Difference between revisions of "Part:BBa K1399004"
m |
Carolinarp (Talk | contribs) |
||
| Line 11: | Line 11: | ||
[3] Purcell, O., Grierson, C. S., Bernardo, M. Di & Savery, N. J. Temperature dependence of ssrA-tag mediated protein degradation. J. Biol. Eng. 6, 10 (2012). | [3] Purcell, O., Grierson, C. S., Bernardo, M. Di & Savery, N. J. Temperature dependence of ssrA-tag mediated protein degradation. J. Biol. Eng. 6, 10 (2012). | ||
| + | |||
| + | ===Contribution=== | ||
| + | Group: Valencia_UPV iGEM 2018 | ||
| + | Authors: Adrián Requena, Carolina Ropero | ||
| + | Part BBa_K2656013 is the GFPmut3b coding sequence with the LVA degradation tag. It is the BBa_K1399004 original sequence adapted to be Golden Gate compatible. Thus, it can be used with Biobrick and Golden Braid assembly methods. | ||
<partinfo>BBa_K1399004 SequenceAndFeatures</partinfo> | <partinfo>BBa_K1399004 SequenceAndFeatures</partinfo> | ||
Revision as of 23:51, 17 October 2018
GFP (mut3b) with LVA-ssrA degradation tag
GFP (mut3b) ( Part:BBa_E0040) with added LVA-ssrA degradation tag. The tag increases GFP turn-over rate, thus providing better temporal resolution of green fluorescence. In the same time, maximal fluorescence amplitudes will be lower as newly formed protein is degraded as soon as it is formed.
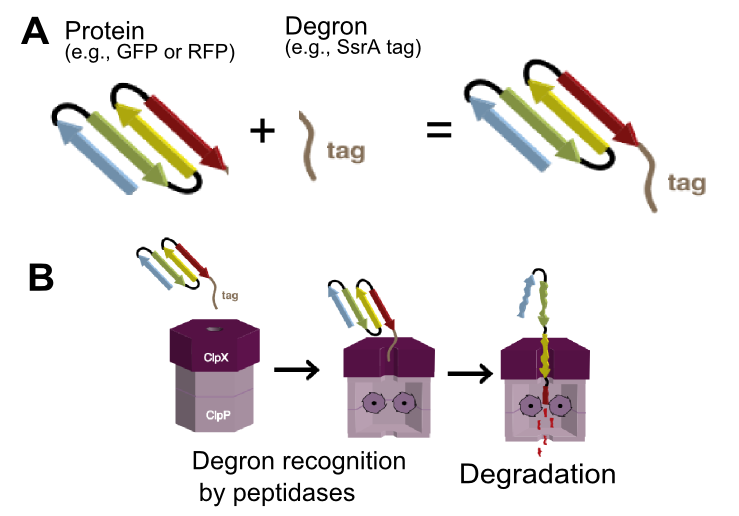
The tag encodes peptide sequence AANDENYALVA and is recognized by ClpA and ClpX unfoldases and ClpX mediator SspB.[1] ClpA and ClpX then form a proteosome-like complex with ClpP protease and the protein is degraded (Figure 1).[1]
The final three residues of the tag determines the strength of interaction with ClpX and thus the final protein degradation rate.[2] The LVA tag is reported to lead to fast protein degradation, degrading GFP with rate -0.018 per minute.[2] However, be aware that exact protein degradation rate depends on multiple factors: ClpXP and ClpAP protease and SspB mediator concentrations, protein stability, Km of binding to the protease, temperature [3].
References
[1] Flynn, J. M. et al. Overlapping recognition determinants within the ssrA degradation tag allow modulation of proteolysis. Proc. Natl. Acad. Sci. U. S. A. 98, 10584–9 (2001). [2] Andersen, J. B. et al. New unstable variants of green fluorescent protein for studies of transient gene expression in bacteria. Appl. Environ. Microbiol. 64, 2240–6 (1998). [3] Purcell, O., Grierson, C. S., Bernardo, M. Di & Savery, N. J. Temperature dependence of ssrA-tag mediated protein degradation. J. Biol. Eng. 6, 10 (2012).
Contribution
Group: Valencia_UPV iGEM 2018 Authors: Adrián Requena, Carolina Ropero Part BBa_K2656013 is the GFPmut3b coding sequence with the LVA degradation tag. It is the BBa_K1399004 original sequence adapted to be Golden Gate compatible. Thus, it can be used with Biobrick and Golden Braid assembly methods.
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 644