Difference between revisions of "Part:BBa K3147003"
| Line 5: | Line 5: | ||
| − | ===I : parts BBa_K3147003 | + | ===I : parts BBa_K3147003 function=== |
This construction produces an mRFP1 (BBA_E1010)[1] fused in C-ter with a ssrA fast degradation tag [2] (BBA_M0050). The TEV cutting site (BBa_J18918) was added between the mRFP1 and the ssrA tag. This is a sister construction to our sfGFP TEV reporter (Ba_K3147000). This construction was used to test the specificity of KARMA using a VHH targeting GFP. In the presence of TEV the ssrA is cleaved and mRFP1 is not degraded anymore. | This construction produces an mRFP1 (BBA_E1010)[1] fused in C-ter with a ssrA fast degradation tag [2] (BBA_M0050). The TEV cutting site (BBa_J18918) was added between the mRFP1 and the ssrA tag. This is a sister construction to our sfGFP TEV reporter (Ba_K3147000). This construction was used to test the specificity of KARMA using a VHH targeting GFP. In the presence of TEV the ssrA is cleaved and mRFP1 is not degraded anymore. | ||
| Line 13: | Line 13: | ||
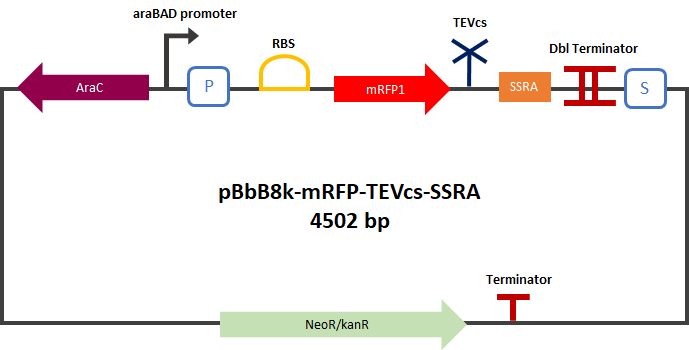
<div align="center">[[File:designK3147003.png|650px]]</div> | <div align="center">[[File:designK3147003.png|650px]]</div> | ||
| − | <div align="center"><b>Figure 1</b>: Construct Design: mRFP1 fused to an | + | <div align="center"><b>Figure 1</b>: Construct Design: mRFP1 fused to an ssrA proteolysis tag with a TEV cutting site between the two.</div> |
===II. Proof of function=== | ===II. Proof of function=== | ||
Revision as of 14:40, 20 October 2019
I : parts BBa_K3147003 function
This construction produces an mRFP1 (BBA_E1010)[1] fused in C-ter with a ssrA fast degradation tag [2] (BBA_M0050). The TEV cutting site (BBa_J18918) was added between the mRFP1 and the ssrA tag. This is a sister construction to our sfGFP TEV reporter (Ba_K3147000). This construction was used to test the specificity of KARMA using a VHH targeting GFP. In the presence of TEV the ssrA is cleaved and mRFP1 is not degraded anymore.
II. Proof of function
This construction was cloned by Gibson Assembly in a pBbB8k-GFP (https://www.addgene.org/35363/ ) backbone under the control of a pBAD promoter.
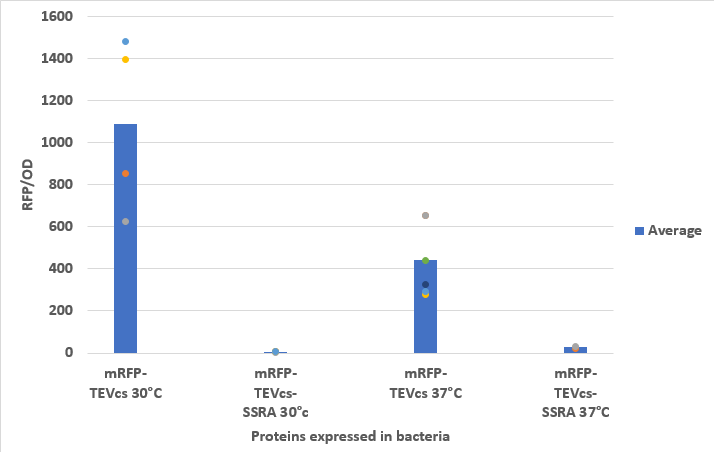
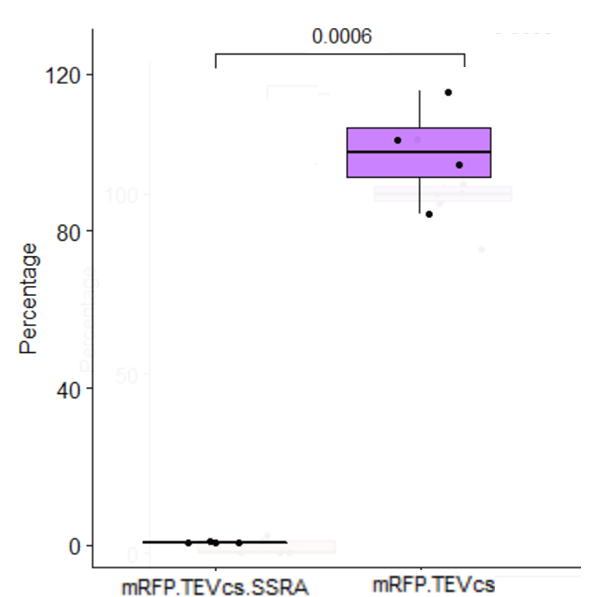
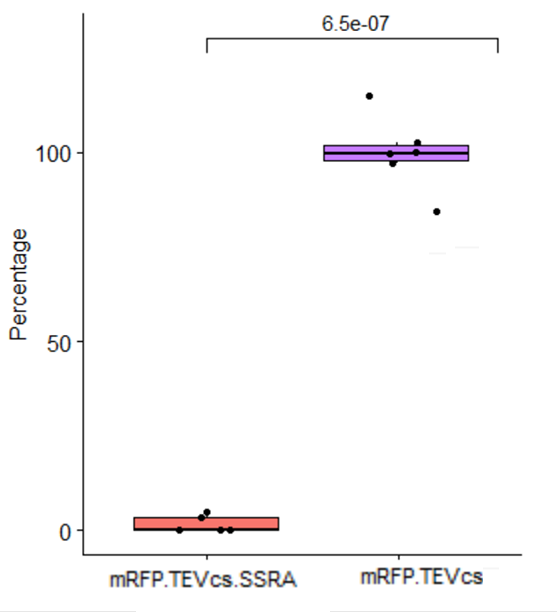
We compared the basal fluorescence of strain E. coli NEB10β transformed with the mRFP1-TEVcs construction to an E. coli NEB10β strain transformed with the mRFP1-TEVcs-ssrA construction. Fluorescence was quantified after overnight induction with 1% arabinose on a plate reader.
Below are the fluorescence measurements of the mRFP1-TEVcs-ssrA and of the mRFP1-TEVcss at 30 and 37°C.
Reference
[1] Levraud, Jean-Pierre et al. 2007. « Identification of the Zebrafish IFN Receptor: Implications for the Origin of the Vertebrate IFN System ». The Journal of Immunology 178(7): 4385‑94. doi:10.4049/jimmunol.178.7.4385
[2] Sunohara, T., Abo, T., Inada, T., & Aiba, H. (2002). The C-terminal amino acid sequence of nascent peptide is a major determinant of SsrA tagging at all three stop codons. RNA (New York, N.Y.), 8(11), 1416–1427. doi:10.1017/s1355838202020198
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 691
- 1000COMPATIBLE WITH RFC[1000]