Difference between revisions of "Part:BBa K3453161"
| Line 6: | Line 6: | ||
===Usage and Biology=== | ===Usage and Biology=== | ||
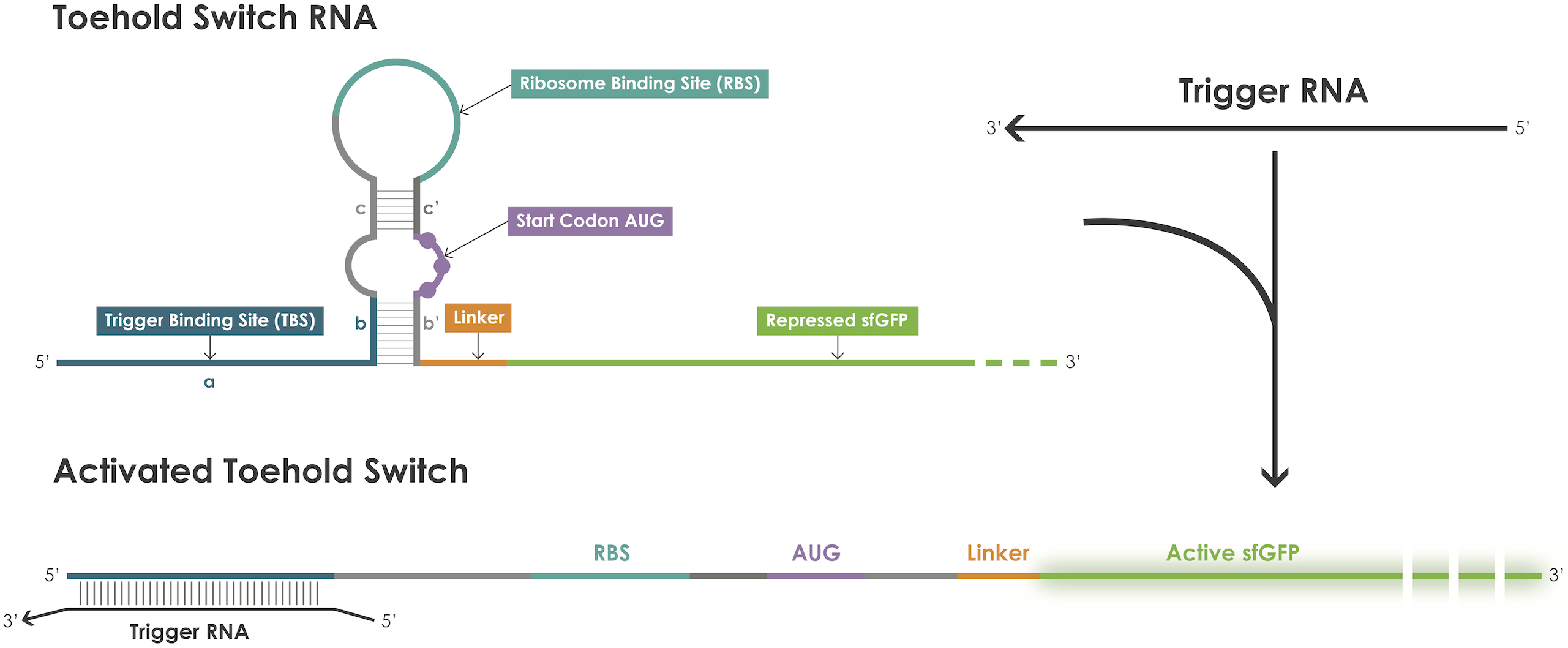
| − | A toehold switch is an RNA–based device containing a ribosome binding site (RBS) and an ATG start codon embedded in a hairpin structure that blocks translation initiation [1]. The hairpin can be unfolded upon binding of a trigger RNA | + | A toehold switch is an RNA–based device containing a ribosome binding site (RBS) and an ATG start codon embedded in a hairpin structure that blocks translation initiation [1]. The hairpin can be unfolded upon binding of a trigger RNA therebyexposing the RBS and the ATG start codon permitting translation of the reporter protein (Figure 1). |
[[File: T--Evry_Paris-Saclay--toehold-switch.png|600px]] | [[File: T--Evry_Paris-Saclay--toehold-switch.png|600px]] | ||
| Line 14: | Line 14: | ||
This part is a transcription unit of [[Part:BBa_K3453061|BBa_K3453061]] which is a trigger for the Rosewood DmTrnL-UAA Toehold Switch 1.1 ([[Part:BBa_K3453051|BBa_K3453051]]) and also contains the switch binding site for the Rosewood DmTrnL-UAA Toehold Switch 1.2 ([[Part:BBa_K3453052|BBa_K3453052]]) and 2.3 ([[Part:BBa_K3453056|BBa_K3453056]]). | This part is a transcription unit of [[Part:BBa_K3453061|BBa_K3453061]] which is a trigger for the Rosewood DmTrnL-UAA Toehold Switch 1.1 ([[Part:BBa_K3453051|BBa_K3453051]]) and also contains the switch binding site for the Rosewood DmTrnL-UAA Toehold Switch 1.2 ([[Part:BBa_K3453052|BBa_K3453052]]) and 2.3 ([[Part:BBa_K3453056|BBa_K3453056]]). | ||
| − | This part proved to be functional: the transcribed rosewood RNA was able to bind to the | + | This part proved to be functional: the transcribed rosewood RNA was able to bind to the Rosewood DmTrnL-UAA Toehold Switch 2.3 and a readable output was generated. However, the Rosewood DmTrnL-UAA Toehold Switches 1.1 and 1.2 turned out to be not functional and the interaction with these switches could not be assessed. |
Full results are available on the [[Part:BBa_K3453151|BBa_K3453151]], [[Part:BBa_K3453152|BBa_K3453152]] and [[Part:BBa_K3453156|BBa_K3453156]] pages in the registry. | Full results are available on the [[Part:BBa_K3453151|BBa_K3453151]], [[Part:BBa_K3453152|BBa_K3453152]] and [[Part:BBa_K3453156|BBa_K3453156]] pages in the registry. | ||
Latest revision as of 21:52, 26 October 2020
Rosewood DmTrnL-UAA Trigger 1.1 transcription unit
This part is a transcription unit of the Rosewood DmTrnL-UAA Trigger 1.1 (BBa_K3453061) for sequence-based detection of rosewood. The transcription is controlled by the T7 promoter (BBa_K2150031) and the strong SBa_000587 synthetic terminator (BBa_K3453000).
Usage and Biology
A toehold switch is an RNA–based device containing a ribosome binding site (RBS) and an ATG start codon embedded in a hairpin structure that blocks translation initiation [1]. The hairpin can be unfolded upon binding of a trigger RNA therebyexposing the RBS and the ATG start codon permitting translation of the reporter protein (Figure 1).
Figure 1. Toehold switches principle.
This part is a transcription unit of BBa_K3453061 which is a trigger for the Rosewood DmTrnL-UAA Toehold Switch 1.1 (BBa_K3453051) and also contains the switch binding site for the Rosewood DmTrnL-UAA Toehold Switch 1.2 (BBa_K3453052) and 2.3 (BBa_K3453056).
This part proved to be functional: the transcribed rosewood RNA was able to bind to the Rosewood DmTrnL-UAA Toehold Switch 2.3 and a readable output was generated. However, the Rosewood DmTrnL-UAA Toehold Switches 1.1 and 1.2 turned out to be not functional and the interaction with these switches could not be assessed.
Full results are available on the BBa_K3453151, BBa_K3453152 and BBa_K3453156 pages in the registry.
References
[1] Green AA, Silver PA, Collins JJ, Yin P. Toehold switches: de-novo-designed regulators of gene expression. Cell (2014) 159, 925-939.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]