Difference between revisions of "Part:BBa K608012"
Night12138 (Talk | contribs) m (Add Tongji_China's works. -- committed by Night12138) |
|||
| Line 82: | Line 82: | ||
|} | |} | ||
| + | |||
| + | =='''Characterization: Average Fluorescence per Cell'''== | ||
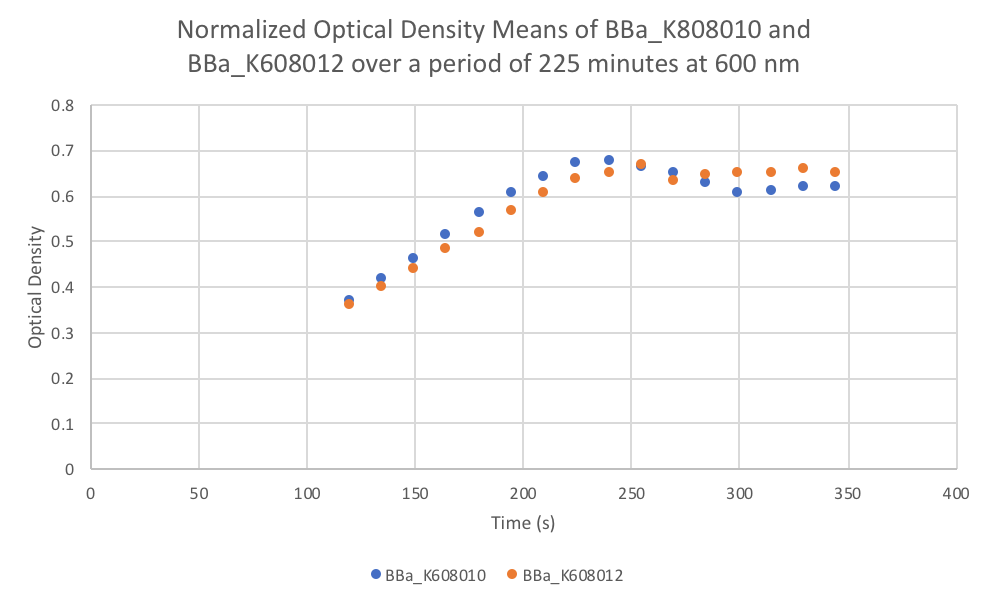
| + | [[File:T--FDR-HB Peru--BBa K608010 and BBa K608012 characterization graph 1.png|thumb|400px|left|<b>Figure 1.</b>]] | ||
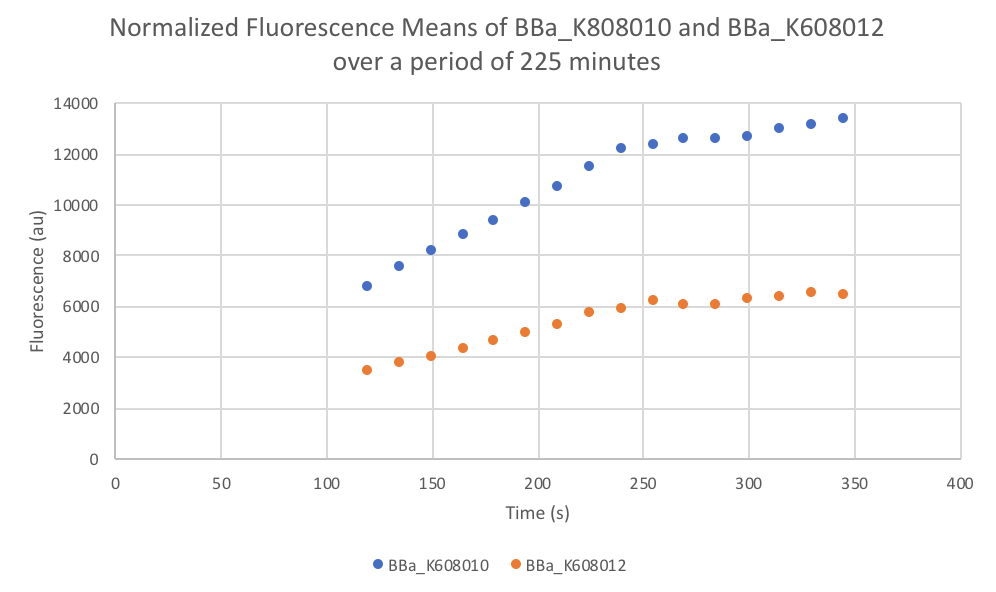
| + | [[File:T--FDR-HB Peru--BBa K608010 and BBa K608012 characterization graph 2.png|thumb|400px|left|<b>Figure 2.</b>]] | ||
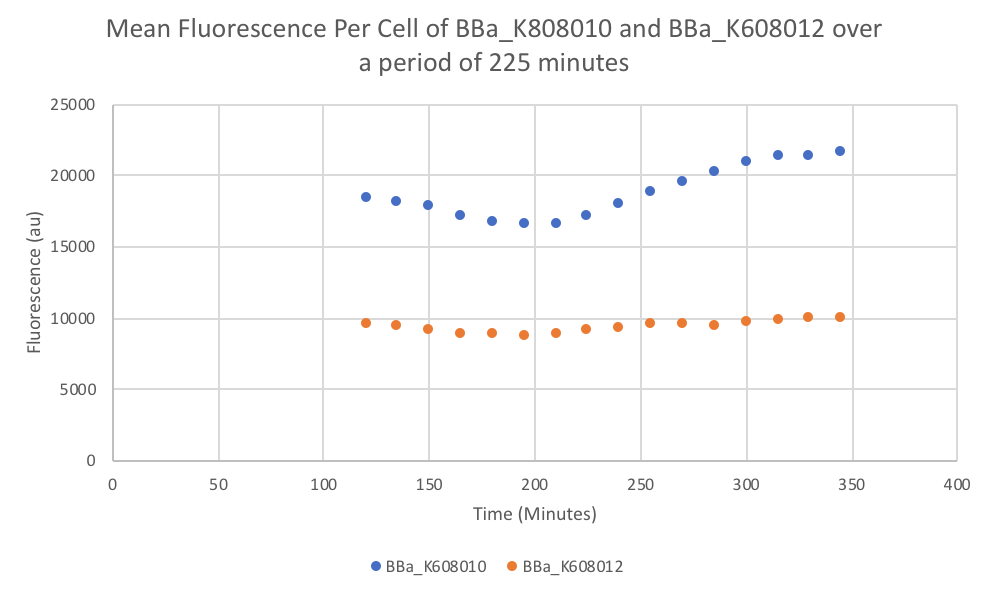
| + | [[File:Graph of "Mean Fluorescence Per Cell of BBa K808010 and BBa K608012 over a period of 225 minutes".png|thumb|400px|left|<b>Figure 3.</b>]] | ||
| + | '''Group:''' FDR-HB_Peru | ||
| + | <br> | ||
| + | <br> | ||
| + | '''Author: '''Marry Xuan | ||
| + | <br> | ||
| + | <br> | ||
| + | '''Summary: '''We compared parts BBa_K608010 and BBa_K608012 to test if the strength of the RBS correlates to the average amount of fluorescence per cell. | ||
| + | <br> | ||
| + | <br> | ||
| + | '''Method: ''' | ||
| + | <br> | ||
| + | <ol> | ||
| + | <li>Dilute 10 ul of the culture (BBa_K608010 and BBa_K608012) into 90ul of M9 + Glucose (0.2% w/v) + CAS amino acids (0.2%) medium in the plate reader wells | ||
| + | <li>Use a plate reader to read the optical density at 600nm (OD). | ||
| + | <li>Use that data to calculate the dilution of the culture to reach an OD reading of 0.1 and dilute with medium (M9 + Glucose + CAS). | ||
| + | <li>Take 200 µl of the previous dilution and add it to a well on the plate reader (fill 3 wells for both K608010 and K608012) | ||
| + | <li>Fill 3 wells with blank medium. | ||
| + | <li>Put the plate in the plate reader and collect the data for 345 minutes, reads both OD and Fluorescence (We used a low gain since the fluorescence was too intense). | ||
| + | <li>Once the data was collected, the OD and Fluorescence data was normalized by subtracting the blank. Then to calculate the average fluorescence per cell, we used the following equation: (FLOnorm)/(ODnorm) | ||
| + | </ol> | ||
| + | <br> | ||
| + | '''Results:''' We found out that after 345 minutes, the average fluorescence of the GFP with a strong RBS (BBa_K608010) was 2.18 times greater than the average fluorescence of the GFP with a weak RBS (BBa_K608012). This data counters the data from the original experiment performed, where the fluorescence intensity of BBa_K608010 was 5.86 times lower than the fluorescence intensity of BBa_K608012. | ||
<!-- Add more about the biology of this part here | <!-- Add more about the biology of this part here | ||
Revision as of 21:00, 18 October 2019
Medium promoter with weak RBS and GFP
Medium promoter from the constitutive promoter family combined with a weak RBS (PR6) and tagged with GFP to quantify the gene expression.
The GFP fluorescence was measured with a plate reader:
The fluorescence intensity and protein concentration were measured with the FLUOstar Omega,
which is a multi-mode microplate reader.
Samples were pipetted into the microplate and analyzed via the plate reader. In this experiment we focused on the protein concentration and the fluorescence intensity of RFP.
We measured the protein concentration with the bradford-assay. This is a method to determine the total protein
concentration. To analyze the protein concentration of the samples, Coomassie Brillant Blue was pippeted to each sample. With the binding of the dye to the proteins the color changes from dark red to blue. The more protein in the solution the more Coomassie dye can bind to proteins and the more the color changes into blue. The absorption of bound Coomassie dye is 595nm. The absorbance is proportional with the amount of bound dye. With a series of Bovine Serum Albumin (BSA) measurements the exact protein concentration of the samples can be determined. BSA acts like a “marker” because the concentration of BSA is known and with a linear calibration line the exact protein concentration can be detected.
GFP served as a reporter of expression. We wanted to know how strong the promoter and RBS activity is. With this reporter gene it was possible to analyze the expression via plate reader. GFP is excited at a wavelength of 509nm and has an emission of 520nm. The plate reader illuminates the samples with a high energy xenon flash lamp. Optical filters or monochromator create the exact wavelength. The more GFP in the sample the higher is the GFP fluorescence intensity. The intensity is collected with the second optical system and is detected with a side window photomultiplier tube.
Promoter and RBS:
PR1: strong Promoter (J23104) strong RBS (B0034)
PR2: strong Promoter (J23104) medium RBS (B0032)
PR3: strong Promoter (J23104) weak RBS (B0031)
PR4: medium Promoter (J23110) strong RBS (B0034)
PR5: medium Promoter (J23110) medium RBS (B0032)
PR6: medium Promoter (J23110) weak RBS (B0031)
| sample | PR2 | PR3 | PR4 | PR5 | PR6 |
| GFP fluorescence intensity | 11378.5 | 1445.0 | 4596.2 | 41221.1 | 26922.7 |
| factor | 7.9 | 1.0 | 3.2 | 28.5 | 18.6 |
The results of this test show that PR6 has 18.6 times higher expression of GFP in comparison with with PR3 which has the lowest expression. The fluorescence intensity of GFP varies, and because of lack of time we could not repeat this experiment. We have also tested the promotor and RBS activity with RFP as a reporter and the results deviate from this experiment. So we are looking forward to test this system another time.
Tongji_China 2019 Characterization
| Sample | PR2(BBa_K608008) | PR5(BBa_K608011) | PR6(BBa_K608012) |
| GFP Fluorescence Intensity | 24541.23 | 62923.91 | 41902.30 |
| Fluorescence Intensity Factor | 1.0 | 2.6 | 1.7 |
| Positive Rate | 0.7563 | 0.9190 | 0.8579 |
Characterization: Average Fluorescence per Cell
Group: FDR-HB_Peru
Author: Marry Xuan
Summary: We compared parts BBa_K608010 and BBa_K608012 to test if the strength of the RBS correlates to the average amount of fluorescence per cell.
Method:
- Dilute 10 ul of the culture (BBa_K608010 and BBa_K608012) into 90ul of M9 + Glucose (0.2% w/v) + CAS amino acids (0.2%) medium in the plate reader wells
- Use a plate reader to read the optical density at 600nm (OD).
- Use that data to calculate the dilution of the culture to reach an OD reading of 0.1 and dilute with medium (M9 + Glucose + CAS).
- Take 200 µl of the previous dilution and add it to a well on the plate reader (fill 3 wells for both K608010 and K608012)
- Fill 3 wells with blank medium.
- Put the plate in the plate reader and collect the data for 345 minutes, reads both OD and Fluorescence (We used a low gain since the fluorescence was too intense).
- Once the data was collected, the OD and Fluorescence data was normalized by subtracting the blank. Then to calculate the average fluorescence per cell, we used the following equation: (FLOnorm)/(ODnorm)
Results: We found out that after 345 minutes, the average fluorescence of the GFP with a strong RBS (BBa_K608010) was 2.18 times greater than the average fluorescence of the GFP with a weak RBS (BBa_K608012). This data counters the data from the original experiment performed, where the fluorescence intensity of BBa_K608010 was 5.86 times lower than the fluorescence intensity of BBa_K608012.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 7
Illegal NheI site found at 30 - 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 707