Part:BBa_K3254000
phiC31attB-BsaI sites-terminator-phiC31attP(r)
This part is an improvement for the part BBa_K2243031. It can be placed between a promoter and a translational unit part and works as a normally open (NO) switch for the downstreamed gene, and switch to ON state by flipping the unidirectional terminator ECK120034435 between the att sites when it reacted with phiC31 integrase BBa_K1039012. In this improved version, two BsaI restriction site were added between attB site and terminator. As a result, it can work as a normally closed (NC) switch for the gene which was inserted between the two BsaI site and switch to OFF state when it flipped.
Usage and Biology
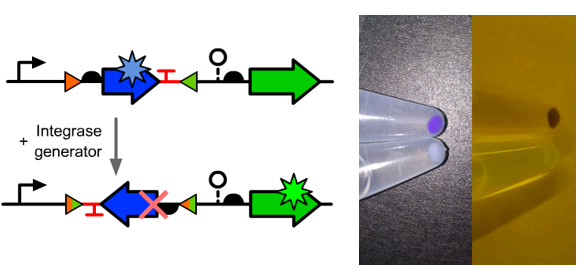
Visual Result as a Normally Closed Switch
- We conducted a simple test to see if our design met the expection.
Experimental Setup
- Genetic design principle of the experimental group is described on the page of BBa_K3254010.
- A P15A-AmpR plasmid was co-transfered into the E.coli DH5α host cell with the reporter plasmid containing this part as the negative control.
- Single colonies were selected from the experimental LB-agar plate and negative control LB-agar plate, then inoculated into EP tubes with 500 μL M9 supplemented medium containing 500 μM IPTG for overnight growth at 37 °C and 200 rpm.
- Tubes were centrifuged at 10000g for 1 min. Then observed the GFP fluorescence of the cell precipitations under blue light.
Results
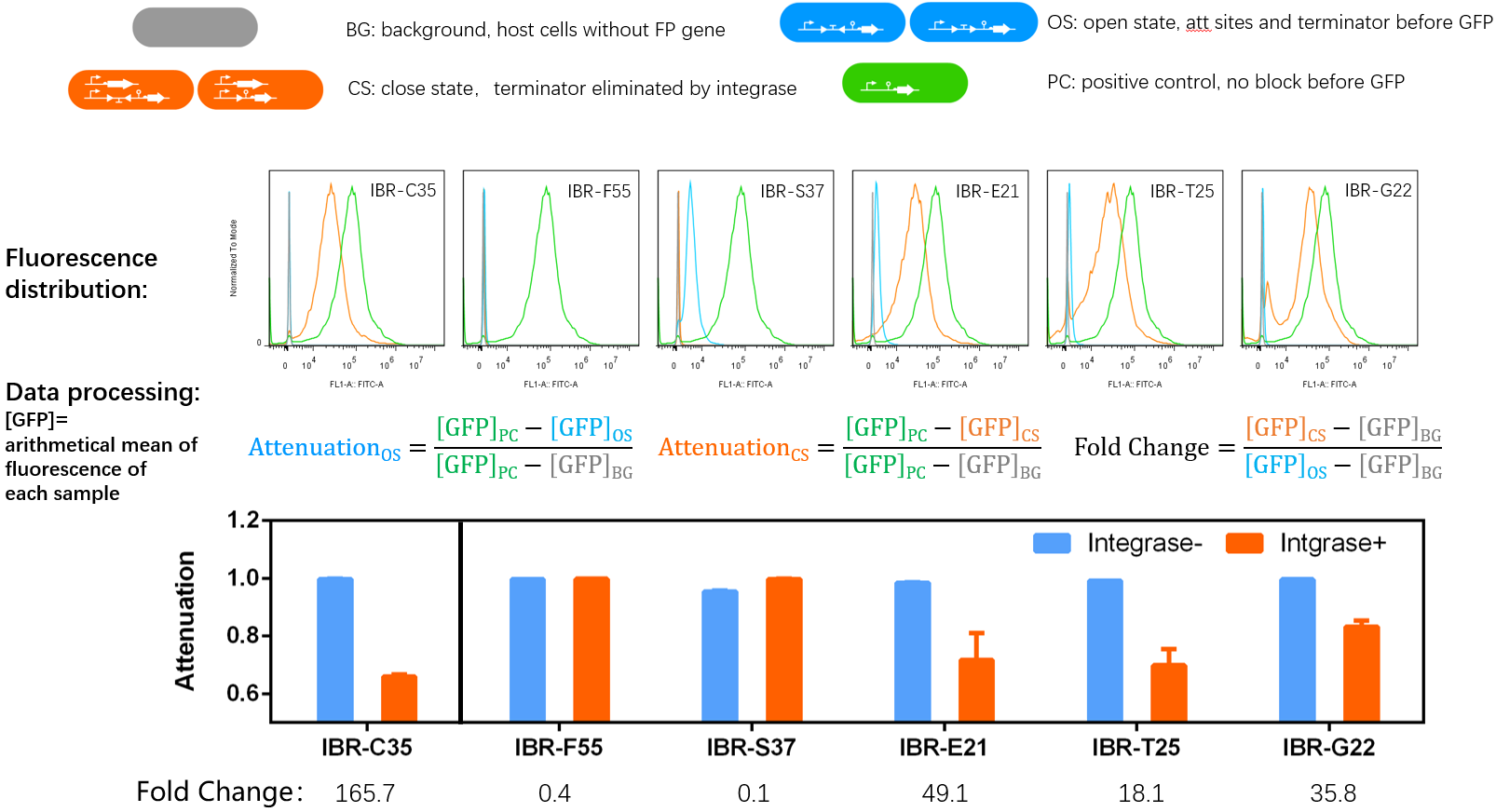
- IBR-C35/F55/S37/E21/T25/G22 indicate the experimental systems for phiC31/Int5/Int7/Int8/Int10/TG1 respctively.
- We observed the GFP fluorescence from the experimental tube as expected.
Quantitative Characterizaion of the Normally Open Switch
Experimental Setup
- Bacteria harboring the circuits (see the top part of the result image) first inoculated from single colonies into a flat-bottom 96-well plate for overnight growth. Then, the cell cultures were diluted 1000-fold with M9 supplemented medium with 500 μM IPTG inducer and growth for another 20 hours. All incubations were carried out using a Digital Thermostatic Shaker maintained at 37 °C and 1000 rpm, using flat-bottom 96-well plates sealed with sealing film. Finally, 3-μL samples each culture were transferred to a new 96-well plate containing 200 μL per well of PBS supplemented with 2 mg/mL kanamycin.
- The fluorescence distribution of each sample was assayed using a flow cytometry. The arithmetical mean of each sample was determined using FlowJo software.
- The principle of data processing is shown on the result image.
Results
- IBR-C35/F55/S37/E21/T25/G22 indicate the experimental systems for phiC31/Int5/Int7/Int8/Int10/TG1 respctively.
- Compared to other parts, this part performed well.
Visual Results as a Normally Closed Switch and Toggle Switch
- We inserted an amilCP translational unit between the two BsaI sites.
- Other experiment setup were the same with "Visual Result as a Normally Closed Switch".
Results
- The normally open switch function well though a light blue color can be observed from the cell precipitations which might due to the incomplete diluted amilCP protein or an unexpected backward promoter.
- At the same time, the downstreamed GFP wasn't expression well which might due to the potential attenuation signal in the reversed amilCP sequence.
Orthogonality Characterization
Genetic Design
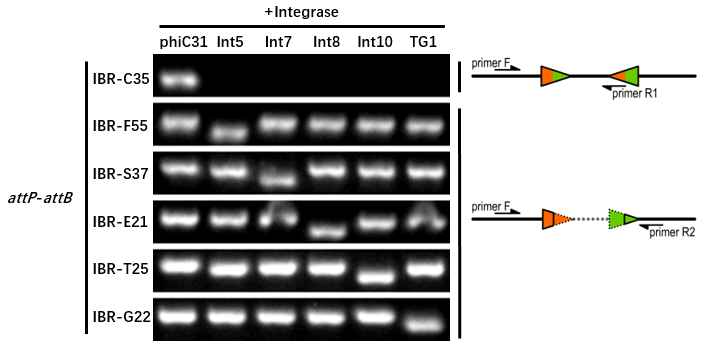
- The composition and principle of the experimental system are indicated below.
Experimental Setup
The reporter plasmid contained this part were co-transferred into E.coli DH5α host with 6 integrase generator plasmids. Then single colonies were inoculated into M9 supplemented medium for overnight growth. Then, the cell cultures were diluted 1000-fold with M9 supplemented medium with 500 μM IPTG inducer and growth for another 20 hours. All incubations were carried out using a Digital Thermostatic Shaker maintained at 37 °C and 1000 rpm, using flat-bottom 96-well plates sealed with sealing film. Finally, The cultures were sampled for genotype PCR testing. The principle of genotype identification was shown on the right of results image.
Results
- IBR-C35 was the plasmid containing this part.
- The result indicates that this part can only be recombined by phiC31 integrase.
- The sequences after recombination are GATCAGCTCCGCGGGCAAGACCGTGCTCTTACCCAGTTGGGCGGGA (attL) and tgcgGGTGCCAGGGCGTGCCCTTGAGTTCTCTCAGTTGGGGG (attR).
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 59
Illegal BsaI.rc site found at 47
| None |