Difference between revisions of "Part:BBa K346003:Experience"
JjunyiJiao (Talk | contribs) |
|||
| Line 4: | Line 4: | ||
| − | To be specific, the entire coding region of the MBP for standard part was amplified by PCR from full length MerR with two pairs of primers. Two of these primers encoded a three-residue bridge, SSG, which does not occur in MerR and was added to | + | To be specific, the entire coding region of the MBP for standard part was amplified by PCR from full length MerR with two pairs of primers. Two of these primers encoded a three-residue bridge, SSG, which does not occur in MerR and was added to facilitate some flexibility in the loop connecting the two dimerization helix. The two PCR products were digested with EcoR I / BamH I, or BamH I / Pst I and cloned into EcoR I / Pst I -digested pSB1K3 in one step (fig), which was verified by sequencing. Then the RBS is prefixed to MBP. |
| − | Based on the same strategy, MBP-His6 was constructed by using two different pairs of primers, which is used for MBP expression test by western blot. The two PCR products were digested with Nde I / BamH I, or BamH I / Xho I and cloned into Nde I / Xho I -digested pET 21a, which contains a region encoding six histines, in one step to construct pET 21a – mbp , which was verified by | + | Based on the same strategy, MBP-His6 was constructed by using two different pairs of primers, which is used for MBP expression test by western blot. The two PCR products were digested with Nde I / BamH I, or BamH I / Xho I and cloned into Nde I / Xho I -digested pET 21a, which contains a region encoding six histines, in one step to construct pET 21a – mbp , which was verified by sequencing. |
Revision as of 04:37, 28 October 2010
Contents
construction of BBa_K346003
To be specific, the entire coding region of the MBP for standard part was amplified by PCR from full length MerR with two pairs of primers. Two of these primers encoded a three-residue bridge, SSG, which does not occur in MerR and was added to facilitate some flexibility in the loop connecting the two dimerization helix. The two PCR products were digested with EcoR I / BamH I, or BamH I / Pst I and cloned into EcoR I / Pst I -digested pSB1K3 in one step (fig), which was verified by sequencing. Then the RBS is prefixed to MBP.
Based on the same strategy, MBP-His6 was constructed by using two different pairs of primers, which is used for MBP expression test by western blot. The two PCR products were digested with Nde I / BamH I, or BamH I / Xho I and cloned into Nde I / Xho I -digested pET 21a, which contains a region encoding six histines, in one step to construct pET 21a – mbp , which was verified by sequencing.
Expression Experiment and Function Test:
To test the function of this part, both expression experiment and function test is necessary. We have verified the size of the expressed proteins with SDS-page and Western blot. Besides, to test the efficiency of mercury binding, we also carried out the function test with ICP-AES with the mercury gradient from 10^-8M to 10^-6M.
Results:
Expression of proteins
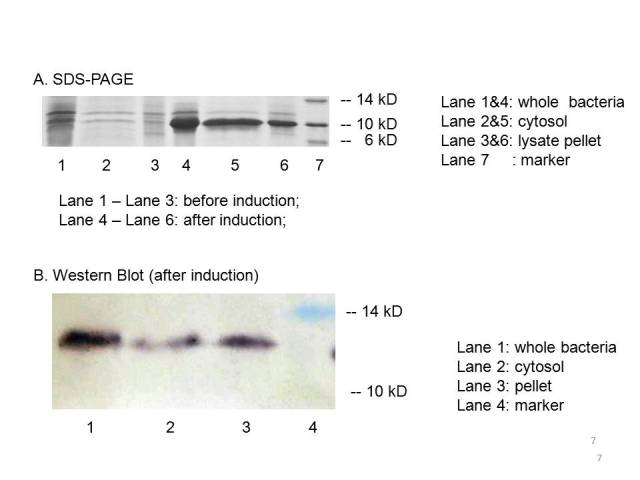
The plasmid PET21a-MBP is transferred to E.coli strain BL21, which can generate T7 polymerase when induced with IPTG. Both induced cells and uninduced cells(as control) are centrifuged to get the cytosol, the periplasm and the membrane separated. The SDS-page and Western blot of the expressed proteins show that induced cells expressed an identical IPTG-inducible protein at the proper place with the size of ~10kD which consists with the predicted size, indicating that the engineered MBP can be expressed in the cytosol.
We can detected overexpression band at about 10 kD, which is similar to MBP. Then we fused a his-tag to our target protein and conducted western blot to further verify their expression. Positive band of the expected molecular weight can be detected in the cytosol, which confirm expression and localization of the target protein. We can also indicate from the SDS-PAGE that there a large number of MBPs exist in cytosol ready to bind mercury ions.
Function test
Having made sure that the protein can express normally in the cytosol, the function tests experiment are carried out with ICP-AES. To test the efficiency of mercury absorption of MBP in different concentration of mercury, the concentration gradient is set from 10^-8M to 10^-6M, the results are shown in figure 3. Our experiment shows that the efficiency of MBP increases with the increase of the mercury concentration and MBP can binding Hg(II) with high efficiency and high sensitivity from the concentration of 10^-7 M(not show in the figure) compared to that of the control.
Figure 3: Overnight cultures were diluted 1:100 and grown to final OD600=0.6-1.0. We then resuspend the culture with 1% PBS and make samples for ICP-AES according to the protocol(See more about this, please visit our wiki.)
User Reviews
UNIQ47dcbbc80eb93baa-partinfo-00000000-QINU UNIQ47dcbbc80eb93baa-partinfo-00000001-QINU