Difference between revisions of "Part:BBa K3196012"
Orange-Huo (Talk | contribs) |
GlacierHOLE (Talk | contribs) |
||
| Line 2: | Line 2: | ||
__NOTOC__ | __NOTOC__ | ||
<partinfo>BBa_K3196012 short</partinfo> | <partinfo>BBa_K3196012 short</partinfo> | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
<!-- --> | <!-- --> | ||
| Line 30: | Line 12: | ||
<partinfo>BBa_K3196012 parameters</partinfo> | <partinfo>BBa_K3196012 parameters</partinfo> | ||
<!-- --> | <!-- --> | ||
| + | The FLO10 was combined with the guiding peptide sequence of saccharomyces cerevisiae to form the signal peptide FLO10- αpro.FLO10 αpro is a combined signal peptide, which enhance the enzyme activity 3 times. | ||
| + | |||
| + | |||
| + | <h1>'''Characterization'''</h1> | ||
| + | This is a four section for degrade and transfer lignin part. | ||
| + | [[File:T--HUST--China--2019-FLO10SLAC.jpg |525px|thumb|center|Figure1. AOX1-FLO10-SLAC-His tag.]] | ||
| + | |||
| + | <h1>'''DNA Gel Electrophoretic'''</h1> | ||
| + | After we link FLO10 and SLAC successfully, we run the PCR with an intention to confirm the expression of SLAC. As the figure shows, we get the genetic stripe about 2208 bp which means the PCR is successful. | ||
| + | [[File:T--HUST-China--2019-DNA Gel Electrophoretic.png|400px|thumb|center|Figure1:This is the result of the SacⅠ single endonuclease digestion plasmid of the TOP10 strain, we will extract the gel and obtain the aimed DNA. ]] | ||
| + | |||
| + | <h1>'''SDS-PAGE'''</h1> | ||
| + | We run the SDS-PAGE to check whether FLO10 help the enzyme transfer to the extracellular. As the figure shows, we get the protein type about 35 KDa which means the SDS-PAGE is successful. | ||
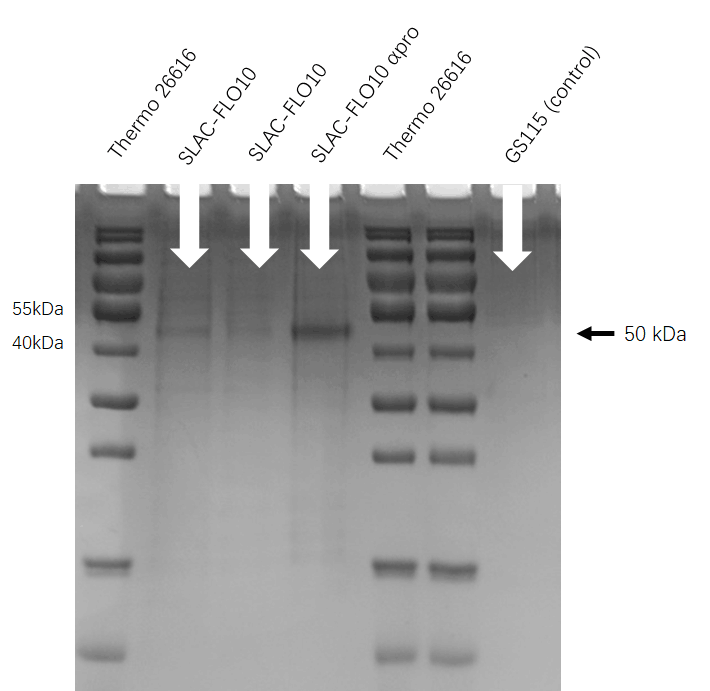
| + | [[File:T--HUST-China--2019-SLAC-SDS-PAGE.jpg |400px|thumb|center|Figure3:the SDS-page result shows that Pichia pastoris have successfully secrete SLAC protein into superfluous liquid.]] | ||
| + | |||
| + | |||
| + | <h1>'''Enzyme Activity'''</h1> | ||
| + | We use ABTS to detect the enzyme activity. As the figure shows, the solution turns green, which confirm the enzyme activity. | ||
| + | [[File:T--HUST-China--2019-SLAC enzyme activity.png|400px|thumb|center|Figure4:these four pictures shows the enzyme activity changes with the time. The four kinds of Pichia pastoris have the different curve, and the most activity strain is FLO10-apro, PHO5apro shows a more delayed increase on enzyme activity.]] | ||
Revision as of 07:14, 20 October 2019
AOX1-Kozak-FLO10 pro-SLAC-His tag-AOX1 Terminator
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 937
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 1586
The FLO10 was combined with the guiding peptide sequence of saccharomyces cerevisiae to form the signal peptide FLO10- αpro.FLO10 αpro is a combined signal peptide, which enhance the enzyme activity 3 times.
Characterization
This is a four section for degrade and transfer lignin part.
DNA Gel Electrophoretic
After we link FLO10 and SLAC successfully, we run the PCR with an intention to confirm the expression of SLAC. As the figure shows, we get the genetic stripe about 2208 bp which means the PCR is successful.
SDS-PAGE
We run the SDS-PAGE to check whether FLO10 help the enzyme transfer to the extracellular. As the figure shows, we get the protein type about 35 KDa which means the SDS-PAGE is successful.
Enzyme Activity
We use ABTS to detect the enzyme activity. As the figure shows, the solution turns green, which confirm the enzyme activity.