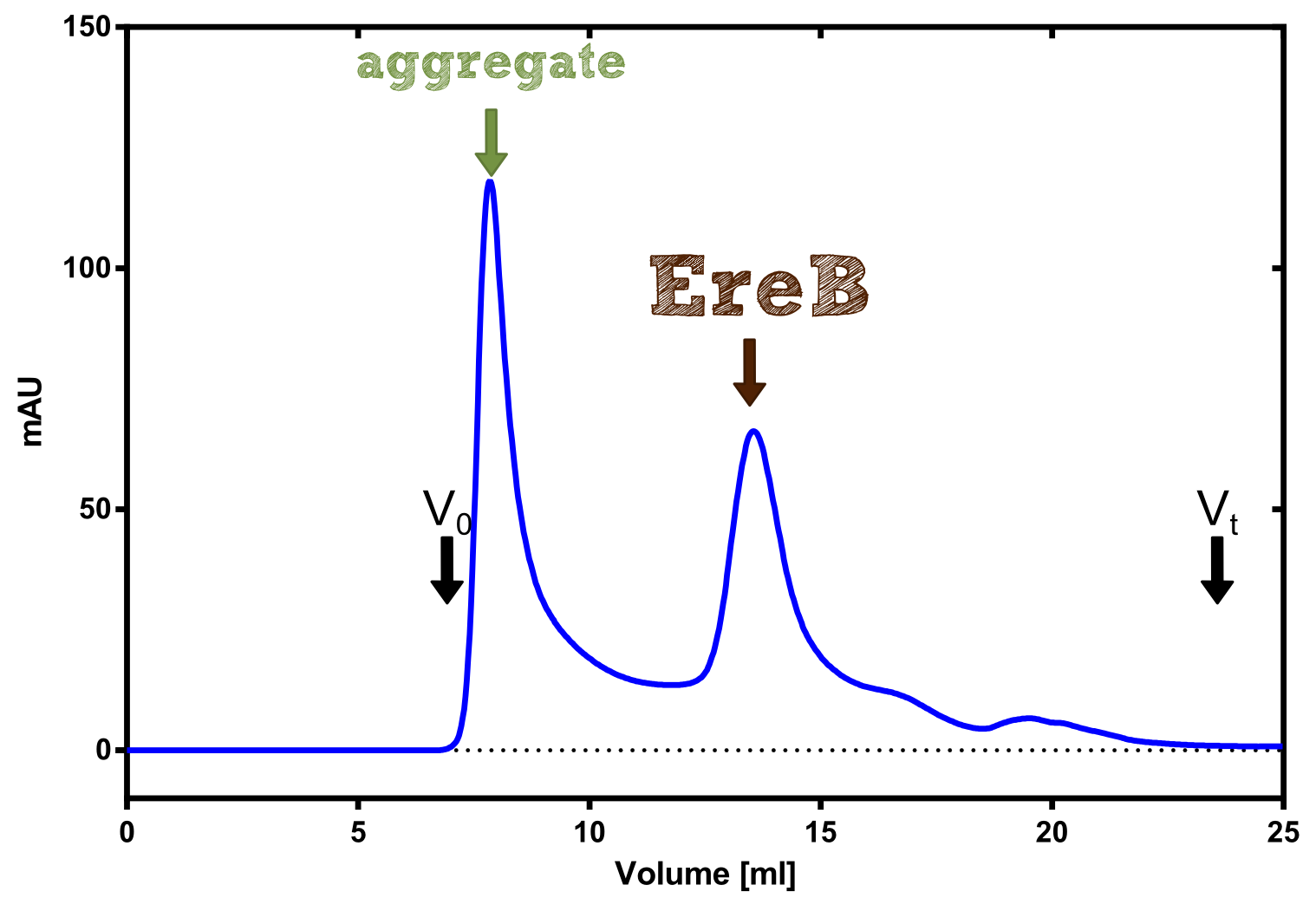
| + | [[File:TUM13_Analytprep_EreB.png|thumb|right|300px| '''Figure B:''' Analytical size exclusion chromatography on a Superdex 200 10/30 column]] | ||
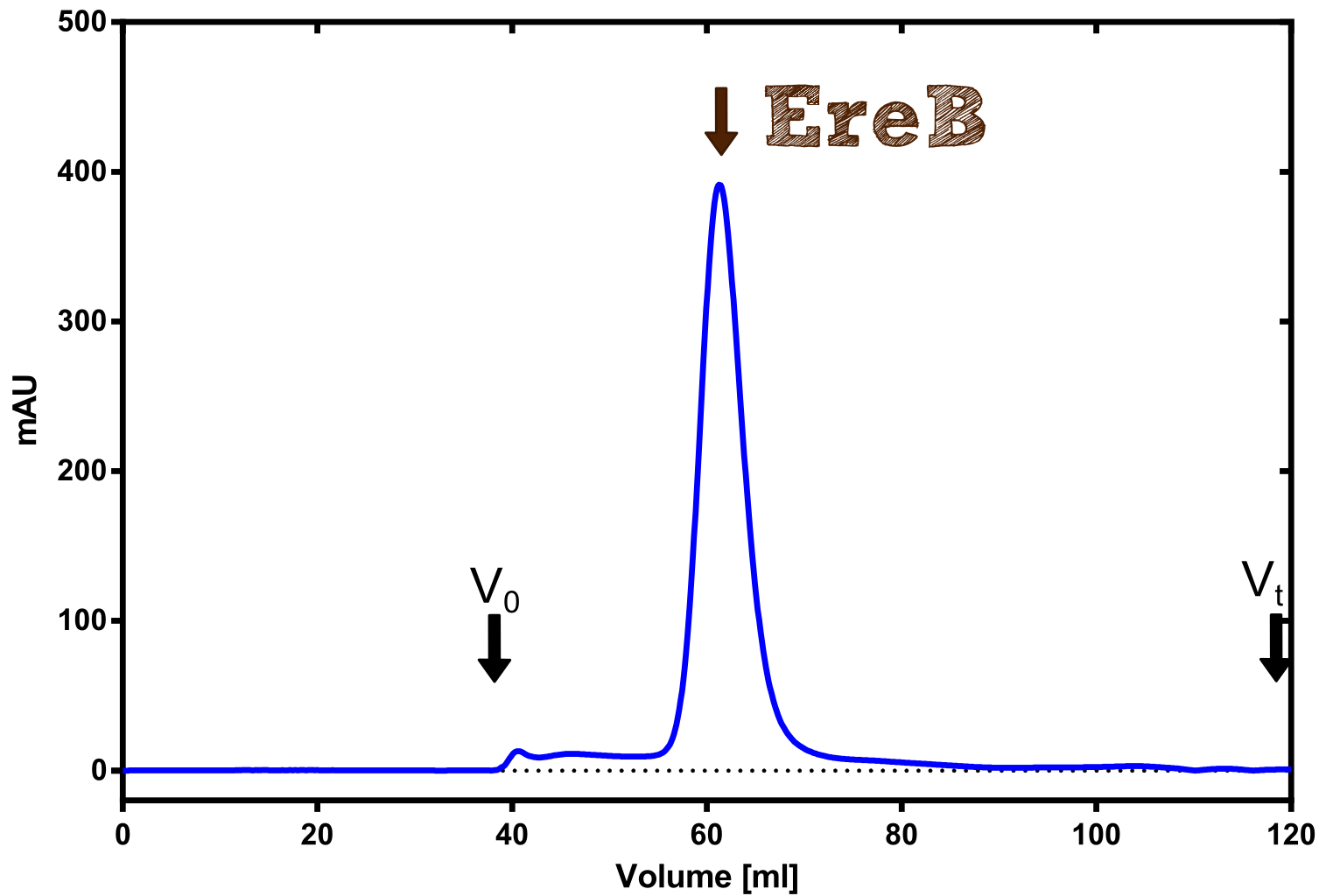
| + | [[File:TUM13_Preparative_EreB.Png|thumb|right|300px| '''Figure C:''' Preparative size exclusion chromatography on a Superdex 75 16/60 column]] | ||
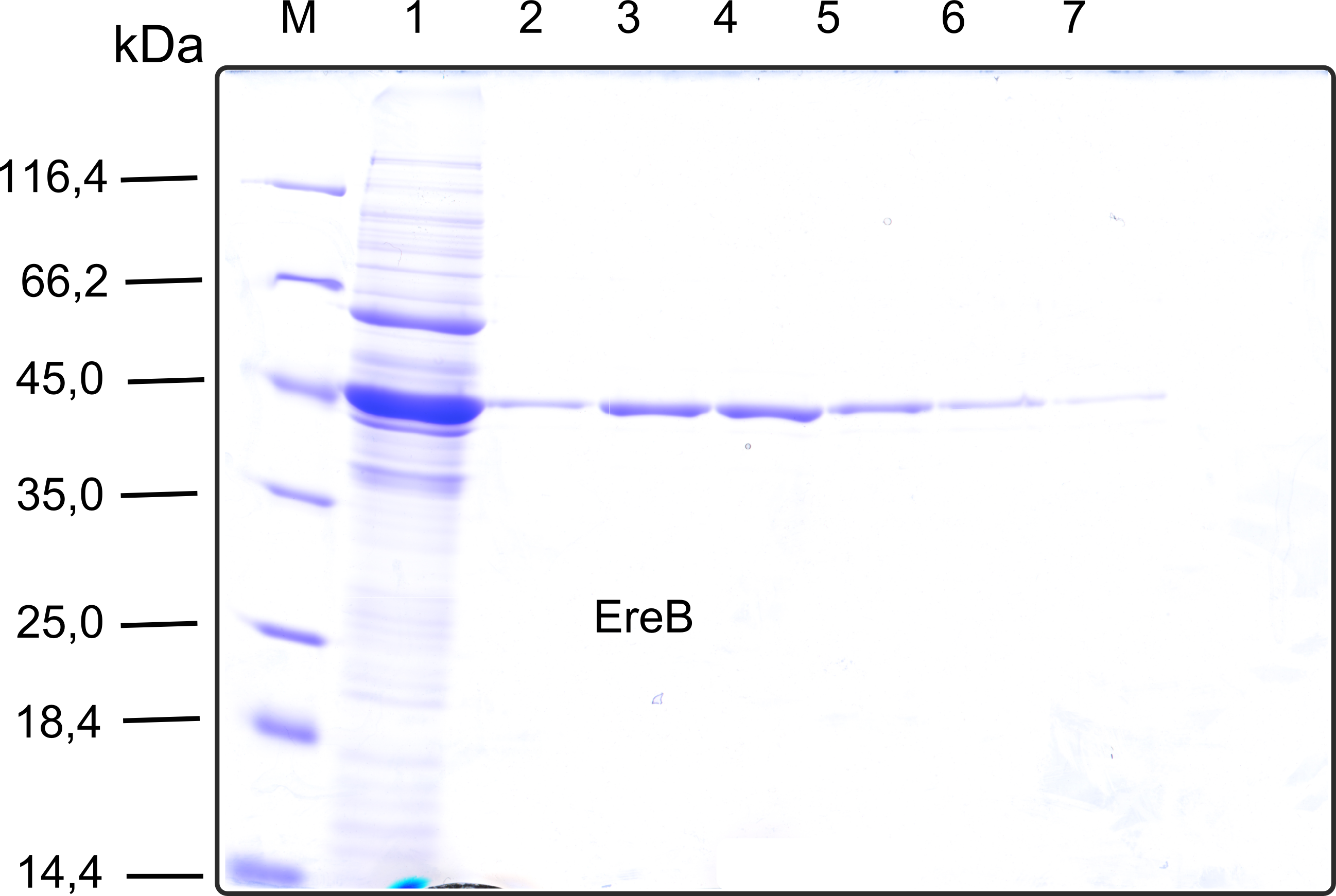
| + | [[File:TUM13_SDS_EreB.png|thumb|right|300px| '''Figure D''': SDS-gel of recombinant EreB with the marker (M) followed by the concentrated throughput of the streptavidin affinity column and 6 fractions collected from the elution peak]] | ||
| + | The recombinant production and purification was carried out twice, in a first attempt 2 L of LB-media were used for an analytical purpose whereas in the second attempt we produced enough purified enzyme for all subsequent experiments. This preparation was carried out in 6 x 2L of LB media. Protein production was in both cases induced at OD = 0.8 by adjusting the cell culture to 5 mM of arabinose and was carried out for 4 h for the first and 5 h for the second preparation. Cell disruption was performed by ultrasonic sound in both cases. The cell lysate was then dialyzed against 5 L of SA-buffer and subsequently applied to streptavidin affinity columns. After the application of the protein, the column was washed with SA-buffer until a base line was reached. Afterwards the protein was eluted using 5 mM biotin. During the first preparation 2-mercapto-ethanol was added after the chromatographic steps. In order to avoid oxidation of cysteine residues to disulphid-bridges, which is not desired for the cytosolic EreB protein, the preparative purification was carried out with buffers containing 5 mM of 2-mercapto-ethanol in all buffers. When comparing the size exclusion chromatograms, obtained from the analytical and the preparative purification, it can be stated that there is still a considerable aggregation peak near the void volume (Fig. B) of the column in the first attempt, which was nearly not the case for the preparative preparation (Fig. C). Therefore we would give the advise to use strictly reducing conditions while working with recombinant EreB. The finally resulting yields of the preparative purification have been determined by absorption measurement of the aromatic amino acids at 280. The total yield was determined to 25 mg of pure protein which is 2.1 mg/L of LB culture. | ||
<!-- --> | <!-- --> | ||
Revision as of 03:48, 5 October 2013
Erythromycin Esterase Type II (EreB) in RFC[25]
Erythromycin esterase type II from Escherichia coli that degrades macrolid antibiotics. Part is RFC[25] compatible and is flanked by RFC[25] pre- and suffix.
Usage and Biology
Protein Data Table for the erythromycin esterase (EreB) BBa_K1159000
| Protein data table for BioBrick BBa_BBa_K1159000 automatically created by the BioBrick-AutoAnnotator version 1.0 | ||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Nucleotide sequence in RFC 25, so ATGGCCGGC and ACCGGT were added (in italics) to the 5' and 3' ends: (underlined part encodes the protein) ATGGCCGGCAGGTTCGAA ... GTTTATGAAACCGGT ORF from nucleotide position -8 to 1260 (excluding stop-codon) | ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid sequence: (RFC 25 scars in shown in bold, other sequence features underlined; both given below)
| ||||||||||||||||||||||||||||||||||||||||||||||
Sequence features: (with their position in the amino acid sequence, see the list of supported features)
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid composition:
| ||||||||||||||||||||||||||||||||||||||||||||||
Amino acid counting
| Biochemical parameters
| |||||||||||||||||||||||||||||||||||||||||||||
| Plot for hydrophobicity, charge, predicted secondary structure, solvent accessability, transmembrane helices and disulfid bridges | ||||||||||||||||||||||||||||||||||||||||||||||
Codon usage
| ||||||||||||||||||||||||||||||||||||||||||||||
Alignments (obtained from PredictProtein.org)
| ||||||||||||||||||||||||||||||||||||||||||||||
| Predictions (obtained from PredictProtein.org) | ||||||||||||||||||||||||||||||||||||||||||||||
Subcellular Localization (reliability in brackets)
| Gene Ontology (reliability in brackets)
| |||||||||||||||||||||||||||||||||||||||||||||
Predicted features:
| ||||||||||||||||||||||||||||||||||||||||||||||
| The BioBrick-AutoAnnotator was created by TU-Munich 2013 iGEM team. For more information please see the documentation. If you have any questions, comments or suggestions, please leave us a comment. | ||||||||||||||||||||||||||||||||||||||||||||||
Production and purification of recombinant EreB
The recombinant production and purification was carried out twice, in a first attempt 2 L of LB-media were used for an analytical purpose whereas in the second attempt we produced enough purified enzyme for all subsequent experiments. This preparation was carried out in 6 x 2L of LB media. Protein production was in both cases induced at OD = 0.8 by adjusting the cell culture to 5 mM of arabinose and was carried out for 4 h for the first and 5 h for the second preparation. Cell disruption was performed by ultrasonic sound in both cases. The cell lysate was then dialyzed against 5 L of SA-buffer and subsequently applied to streptavidin affinity columns. After the application of the protein, the column was washed with SA-buffer until a base line was reached. Afterwards the protein was eluted using 5 mM biotin. During the first preparation 2-mercapto-ethanol was added after the chromatographic steps. In order to avoid oxidation of cysteine residues to disulphid-bridges, which is not desired for the cytosolic EreB protein, the preparative purification was carried out with buffers containing 5 mM of 2-mercapto-ethanol in all buffers. When comparing the size exclusion chromatograms, obtained from the analytical and the preparative purification, it can be stated that there is still a considerable aggregation peak near the void volume (Fig. B) of the column in the first attempt, which was nearly not the case for the preparative preparation (Fig. C). Therefore we would give the advise to use strictly reducing conditions while working with recombinant EreB. The finally resulting yields of the preparative purification have been determined by absorption measurement of the aromatic amino acids at 280. The total yield was determined to 25 mg of pure protein which is 2.1 mg/L of LB culture.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 189
Illegal XhoI site found at 599
Illegal XhoI site found at 1189 - 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal SapI.rc site found at 316