Difference between revisions of "Part:BBa K3349004"
Kristiturton (Talk | contribs) |
Kristiturton (Talk | contribs) |
||
| Line 9: | Line 9: | ||
A GFP reporter was added to the C-terminus of the gene. As well, a C-terminal his tag was added to the GFP for nickel affinity purification. | A GFP reporter was added to the C-terminus of the gene. As well, a C-terminal his tag was added to the GFP for nickel affinity purification. | ||
| + | |||
| + | |||
| + | |||
| + | For wetlab results we were able to start preparing for subcloning our parts into pSB1C3. The following Figure shows our digestion of the genes from pUC57-Kan using EcoRI and PstI. | ||
| + | |||
| + | <html> | ||
| + | <img src= "https://2020.igem.org/wiki/images/c/c9/T--Lethbridge_HS--insertdigest2020.png" alt="pnl modelling" style="width:500px;height:700px;"> | ||
| + | </html> | ||
| + | |||
| + | |||
| + | <b>Figure 3: </b> Restriction digest of PelB and PelC DNA from the pUC57 plasmid using EcoRI and PstI enzymes run on a 1% agarose gel. The red boxes show the band corresponding to that of our DNA constructs. | ||
| + | |||
<!-- Add more about the biology of this part here | <!-- Add more about the biology of this part here | ||
Revision as of 16:46, 24 October 2020
Pectin Lyase PelB-GFP
This Pectin Lyase is a protein from Paenibacillus amylolyticus. With Pnl and PelC, PelB is able to cleave and therefore breakdown pectin or homogalacturonan substrates. This protein has been characterized to cleave methylated pectin (at unmethylated sites) and polygalacturonic acid. Studies by Keggi and Doran-Peterson determined that PelB has a Km of 50.1+/- 140 mg/ml and a Vmax of 745 +/-2024 IU/mg with a 80% methylated substrate. The optimal temperature for PelB activity was determined to be 70°C [1].
For more information, please visit our website https://2020.igem.org/Team:Lethbridge_HS or the basic gene part BBa_K3349001.
A GFP reporter was added to the C-terminus of the gene. As well, a C-terminal his tag was added to the GFP for nickel affinity purification.
For wetlab results we were able to start preparing for subcloning our parts into pSB1C3. The following Figure shows our digestion of the genes from pUC57-Kan using EcoRI and PstI.
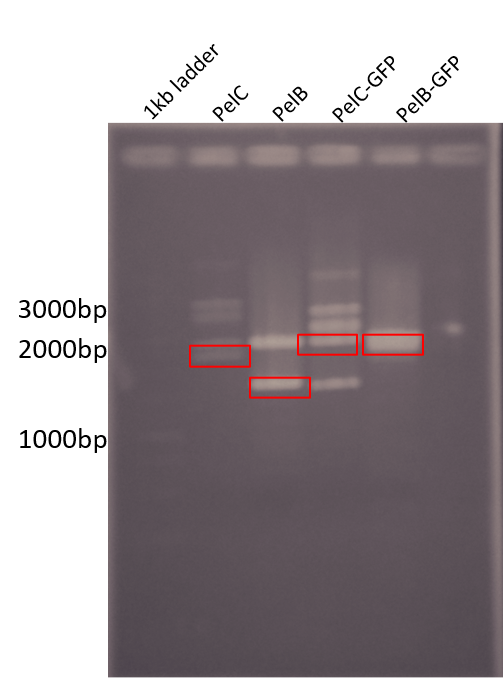
Figure 3: Restriction digest of PelB and PelC DNA from the pUC57 plasmid using EcoRI and PstI enzymes run on a 1% agarose gel. The red boxes show the band corresponding to that of our DNA constructs.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 261
Illegal BamHI site found at 1146 - 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
