Difference between revisions of "Part:BBa K3187014"
PeterGockel (Talk | contribs) |
|||
| Line 112: | Line 112: | ||
<hr class="head"> | <hr class="head"> | ||
<p> | <p> | ||
| − | Both the original and the improved version were cloned in pSB1C3. <i>E. coli</i> BL21 (DE3) was transformed using BBa_K1073024 and cultured using 25 mL LB medium supplemented with | + | Both the original and the improved version were cloned in pSB1C3. <i>E. coli</i> BL21 (DE3) was transformed using BBa_K1073024 and cultured using 25 mL LB medium supplemented with chloramphenicol (based on the pSB1C3 resistance). The expression was measured (Ex: 587 nm, Em: 610 nm) over a period of 6 hours after induction with 0.5 mM IPTG (<b>Fig. 2</b>) using a SpectraMax M5E. |
</p> | </p> | ||
<img | <img | ||
Latest revision as of 17:46, 21 October 2019
Plac-RBS-mCherry-double terminator (IPTG-inducible) with Translation Enhancing 5'UTR
Improved Version of BBa_K1141000
Usage and Biology
BBa_K3187014 (improved BBa_K1141000) encodes the fluorescent protein mCherry under the control of an isopropyl-β-D-thiogalactopyranosid (IPTG)-inducible T7 promoter. The parts was improved in terms of its expression levels, based on the insertion of a 5’-untranslated region (5’-UTR) upstream of the coding sequence. This 5'-UTR was adapted from iGEM Bielefeld 2015 ( BBa_K1758100) and is based on the research of Olins et al [1] and Takahashi et al. [2] . It contains the strong ribosomal binding site (RBS) g10-L from the T7 bacteriophage and a sequence that plays a role in the regulation of mRNA binding to and release from the 30S ribosomal subunit. The 5'-UTR therefore enhances the translation efficiency of the following coding sequence (CDS) (Fig. 1).
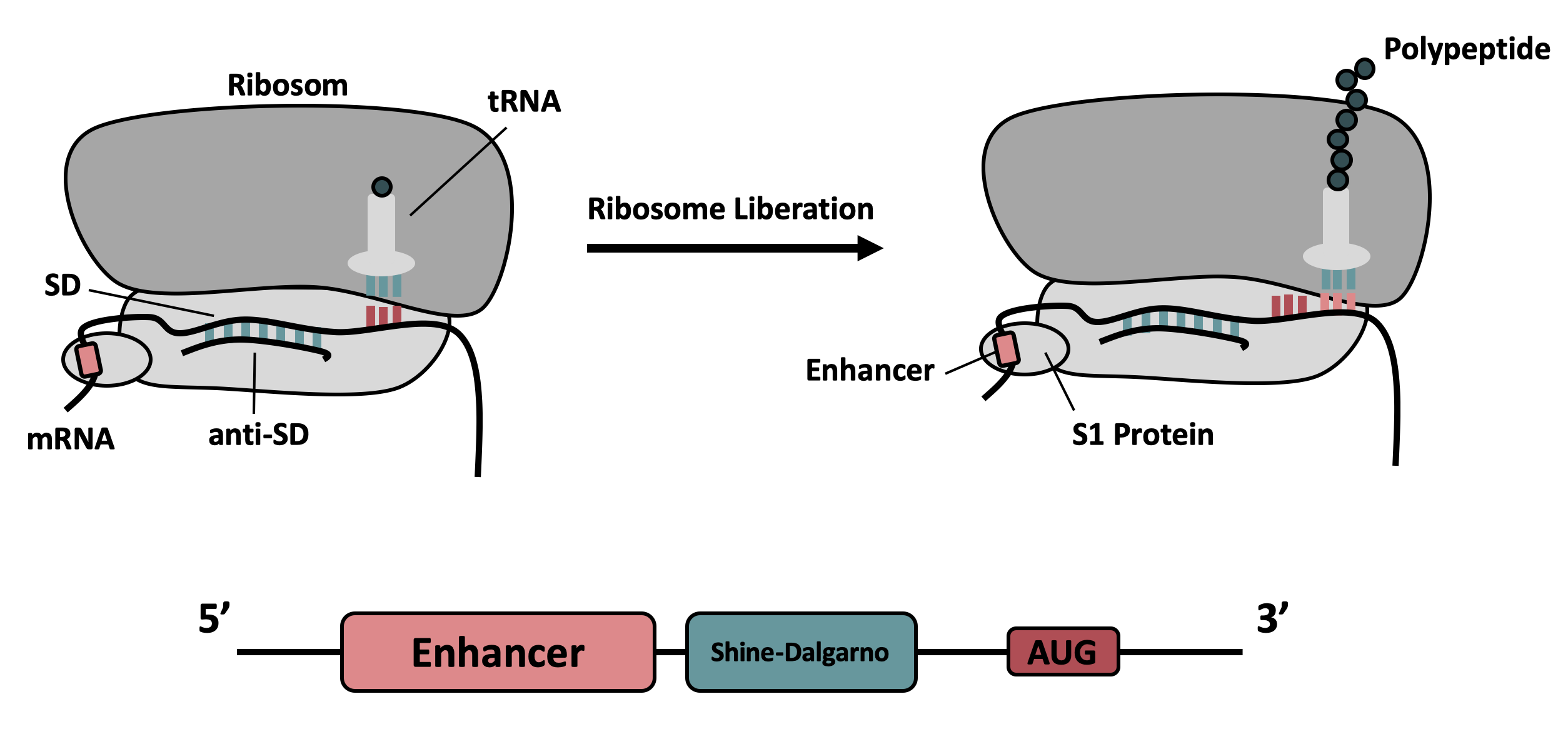
The sequence of the translation enhancing 5’-UTR can be divided into the four main features listed below:

| Sequence | Function |
|---|---|
|
AATAATTTTGTT TTAACTTTAA |
The T7 g10 leader sequence (first described by Olins et al [1] )increases the efficiency of translation initiation. This sequence contains the epsilon motif TTAACTTTA which enhances the binding of the mRNA to the 16S rRNA. |
| poly-A | Referring to Takahashi et al. [2] a spacer between the epsilon motive and the RBS improves the translation rate. |
| GAAGGAG | According to Karig et al. [3] and Lentini et. al [4] a distance of 4-9 bases between RBS and start codon increases the translation efficiency. |
| AATAATCT | According to Lentini et. al [4] an AT-rich composition between the RBS and the start codon results in the best expression results. |
Results
Both the original and the improved version were cloned in pSB1C3. E. coli BL21 (DE3) was transformed using BBa_K1073024 and cultured using 25 mL LB medium supplemented with chloramphenicol (based on the pSB1C3 resistance). The expression was measured (Ex: 587 nm, Em: 610 nm) over a period of 6 hours after induction with 0.5 mM IPTG (Fig. 2) using a SpectraMax M5E.

Figure 2 shows an increased expression of the improved BBa_K1141001 in comparison to BBa_K1141001. The increased expression rate of the improved version (left) after 6 hours can also be observed with the naked eye (Fig. 3).

References
- ↑ Peter O. Olins and Shaukat H. Rangwala, A Novel Sequence Element Derived from Bacteriophage T7 mRNA Acts as an Enhancer of Translation of the lacZ Gene in Escherichia coli, The Journal of Biological Chemistry, Vol. 264, No. 29, Issue of Ovtobet 15, pp. 16973-16976, 1989 [1]
- ↑ Shuntaor Takahashi, Hiroyuki Furusawa, Takuya Ueda and Yoshio Okahata, Translation Enhancer Improves the Ribosome Liberation from Translation Initiation, J. Am. Chem. Soc. 2013, 135 35, 13096-13106 [2]
- ↑ David K. Karig, Sukanya Iyer, Michael L. Simpson and Mitchel J. Doktycz, Expression optimization and synthetic gene networks in cell-free systems, Nucleic Acids Res. 2012 Apr; 40(8): 3763.3774 [3]
- ↑ Roberta Lentini, Silvia Perez Santero, Fabio Chizzolini, Dario Cecchi, Jason Fontana, Marta Marchioretto, Christina Del Bianco, Jessica L. Terrell, Amy C. Spencer, Laura Martini, Michele Forlin, Michael Assfalg, Mauro Dalla Serra, William E. Bentley and Sheref S. Mansy, Integrating artificial with natural cells to translate chemical messages that direct E. coli behavior, Nature Communications 5, Article number: 4012 (2014) [4]
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
