Difference between revisions of "Part:BBa J23119"
| Line 16: | Line 16: | ||
| − | <p>GreatBay_China have made two <b> improved parts </b> based on J23119. <b> UP119</b> is modified by addition of an up-element. ( | + | <p>GreatBay_China have made two <b> improved parts </b> based on J23119. <b> UP119</b> is modified by addition of an up-element. ([[Part:BBa_K2753055]])<b>TALE2 sp6 </b>is a TALE stabilized variant of J23119. ([[Part:BBa_K2753023]])</p><br> |
<!-- --> | <!-- --> | ||
Revision as of 15:08, 13 October 2018
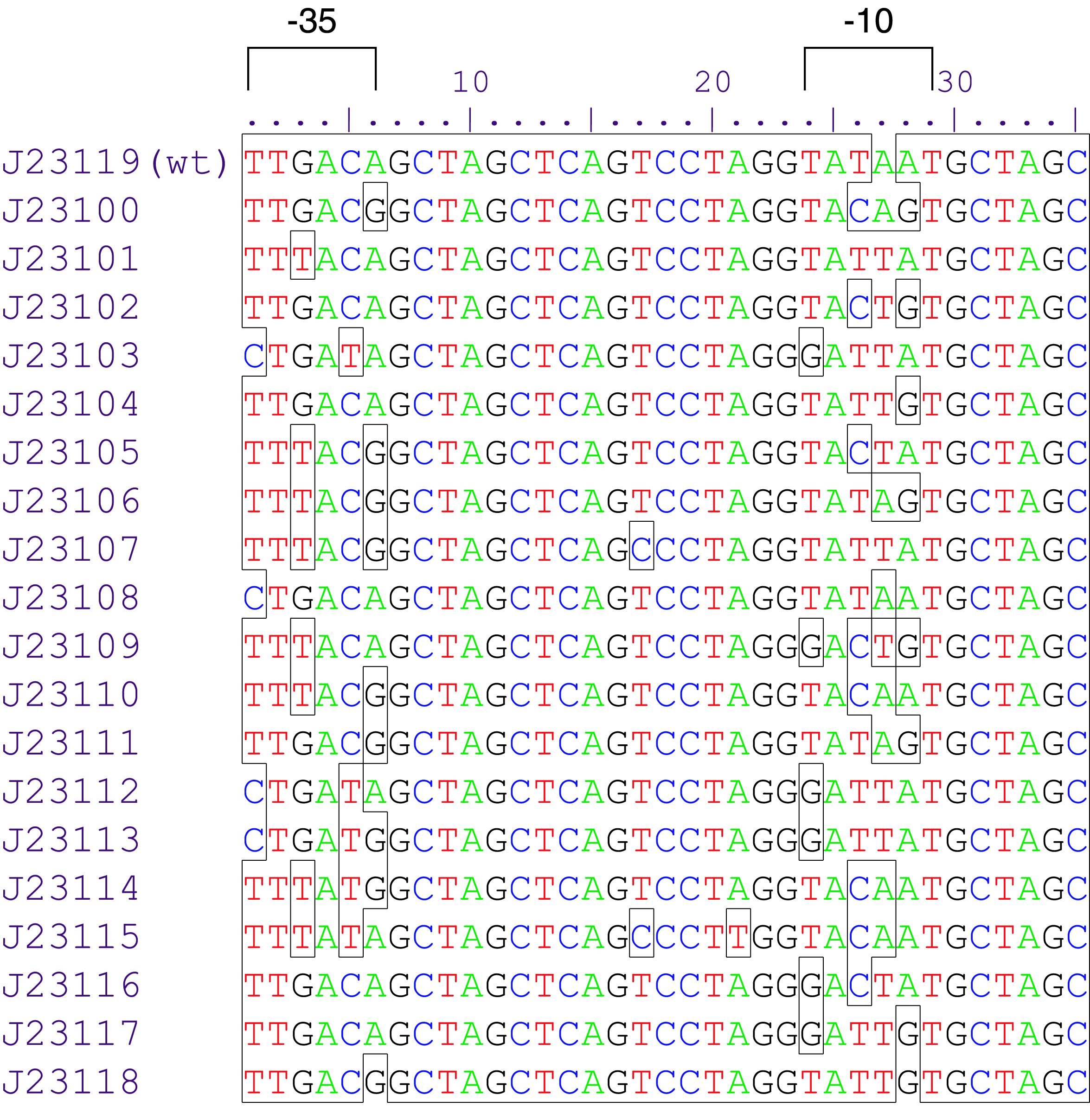
constitutive promoter family member
Variant RFP (au) J23112 1 J23103 17 J23113 21 J23109 106 J23117 162 J23114 256 J23115 387 J23116 396 J23105 623 J23110 844 J23107 908 J23106 1185 J23108 1303 J23118 1429 J23111 1487 J23101 1791 J23104 1831 J23102 2179 J23100 2547 |
Constitutive promoter family
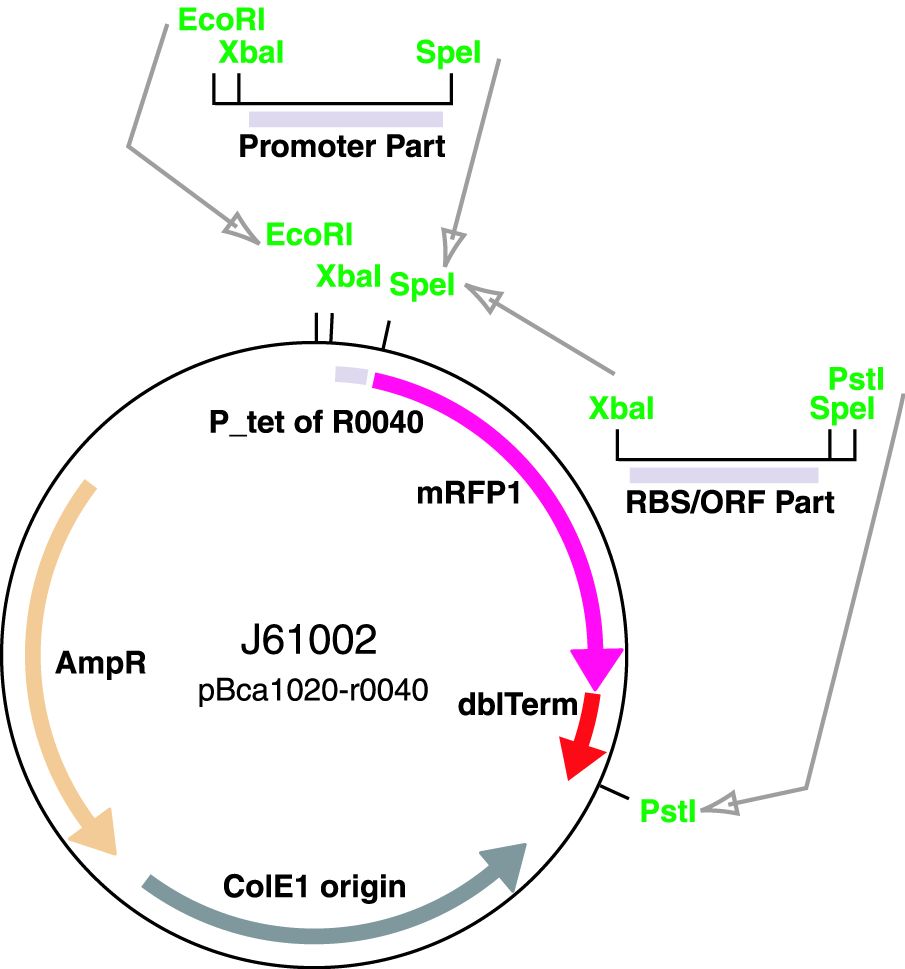
Parts J23100 through J23119 are a family of constitutive promoter parts isolated from a small combinatorial library. J23119 is the "consensus" promoter sequence and the strongest member of the family. All parts except J23119 are present in plasmid J61002. Part J23119 is present in pSB1A2. This places the RFP downstream of the promoter. Reported activities of the promoters are given as the relative fluorescence of these plasmids in strain TG1 grown in LB media to saturation. See part BBa_J61002 for details on their use.
These promoter parts can be used to tune the expression level of constitutively expressed parts. The NheI and AvrII restriction sites present within these promoter parts make them a scaffold for further modification. JCAraw
Usage and Biology
iGEM16-CLSB-UK:The consensus promoter Part:BBa_J23119 was assembled with AmilCP blue-purple chromoprotein Part:BBa_K592025. The construct Part:BBa_K2078002 was carried on the pSB1C3 plasmid for expression and amplification in E-Coli.
GreatBay_China 2018:
Team GreatBay_China 2018 characterized J23119, Part:BBa_J23105, and Part:BBa_J23101 by assembling them with Part:BBa_B0034 and a sfGFPPart:BBa_I746916 on three vectors: pUC20 (copy number about 500/cell), pR6K (copy number about 15/cell), pSC101 (copy number about 2/cell). Then we measured the fluorescence by Flow Cytometry as a reference for the TALE stabilized promoter library.
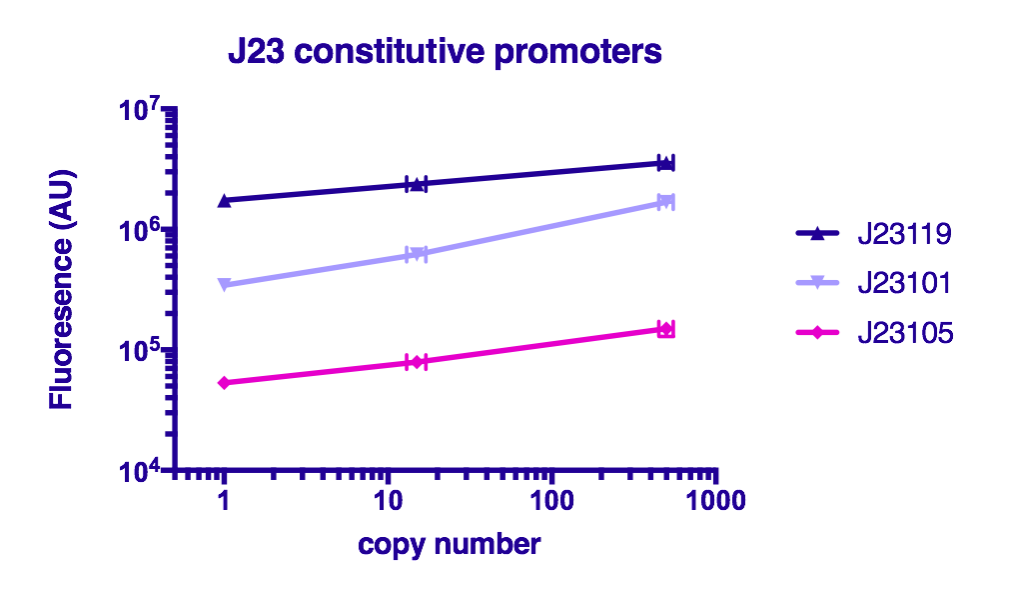
The result indicate that the strength of J23119, J23105, and J23101 are about the same as described by team iGEM2006_Berkeley, and the fluorescence increases as the copy number of the vector increases
GreatBay_China have made two improved parts based on J23119. UP119 is modified by addition of an up-element. (Part:BBa_K2753055)TALE2 sp6 is a TALE stabilized variant of J23119. (Part:BBa_K2753023)
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 7
Illegal NheI site found at 30 - 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
RBS
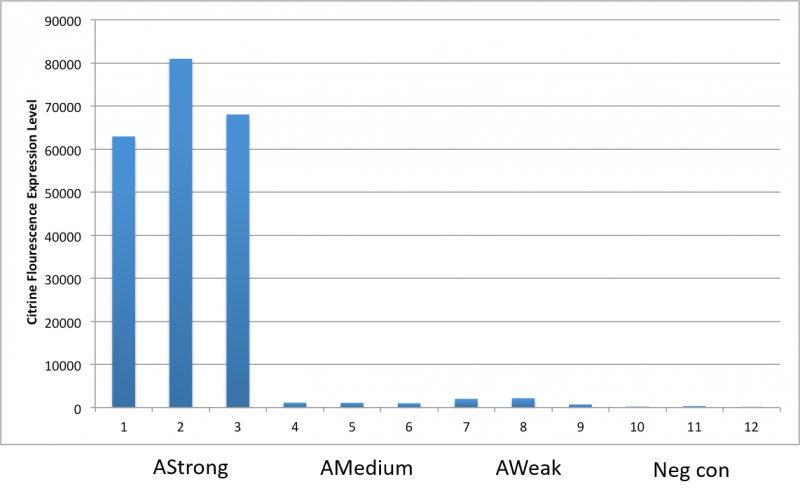
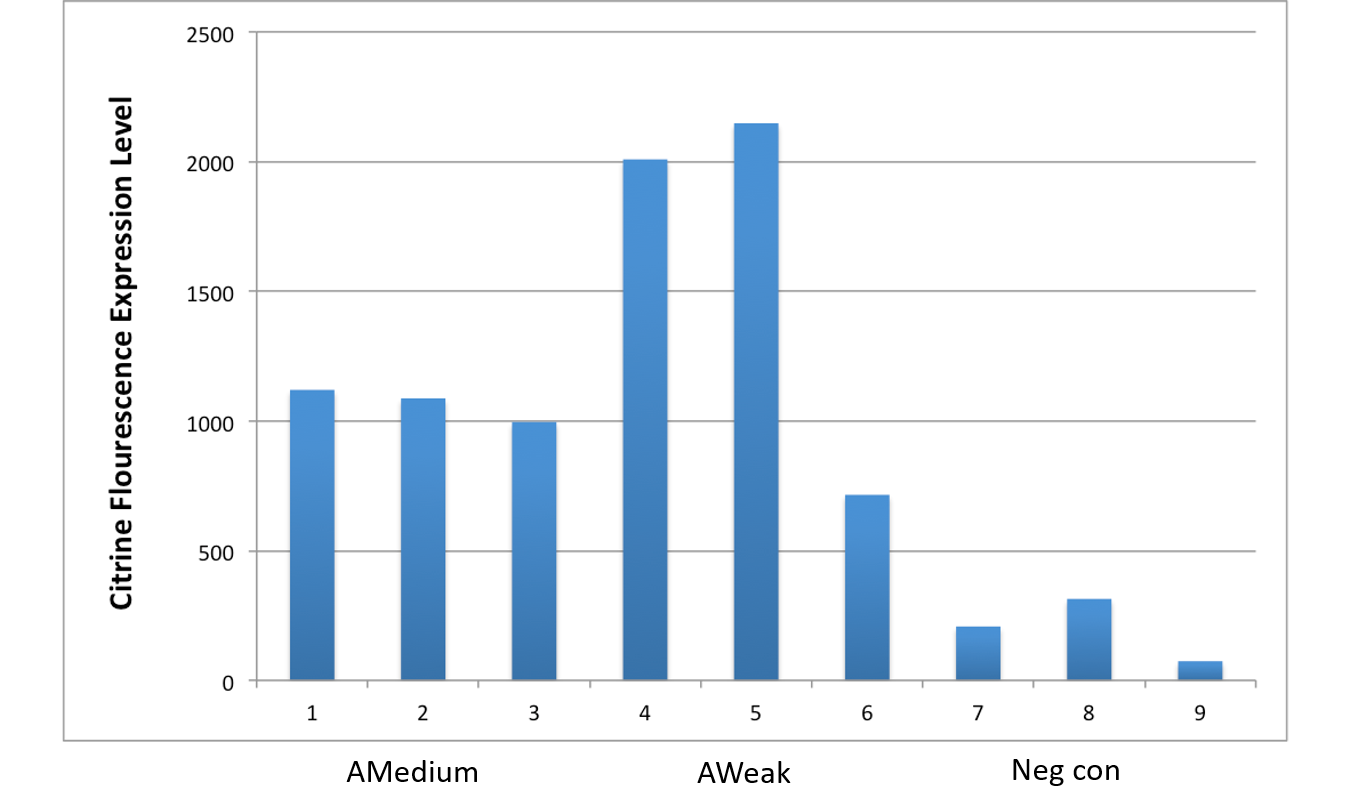
After an attempt to clone this promoter upstream of the Citrine reporter gene yielded poor expression, the Yale iGEM teaminspected the Ribosome Binding Site used with the promoters and realized that it would need to be improved. We used the Salis Lab Ribosome Binding Site Calculator to design a stronger Ribosome Binding Site.
Below, we demonstrate that the newly designed Ribosome Binding Site is appropriately compatible with the Anderson Promoter collection.