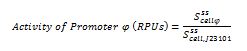Difference between revisions of "Part:BBa J23102:Experience"
(→Unexpected LuxR-AHL repressible behaviour) |
(→User Reviews) |
||
| Line 69: | Line 69: | ||
With the previous results of the characterization of the promoters there is concluded that the promoter J23107, is the strongest because it produces more RPUs” | With the previous results of the characterization of the promoters there is concluded that the promoter J23107, is the strongest because it produces more RPUs” | ||
| + | |||
| + | <br /><br /> | ||
| + | <I>'''iGEM Bulgaria 2017''' Regulation of the J23102 promoter activity by CRISPi</I> | ||
| + | |||
| + | |||
| + | We offer an improvement to the J23102 part. We designed 2 gRNAs that target this promoter (TTGACAGCTAGCTCAGTCCT and TAGTAGCTAGCACAGTACCT) and cloned them into our gRNA expression vector (Part:BBa_K2515002). These constructs gave us the ability to control the J23102 promoter activity in a strain with dCas9 unther teh control of an inducible arabinose promoter. We believe that this type of regulation can be transffered to the other members of this promoter family if one redesign the gRNAs. | ||
Revision as of 22:19, 1 November 2017
This experience page is provided so that any user may enter their experience using this part.
Please enter
how you used this part and how it worked out.
Applications of BBa_J23102
Unexpected LuxR-AHL repressible behaviour
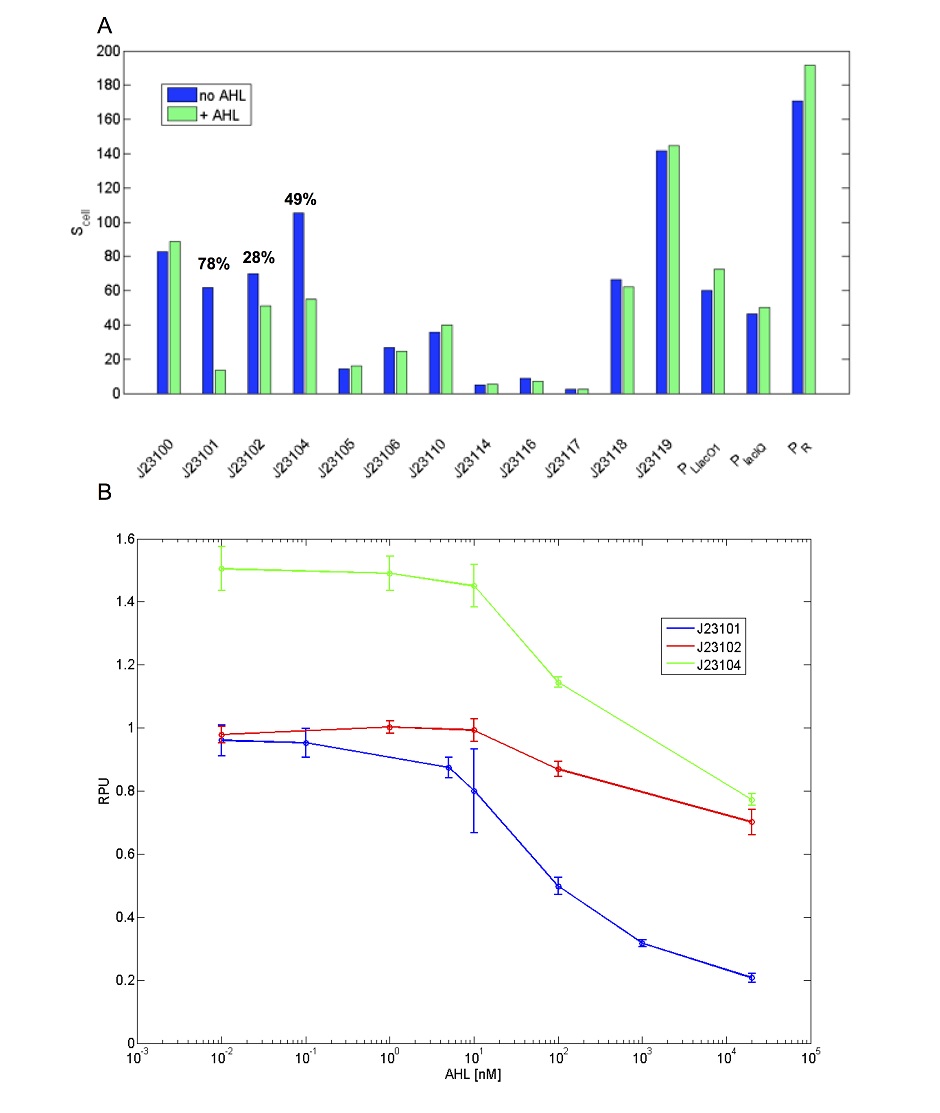
This promoter shows activity repression as a function of AHL in presence of LuxR [ADD REFERENCE LINK]. Results are shown below for a set of widely used promoters tested in the same conditions (TOP10, M9 medium supplemented with casamino acids and glycerol, assayed in a microplate reader). Promoters drive the BBa_I13507 RFP expression device. The TACTAGTG scar is present between promoter and RBS, except for PLlacO1 (BBa_R0011), PlacIQ (BBa_I14032) and PR (BBa_R0051) where the scar is TACTAGAG. RFP was measured and used to compute Scell (arbitrary units) or RPU values. Promoters were assembled in pSB4C5 and were co-transformed with BBa_S03119 in pSB3K3 in TOP10. Blue bars represent the activity when no AHL was added to the medium, while green bars represent the repressed activity (AHL 20 µM was added to the medium). Only BBa_J23101, BBa_J23102 and BBa_J23104 showed repression and, for them, percent repression is reported.
Evaluation of Anderson promoter J23102 in B. subtilis by iGEM-Team LMU-Munich 2012
This Anderson promoter was evaluated without fused RFP with the lux operon as a reporter in B. subtilis. See the new BioBrick BBa_K823006 without RFP and have a look at the [http://2012.igem.org/Team:LMU-Munich/Data/Anderson Data] from the evaluation in B. subtilis.
User Reviews
UNIQ10aef75a184656ee-partinfo-00000000-QINU UNIQ10aef75a184656ee-partinfo-00000001-QINU
iGEM WHU-China 2013 construction of tandem double promoters

figure 1. The seven different combinations of double tandem promoters,from left to right, are J23106,J23102-J23102,J23102-J23106,J23106-J23102,J23106-J23106,J23106-J23116,J23116-J23102 and J23116-J23106.
We use consecutive promoter BBa_J23102 as a basic part to construct tandem double promoters BBa_k1081002(J23102-J23102),BBa_k1081003(J23102-J23106),BBa_k1081004(J23106-J23102) and BBa_k1081007(J23116-J23102).
iGEM CINVESTAV_IPN_UNAM CHARACTERIZATION OF IGEM DISTRIBUTION BIOPARTS
For contribute to the parts registry our team decided to make the characterization of constitutive promoters belonging to the family isolated from a small combinatorial library (J23101 , J23102, J23104, J23107, J23108, J2311, and J23115) which were attached to GFP to determine promoter activity, using the equipment Victor X3 Multilabel Plate Reader.
Fig. 1 Construction of the promoter J23102 expressing GFP.
Methods
With the selected colonies, an overnight culture was made in M9 media(minimal media supplemented with 0.2% CAA). After 12 hours the culture was transferred to a 96 well plate at a 1:10 dilution (20 μl of culture and 180 μL of fresh M9 medium). OD and fluorescence measurements of the selected colonies were performed at intervals of 30 minutes for 16 h. From the results the PopS were calculated (polymerases per second).
Modeling
The ecuations used for calulated de promoter activity were based on (R. K. Jason et. al 2009).
Results
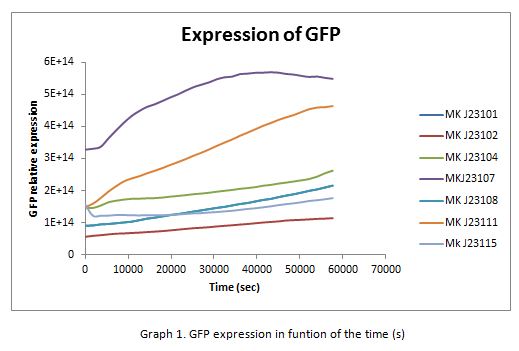
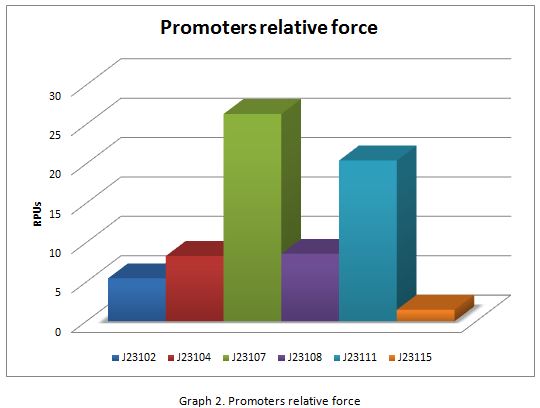
In the following graphs there is shown the GFP expression in function of th time and the realtive promotor intensity.
With the previous results of the characterization of the promoters there is concluded that the promoter J23107, is the strongest because it produces more RPUs”
iGEM Bulgaria 2017 Regulation of the J23102 promoter activity by CRISPi
We offer an improvement to the J23102 part. We designed 2 gRNAs that target this promoter (TTGACAGCTAGCTCAGTCCT and TAGTAGCTAGCACAGTACCT) and cloned them into our gRNA expression vector (Part:BBa_K2515002). These constructs gave us the ability to control the J23102 promoter activity in a strain with dCas9 unther teh control of an inducible arabinose promoter. We believe that this type of regulation can be transffered to the other members of this promoter family if one redesign the gRNAs.