Difference between revisions of "Part:BBa J64997"
| Line 26: | Line 26: | ||
====Improvements==== | ====Improvements==== | ||
| − | This part was used as a promoter in the following parts : BBa_K2203001, BBa_K2203002, BBa_K2203003, BBa_K2203004, BBa_K2203005, BBa_K2203006 ; BBa_K3100022 (by SCUT_China | + | This part was used as a promoter in the following parts : BBa_K2203001, BBa_K2203002, BBa_K2203003, BBa_K2203004, BBa_K2203005, BBa_K2203006 ; BBa_K3100022 (by SCUT_China 2019https://parts.igem.org/Part:BBa_K3100022). |
<h4><strong>Improvement of BBa_K3100022 by SCUT_China 2019</strong></h4><br> | <h4><strong>Improvement of BBa_K3100022 by SCUT_China 2019</strong></h4><br> | ||
Revision as of 03:53, 22 October 2019
T7 consensus -10 and rest
Characterization in vivo
This part has been characterized for GFP expression in the pCDF plasmid by the 2015 LASATX iGEM team.
We placed the T7 promoter in front of the sfGFP gene (BBa_K1624004). The experimental construct was placed in a pCDFDuet-1 vector and transformed into BL21(DE3) cells. We used BL21(DE3) cells without the plasmid containing T7 as our negative control. Cells were grown overnight. A 1:100 dilution was made, and cultures were grown with 20% glucose to 0.6 OD. At 0.6 OD, cells were resuspended in IPTG and induced for 2 hours. After induction, cells were washed and resuspended in PBS buffer. Fluorescence was then measured at 600 nm for 25 flashes. Bars show average of triplicates. Error bars show standard deviation.

The sfGFP gene placed downstream of the T7 promoter was successfully expressed, with fluorescence from the experimental construct significantly higher than the control.
Characterization in Cell-Free Environment
The T7 promoter was characterized in self-made E.Coli lysate from strain BL21(DE). Cell-free T7-GFP-mut3b versus GFP-mut3b (BBa_E0040) synthesis was analyzed. Fluorescence was measured at 37°C for five hours on a plate reader. For details on how the lysate and the energy solution were made and which components went into the final reaction volume of 10uL, check out our [http://2017.igem.org/Team:EPFL/Protocols protocols]. Shown are two times three repeats and a negative control, a shaded error graph (control was subtracted) and a bar plot summarizing the result.
GFP-mut3b expression cannot be dinstinguished from the no DNA control. T7-GFP-mut3b expression however is high. Saturation occurs after about five hours. Adding the T7 promoter in front of the GFP-mut3b enables transcription by the T7 polymerase, a much more efficient polymerase than the E.Coli polymerase found natively in E.Coli cells.



Improvements
This part was used as a promoter in the following parts : BBa_K2203001, BBa_K2203002, BBa_K2203003, BBa_K2203004, BBa_K2203005, BBa_K2203006 ; BBa_K3100022 (by SCUT_China 2019https://parts.igem.org/Part:BBa_K3100022).
Improvement of BBa_K3100022 by SCUT_China 2019
In order to achieve a larger scale and more accurate regulation range, we have improved T7 promoters with different strength providing more options for the precise regulation.
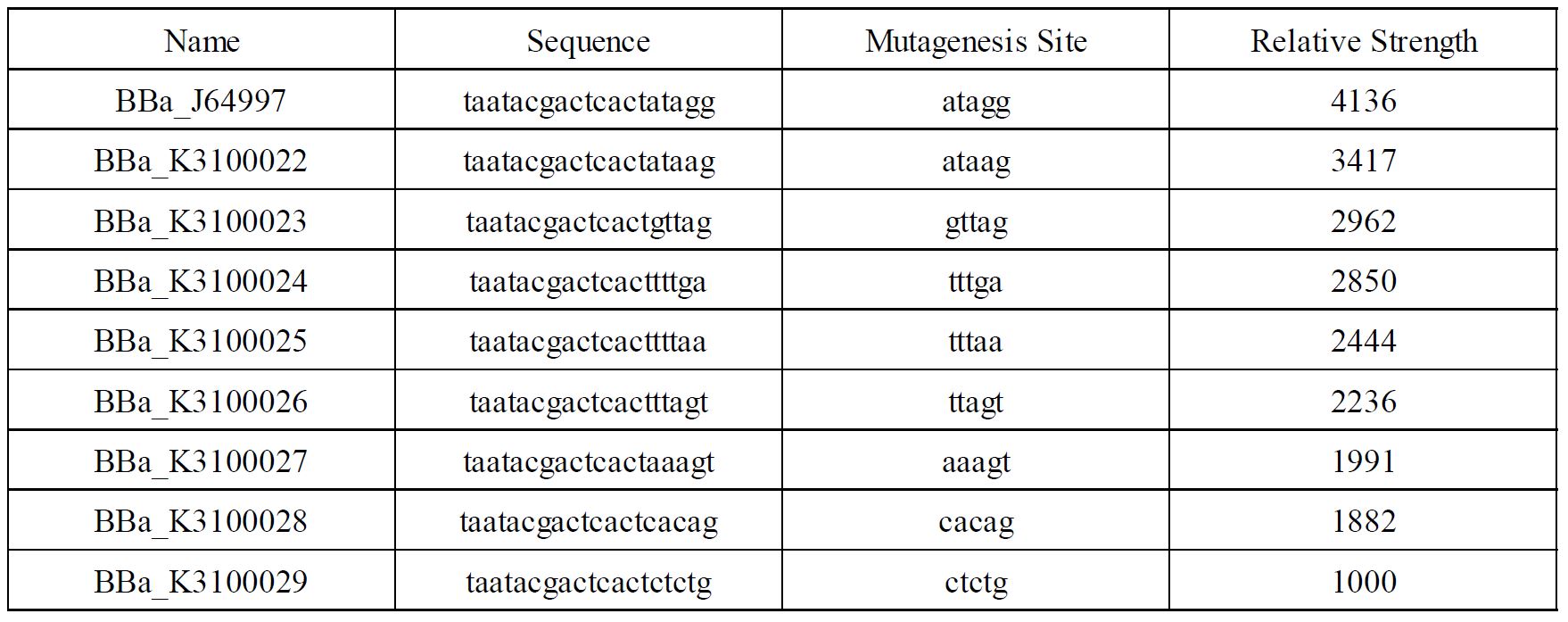
Fig1. T7 promoters with different strength
Characterization:
1. Construction of the T7 Promoter Library
A T7 promoter library was constructed through site-saturation mutagenesis, and an efficient screening method based on GFP fluorescence detection was established.
The site-saturation mutagenesis DNA fragment pET30a(+)-PT7-RFP vector 1 was obtained using the plasmid pET30a(+)-PT7-RFP as the template and vector 1.F (the sequence containing the mutation in promoter)and vector 1.R as the primers. At the same time, vector 2.F and vector 2.R as the primers to obtain pET30a(+)-PT7-RFP vector 1 by PCR. And then construct pET30a(+)-PT7 mutagenesis -RFP with pET30a(+)-PT7-RFP vector 1 and vector 2 by Gibson Assembly.
Fig 2. Site-saturation mutagenesis was conducted to obtain the T7 mutants
vector 1.F |
CCCGCGAAATTAATACGACTCACTNNNNNGGAATTGTGAGCGGATAAC |
vector 1.R |
TACCGCACAGATGCGTAAG |
vector 2.F |
TTCTCCTTACGCATCTGT |
vector 2.R |
AGTGAGTCGTATTAATTTCG |
Table 1: Primers for site-saturation mutagenesis
2. T7 Mutant Screening and Determination of GFP Fluorescence
T7 mutants were preliminarily selected by fluorescence of colonies grown on LB agar plates with Lactose. These mutants were inoculated in 2ml LB medium and cultured at 37 °C with agitation at 250 rpm for 12 h. Then, 1:100 inoculate to 200μl LB medium (with 0.2mM IPTG) in 96 - well plates at 30°C with agitation at 250 rpm for 16 h. The excitation wavelength was set to 553 nm, and the emission wavelength was set to 583 nm. The optical density of bacteria at 600 nm was detected with a spectrophotometer.
Finally, we have screened 8 different strength T7 mutants. The following figure is fluorescence/OD600 of them.
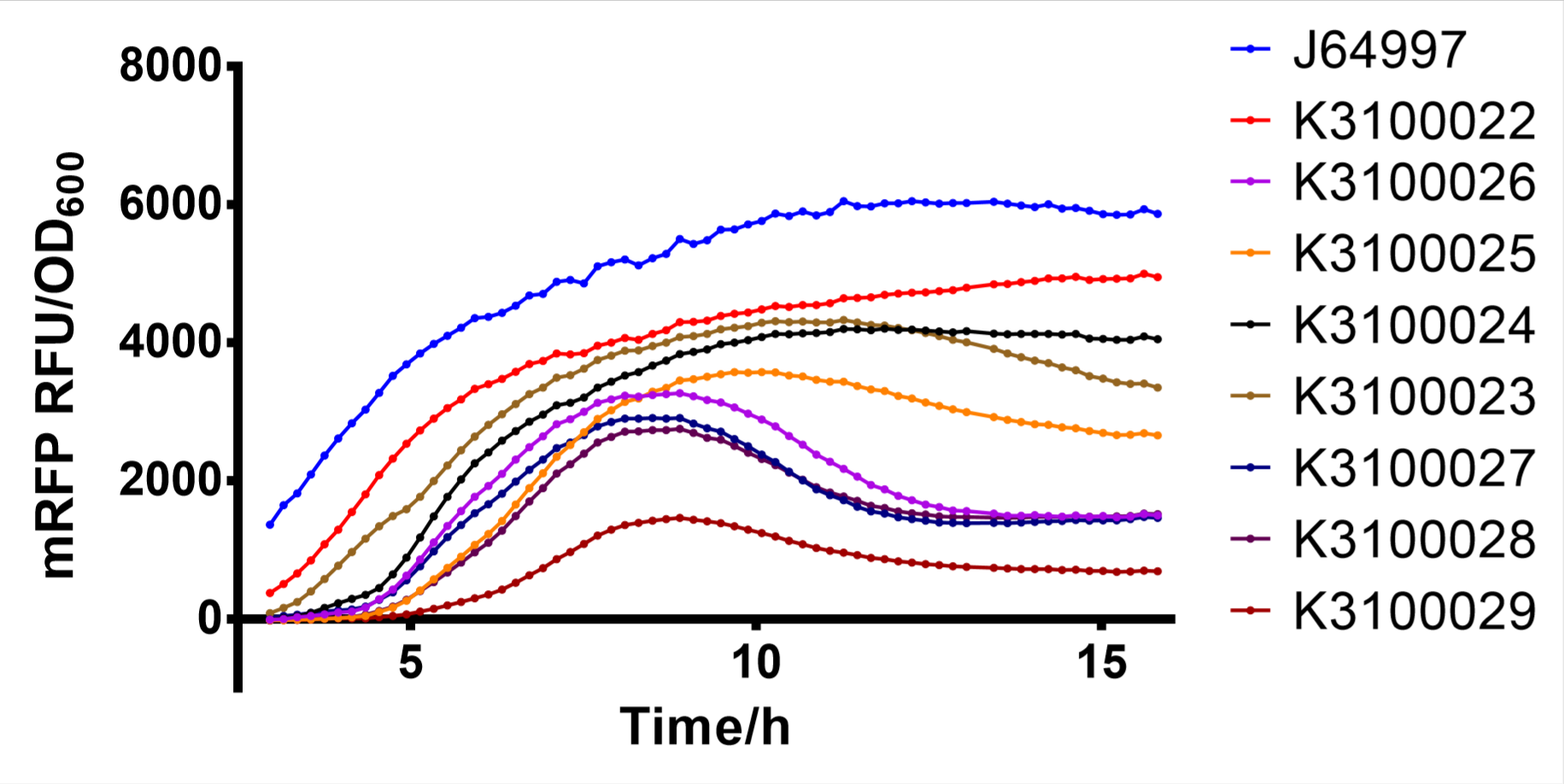
Fig. 2: fluorescence/OD600 of T7 mutants
Reference:
[1] Nie, Z., Luo, H., Li, J. et al. Appl Biochem Biotechnol (2019). https://doi.org/10.1007/s12010-019-03113-y
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]

