Part:BBa_K1761006
SmallBit Split Luciferase
SmallBit is the small part of a Split Luciferase. Its molecular weight is 1 kDa. This construct on its own has no function since SmallBit on itself has no luminescence activity. For luminescence activity, the LargeBit of the Split Luciferace is needed.
- Sequence will be published later.
Usage adn Biology
Split luciferase can be used as signaling components. This split luciferase consists of two parts, namely LargeBit and SmallBit. These parts were both inserted in a pETDuet-1 vector together with OmpX and a linker. LargeBit and SmallBit have an affinity towards each other, so when in close proximity they will come together and will give a luminescence signal. Followed is a short description of each part and of the NanoBit construct.
OmpX - LargeBit BBa_K1761005 [1]
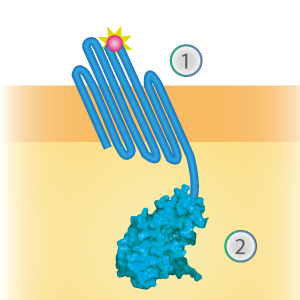
LargeBit is the big part of this Split Luciferase. Its molecular weight is 18 kDa. OmpX (1) (with a correct mutation for the amber stop codon TAG), a BamHI-linker and LargeBit (2) together will look like Figure 1. This construct on its own has no function since LargeBit on itself has no luminescence activity.
Figure 1: Schematical overview of the OmpX - LargeBit construct.
OmpX - SmallBit BBa_K1761006
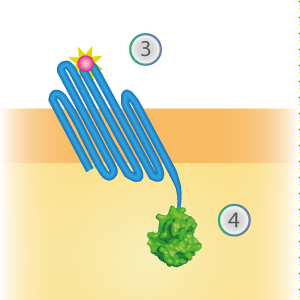
SmallBit is the small part of this Split Luciferase. Its molecular weight is 1 kDa, it is only 11 amino acids long. OmpX (1) (with a correct mutation for the amber stop codon TAG), a BsoBI-linker and SmallBit (2) together will look like Figure 2. This construct on its own has no function since SmallBit on itself has no luminescence activity.
Figure 2: Schematical overview of the OmpX - SmallBit construct.
NanoBit construct
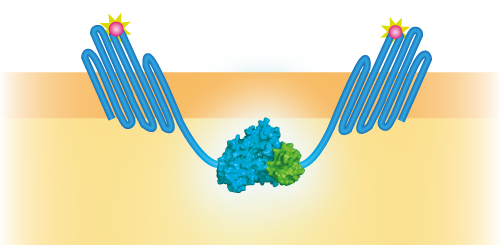
NanoBit utilizes a structural complementation-based approach to monitor protein interactions within living cells. Protein interaction promotes structural complementation and generation of a bright, luminescent enzyme. Protein dynamics can be followed in real-time in living cells following addition of the Nano-Glo Live Cell Reagent, a non-lytic detection reagent containing the cell-permeable furamizine substrate. See Figure 3 for the whole construct.
Figure 3: Schematical overview of the NanoBit construct.
Characterization
When mutated with the amber stop codon TAG, a unnatural amino acid with an azide-functionalized group can be expressed. After the expression of this amino acid, OmpX can covalently bind almost anything, as long as it contains a DBCO-functionalized group. The binding finds place by using a bio-orthogonal “click” reaction (SPAAC chemistry). To test the functionality of this “click” reaction, some experiments were done by clicking DBCO-PEG4-TAMRA at the surface. For all the experiments, the following vectors were used: pETDuet-1 with one or two construct(s) inserted (OmpX + intracellular protein) and pEVOL-pAzF (tRNA + tRNA synthetase). Both vectors were transformed into BL21(DE3). The expression was introduced by adding arabinose, IPTG and the unnatural amino acid.
DBCO-PEG4-TAMRA Confirmation
To confirm whether OmpX is in the membrane and whether or not the unnatural amino acid is being incorporated into OmpX, DBCO-PEG4-TAMRA was used. TAMRA is a fluorescent dye that can be used to verify the “click” reaction. If the unnatural amino acid is present, DBCO-PEG4-TAMRA should “click” to the transmembrane protein OmpX and stay there. This can be analyzed with FACS. For more information about how to perform FACS experiments, see our Protocol Page [http://2015.igem.org/Team:TU_Eindhoven/Project/Protocols].
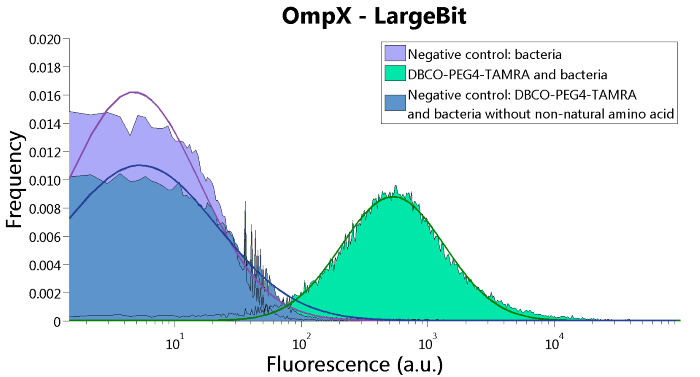
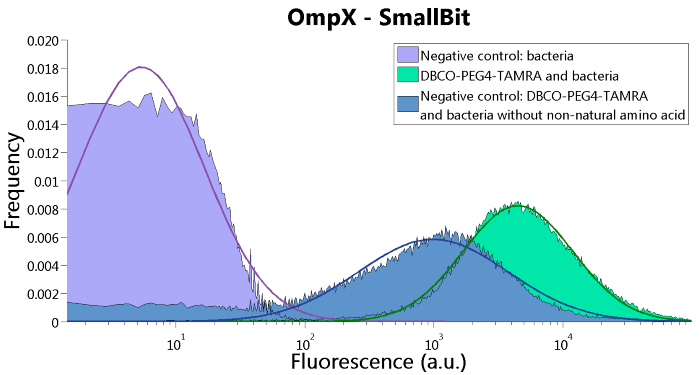
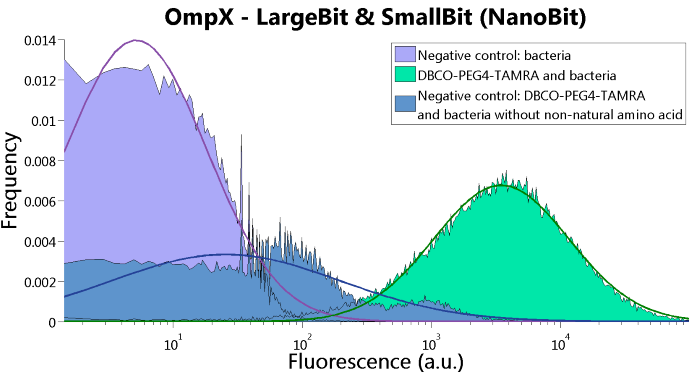
To verify that OmpX is in the membrane, we used the OmpX – LgBiT and OmpX – SmBiT constructs. These gave the following results after clicking with DBCO-PEG4-TAMRA (see Figure 4, 5 and 6). From this it can be concluded that OmpX is in the membrane and that the “click” reaction works.
Figure 4: FACS results of OmpX - LargeBit.
Figure 5: FACS results of OmpX - SmallBit
Figure 6: FACS results of OmpX - LargeBit and OmpX - SmallBit.
Bioluminescence Confirmation
To confirm whether NanoBit is present and is working, a bioluminescence measurement was performed. The results of this experiment are shown in Figure 7.
File:TU Eindhoven Bioluminescence NanoBit.png
Figure 7: Bioluminescence of results of the NanoBit construct. Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
//function/reporter/light
| emission | 460 nm |