Difference between revisions of "Part:BBa K2201004"
| Line 366: | Line 366: | ||
</html> | </html> | ||
| − | [[file:T--Bielefeld-CeBiTec--Parts µmaxMicrocultivation.jpeg|500px|thumb|left|<b>Figure (5): Graphical determination of the maximum specific growth rate | + | [[file:T--Bielefeld-CeBiTec--Parts µmaxMicrocultivation.jpeg|500px|thumb|left|<b>Figure (5): Graphical determination of the maximum specific growth rate µ<sub>max</sub> for the microcultivations. </b>]] |
<html> | <html> | ||
Revision as of 16:41, 30 October 2017
Nucleotide Transporter PtNTT2 from Phaeodactylum tricornutum
Usage and Biology
Phaeodactylum tricornutum, a diatom of the genus Phaedactylum, features six putative nucleotide transporters (NTTs). Two isoforms of these NTTs have been characterized by Ast et al. 2009 and it was shown that both isoforms facilitate transport across the plastid membrane. While isoform 1 (NTT1) acts as a proton-dependent adenine nucleotide importer, NTT2 facilitates the counter exchange of (deoxy-)nucleoside triphosphates (Ast et al., 2009).The isoform 2 of the nucleotide transporter was shown to be a broad range (deoxy-)nucleoside transporter, facilitating the uptake of CTP, GTP, dCTP, ATP, UTP, dGTP, dATP and TTP when expressed in E. coli.Zhang et al. 2017 investigated the use of PtNTT2 for the uptake of the unnatural bases dNaM and dTPT3. Therefore, the expression of PtNTT2 was investigated in different strains, under control of different promotors, and plasmid-bound as well as integrated into the chromosome. In their final design, Zhang and colleagues integrated PtNTT2 chromosomally in E. coli BL21(DE3) under control of the lacUV5 promoter. To demonstrate its feasibility for the uptake of nucleotides in E. coli from the media, uptake of [α 32P]-dATP was measured. The native sequence of PtNTT2 features an N-terminal signal sequence directing the subcellular localization to the plastid membrane. In E. coli, this signal sequence is likely to be retained, leading to a growth defect in cells expressing the native PtNTT2 transporter.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Contents
Computational Analysis of PtNTT2
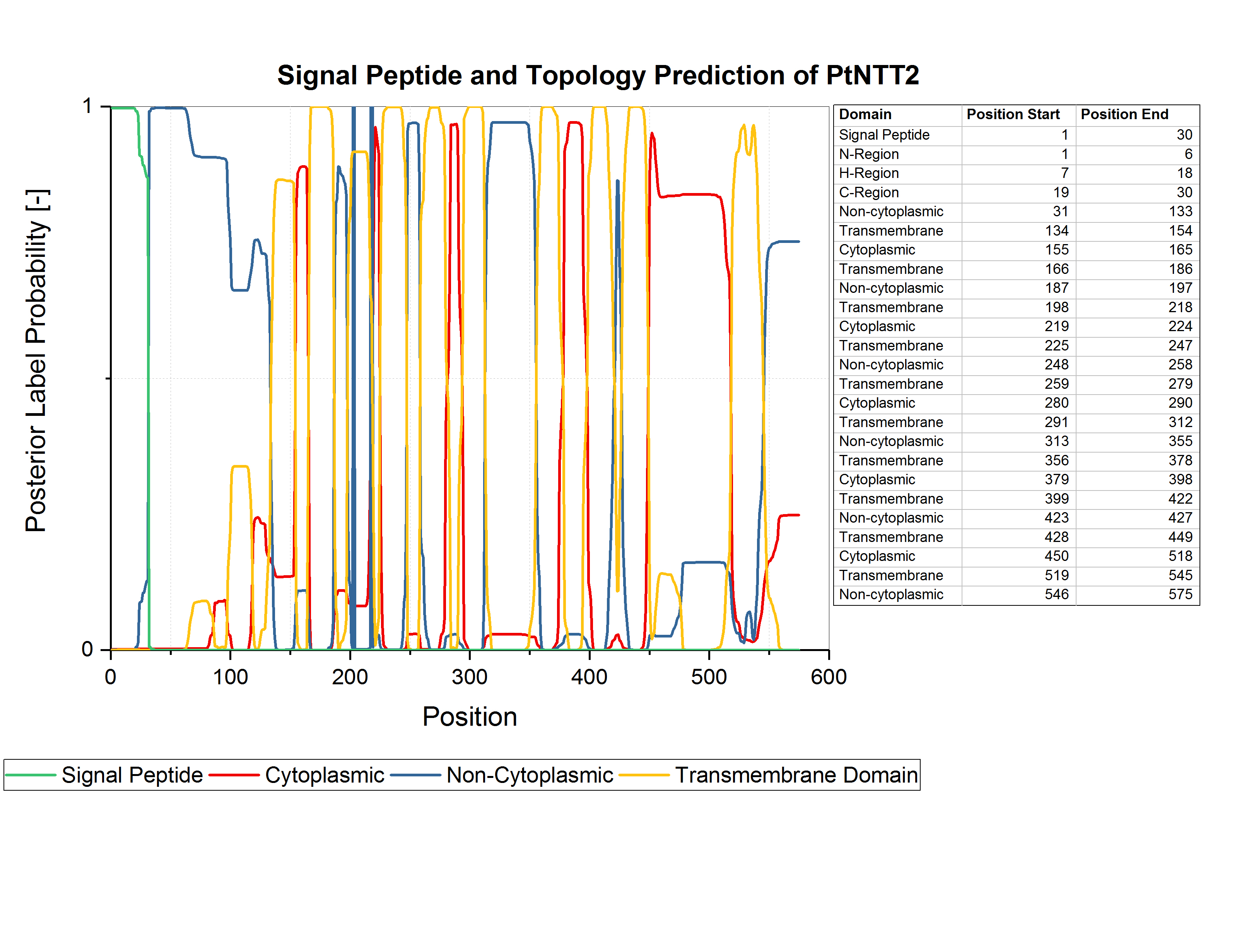
Zhang et al. used an N-terminal truncated version of PtNTT2, lacking the first 65 amino acids, since they observed some kind of toxicity resulting from the native N-terminal sequence (Zhang et al., 2017). For our project, we analyzed the amino acid sequence of PtNTT2 using the prediction software Phobius (Käll et al., 2007). Using Phobius, we analyzed the signal peptide and the transmembrane topology of PtNTT2. The analysis revealed that the native signal peptide is formed by amino acids 1 30, which means that Zhang et al. removed more than the native signal peptide for their experiment. The results of the prediction are shown in figure (1). Analysis of the transmembrane topology of the transporter, which is integrated into the plastid membrane in its native algal cell, shows iterative non-cytoplasmatic, transmembrane and cytoplasmic regions. The topology might indicate, that the transporter will be integrated into the inner membrane when expressed in E. coli.
Plasmid Design
For the analysis and characterization of PtNTT2, a total of eleven plasmids were designed and cloned based on initial research and the computational analysis. The coding sequence of PtNTT2 was codon optimized using the IDT Codon Optimization Tool and ordered as two gBlocks. Using overlap extension PCR, the two gBlocks were put together and inserted into pSB1C3 using Gibson Assembly. The truncated versions of the transporter as well as the versions with new signal peptides were constructed using primers and Gibson Assembly.
Table (1): Designed and cloned plasmids for the analysis and characterization of PtNTT2.
| Plasmid Name | BioBrick Number | Characteristics |
|---|---|---|
| pSB1C3-PtNTT2 | BBa_K2201004 | Only the cds |
| pSB1C3-PlacUV5-PtNTT2 | BBa_K2201000 | cds with lacUV5 promotor and a strong RBS ( BBa_B0034 ) |
| pSB1C3-PlacUV5-PtNTT2(66-575) | BBa_K2201001 | cds with lacUV5 promotor and a strong RBS ( BBa_B0034 ),truncated version lacking the first 65 amino acids |
| pSB1C3-PlacUV5-PtNTT2(31-575) | BBa_K2201005 | cds with lacUV5 promotor and a strong RBS ( BBa_B0034 ), truncated version lacking the first 30 amino acids |
| pSB1C3-PlacUV5-pelB-SP-PtNTT2 | BBa_K2201006 | cds with lacUV5 promotor and a strong RBS ( BBa_B0034 ), native signal peptide replaced with the pelB signal peptide |
| pSB1C3-PlacUV5-TAT-SP-PtNTT2 | BBa_K2201007 | cds with lacUV5 promotor and a strong RBS ( BBa_B0034 ), native signal peptide replaced with a TAT signal peptide |
| pSB1C3-PlacUV5-PtNTT2-GFP | BBa_K2201002 | Fusion protein of BBa_ K2201000 with GFP ( BBa_E0040 ), cMyc epitope tag as linker ( BBa_K2201181 ) |
| pSB1C3-PlacUV5-PtNTT2(66-575)-GFP | BBa_K2201003 | Fusion protein of BBa_ K2201001 with GFP ( BBa_E0040 ), cMyc epitope tag as linker ( BBa_K2201181 ) |
| pSB1C3-PlacUV5-PtNTT2(31-575)-GFP | BBa_K2201011 | Fusion protein of BBa_K2201005 with GFP ( BBa_E0040 ), cMyc epitope tag as linker ( BBa_K2201181 ) |
| pSB1C3-PlacUV5-pelB-SP-PtNTT2-GFP | BBa_K2201012 | Fusion protein of BBa_K2201006 with GFP ( BBa_E0040 ), cMyc epitope tag as linker ( BBa_K2201181 ) |
| pSB1C3-PlacUV5-TAT-SP-PtNTT2-GFP | BBa_K2201013 | Fusion protein of BBa_K2201007 with GFP ( BBa_E0040 ), cMyc epitope tag as linker ( BBa_K2201181 ) |
Cultivations
Shake Flask Cultivation
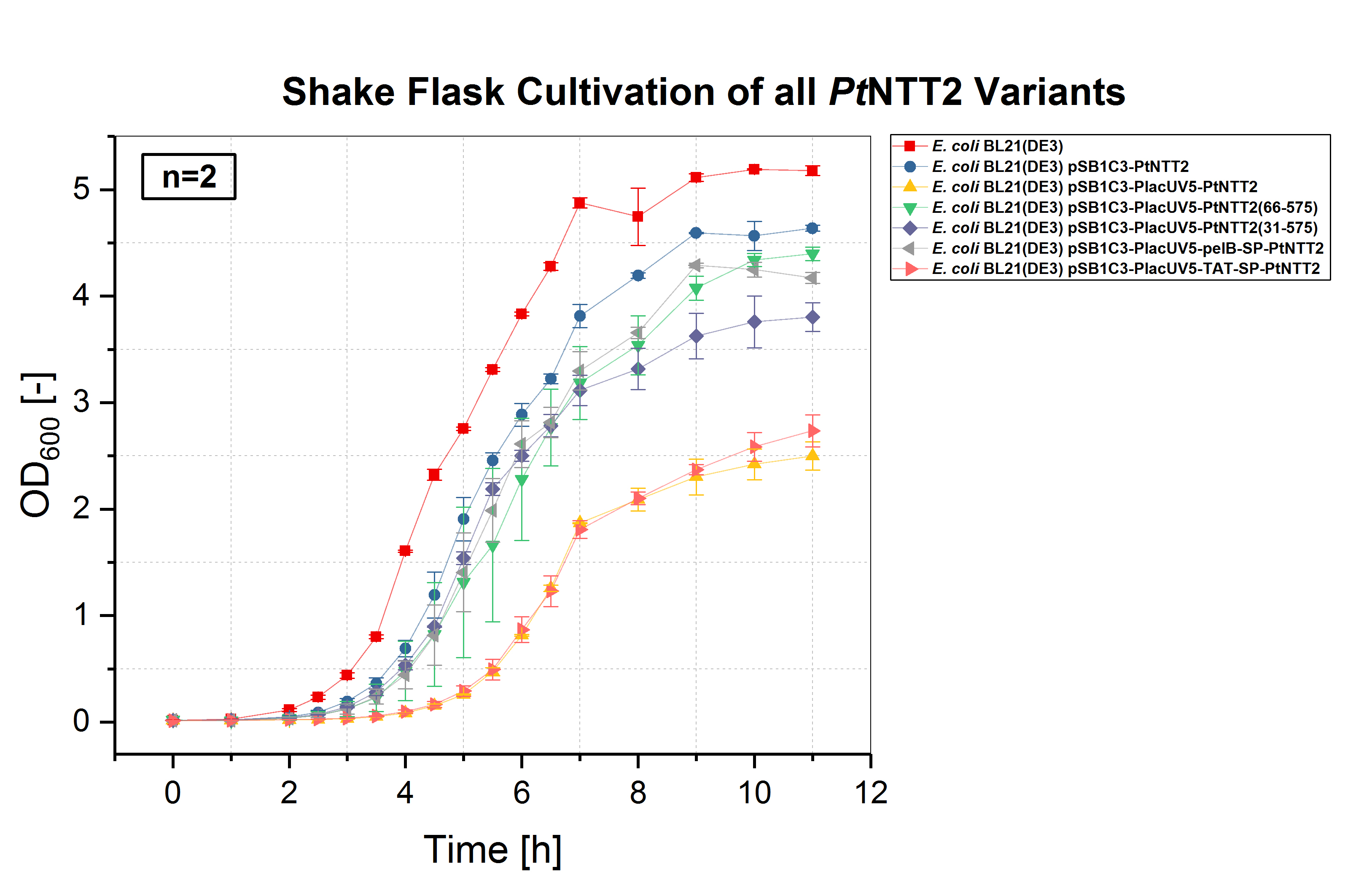
E. coli BL21(DE3) and E. coli BL21(DE3) pSB1C3-PtNTT2, not expressing PtNTT2, were used as negative controls. Two biological replicates of each strain were cultivated and three technical replicates taken for each measurement. A clear difference in the growth rates can be observed, with E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2 and E. coli BL21(DE3) pSB1C3-PlacUV5-TAT-SP-PtNTT2 showing the weakest growth. Both strains also show the longest lag phase, which is nearly twice as long as the lag phase of E. coli BL21(DE3). E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2(66-575) and E. coli BL21(DE3) pSB1C3-PlacUV5-pelB-SP-PtNTT2 show the best growth of all PtNTT2 variants, reaching the highest OD600.
Given that Zhang et al., 2017 reported some kind of toxicity resulting from the N-terminal sequence of PtNTT2, we first investigated if we could also observe the same toxicity associated with the N-terminal sequence. We also tested whether our own versions of the transporter result in better growth and reduced toxicity compared to the native transporter. Therefore, after cloning the plasmids in E. coli DH5α and verifying the correct assembly via sequencing, all plasmids were transformed into E. coli BL21(DE3). The presence of the correct plasmids was again verified by colony PCR.
The first cultivations were carried out in shake flasks in LB media. A total cultivation volume of 50 mL was used. The cultures were incubated at 37 °C and 180 rpm. All cultures were inoculated with an OD600 of 0.01. OD600 was measured every hour during lag and stationary phase and every 30 minutes during the exponential phase. The optical density was measured using an Eppendorf Photometer and standard cuvettes. E. coli BL21(DE3) without a plasmid and E. coli BL21(DE3) harboring pSB1C3-PtNTT2 were used as negative controls. Two biological replicates of each strain were tested and three technical replicates were measured for each timepoint.
E. coli BL21(DE3) without any plasmid shows the best growth, with the highest specific growth rate and the highest final OD600 of 5.178 ± 0.046. The second negative control, E. coli BL21(DE3) harboring pSB1C3-PtNTT2 (BBa_K2201004) reached the second highest OD600 with 4.638 ± 0.029. Of the functional PtNTT2 variants, strains harboring pSB1C3-PlacUV5-PtNTT2(66-575) (BBa_K2201001) and pSB1C3-PlacUV5-pelB-SP-PtNTT2 (BBa_K2201006) reached the highest ODs with 4.397 ± 0.062 and 4.171 ± 0.051, respectively. E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2(31-575) (BBa_K2201005) showed similar growth to the two previous strains during the lag phase and early exponential phase, but reaching a lower OD600 of 3.802 ± 0.135. E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2 (BBa_K2201000) and E. coli BL21(DE3) pSB1C3-PlacUV5-TAT-SP-PtNTT2 (BBa_K2201007) grew significantly weaker compared to all previous strains, with a lag phase nearly four hours long and final ODs of 2.499 ± 0.134 and 2.735 ± 0.150, respectively. The difference in growth between the two negative controls can be explained by the metabolic burden caused by plasmid replication and expression of the chloramphenicol resistance. Therefore, E. coli BL21(DE3) pSB1C3-PtNTT2 is the more accurate control, since all samples contain the same plasmid backbone and were also grown in LB media supplemented with chloramphenicol. The difference in growth between strains expressing the native, full length transporter and the truncated version PtNTT2(66-575), observed by Zhang et al. 2017, could also be shown. This negative effect might be associated with the native signal peptide of PtNTT2, which E. coli might not be able to process correctly. Another explanation for the weak growth could be that the native transporter variant has a higher activity compared to the other variants. If the activity is too high, this might lead to a toxic effect and to the observed weak growth.
Table (2): Final OD600 of all cultures. The highest OD600 was reached by the wildtype E. coli BL21(DE3), the lowest by E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2.
| Strain | Final OD600 [-] |
|---|---|
| E. coli BL21(DE3) | 5.178 ± 0.046 |
| E. coli BL21(DE3) pSB1C3-PtNTT2 | 4.638 ± 0.029 |
| E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2 | 2.499 ± 0.134 |
| E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2(66-575) | 4.397 ± 0.062 |
| E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2(31-575) | 3.802 ± 0.135 |
| E. coli BL21(DE3) pSB1C3-PlacUV5-pelB-SP-PtNTT2 | 4.171 ± 0.051 |
| E. coli BL21(DE3) pSB1C3-PlacUV5-TAT-SP-PtNTT2 | 2.735 ± 0.150 |
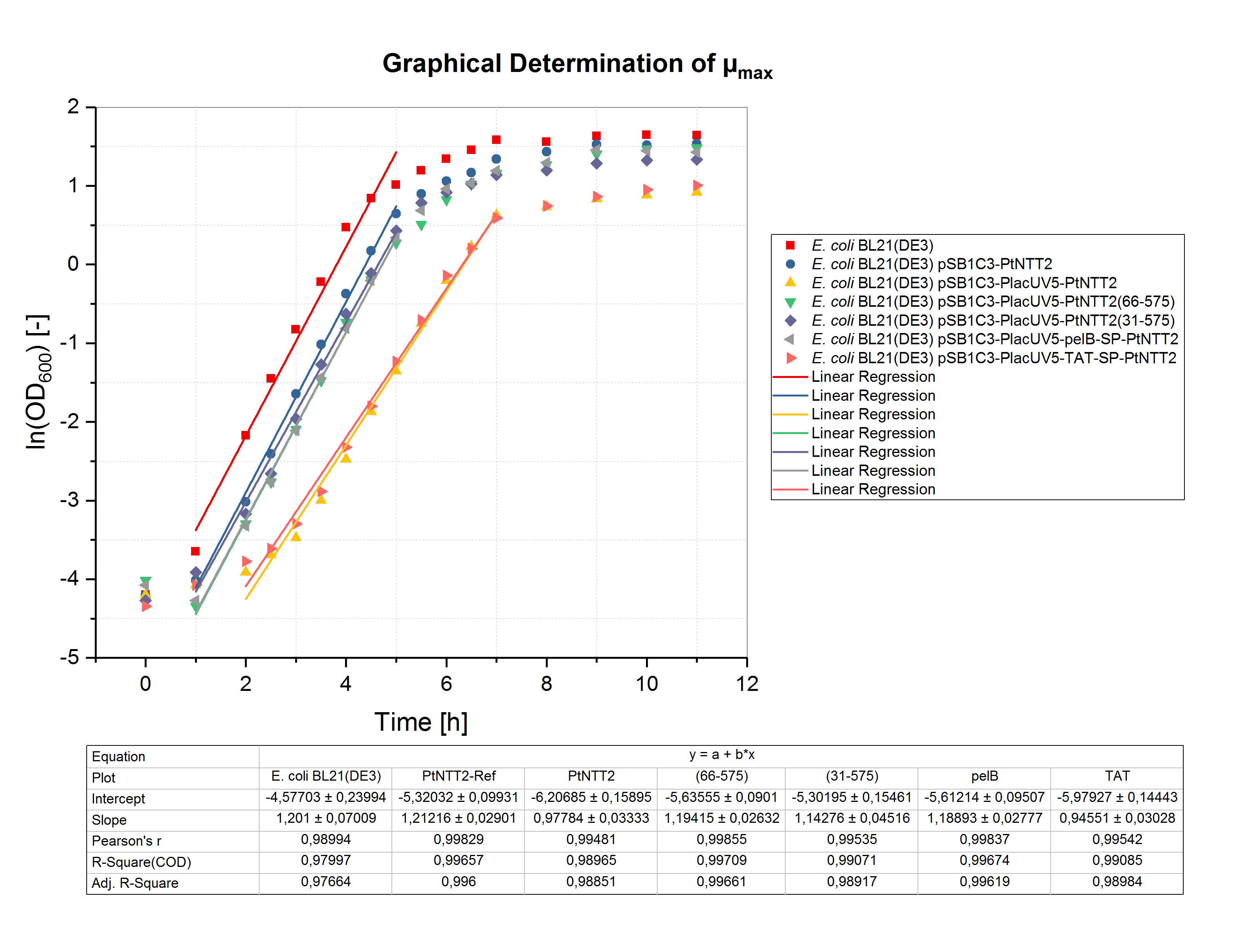
To determine the maximum specific growth rate (µmax), the natural logarithm of the OD600 values was plotted against the cultivation time. The slope of the linear regression through the exponential phase gives µmax. The graphical determination of µmax for the shake flask cultivation is shown in figure (4).Unsurprisingly, the highest specific growth rate can be observed for the negative controls E. coli BL21(DE3) and E. coli BL21(DE3) pSB1C3-PtNTT2 with values of 1.201 ± 0.070 h-1 and 1.212 ± 0.029 h-1, respectively. The lowest specific growth rate could be observed for E. coli BL21(DE3) pSB1C3-PlacUV5-TAT-SP-PtNTT2 with a value of 0.946 ± 0.030 h-1. Based on the specific growth rate, the minimal doubling time was calculated using the following equation:
The maximum specific growth rates and minimal doubling times are show in table (3) for all cultures.These results clearly show that expression of PtNTT2 leads to a reduced final cell density and slower growth. Furthermore, the different variants of PtNTT2 differ highly, indicating that some variants of PtNTT2 negatively affect the growth rate and final cell density.
Table (3): Maximum specific growth rates and minimum doubling times for all cultures.
| Strain | µmax [h-1] | td [h] |
|---|---|---|
| E. coli BL21(DE3) | 1.201 ± 0.070 | 0.577 ± 0.058 |
| E. coli BL21(DE3) pSB1C3-PtNTT2 | 1.212 ± 0.029 | 0.572 ± 0.024 |
| E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2 | 0.978 ± 0.033 | 0.709 ± 0.034 |
| E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2(66-575) | 1.194 ± 0.026 | 0.581 ± 0.022 |
| E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2(31-575) | 1.143 ± 0.045 | 0.606 ± 0.039 |
| E. coli BL21(DE3) pSB1C3-PlacUV5-pelB-SP-PtNTT2 | 1.189 ± 0.028 | 0.583 ± 0.024 |
| E. coli BL21(DE3) pSB1C3-PlacUV5-TAT-SP-PtNTT2 | 0.946 ± 0.030 | 0.733 ± 0.032 |
Microcultivations
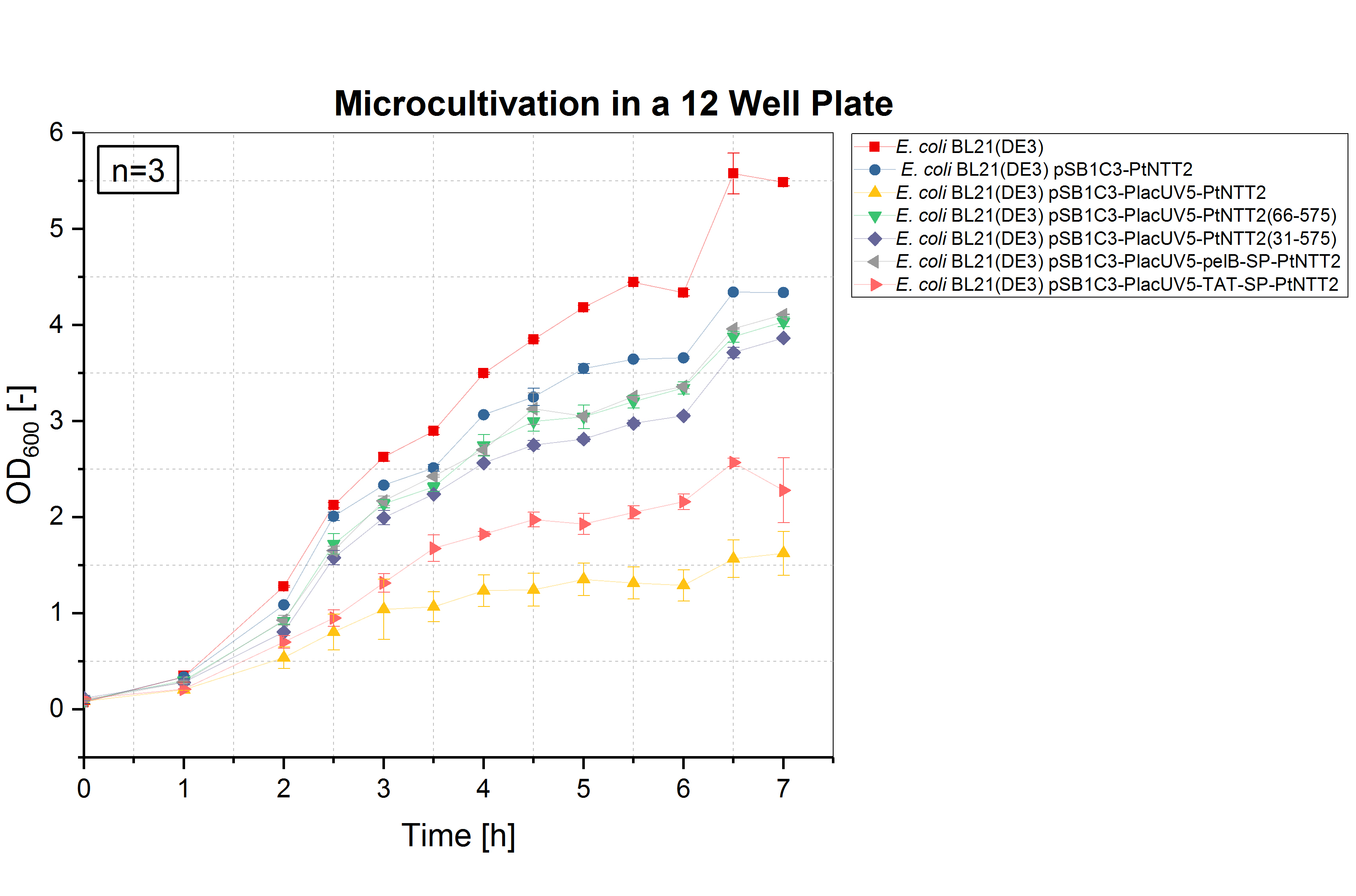
Due to the high price of unnatural nucleoside triphosphates, performing experiments such as cultivations in a small volume is desirable. Therefore, we validated whether cultivations in micro well plates can achieved the same results as cultivations performed in shake flasks and optimized the method. The optimal conditions for cultivations in a small volume were determined as follows: 12 well plates with a cultivation volume of 1 mL, 37 °C and 600 rpm. To analyze the growth of the differentPtNTT2 variants in a micro cultivation, the shake flask cultivation was repeated in a 12 well plate. All cultures were inoculated with an OD600 of 0.1. Three biological replicates of each strain were tested and three technical replicates were measured at each time point. The results are shown in figure (4).
The results of the microcultivations show the same growth pattern observed in the shake flask cultivations. E. coli BL21(DE3) and E. coli BL21(DE3) pSB1C3-PtNTT2 reached the highest ODs with 5.487 ± 0.038 and 4.337 ± 0.01, respectively. E. coli BL21(DE3) pSB1C3-PlacUV5-pelB-SP-PtNTT2 reached a final OD600 of 4.110 ± 0.005, followed by E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2(66-575) with a final OD600 of 4.035 ± 0.051. E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2(31-575) reached the third highest OD600 of allPtNTT2 variants with a final OD600 of 3.865 ± 0.008. The lowest final ODs were again reached by E. coli BL21(DE3) pSB1C3-PlacUV5-TAT-SP-PtNTT2 and E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2, reaching a final OD600 of 2.28 ± 0.337 and 1.623 ± 0.481. Therefore, we were able to demonstrate that the results of the shake flask cultivations can be transferred to a smaller format, such as a microcultivation in 1 mL. The reproduction of the results in a 50 times smaller volume was important for further experiments.
Table (4): Final OD600 of all cultures.
The highest OD600 was reached by the wildtype E. coli BL21(DE3) with 5,487 ± 0.038, the lowest by E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2 with 1.623 ± 0.481.
| Strain | Final OD600 [-] | |
|---|---|---|
| E. coli BL21(DE3) | 5.487 ± 0.038 | |
| E. coli BL21(DE3) pSB1C3-PtNTT2 | 4.337 ± 0.010 | |
| E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2 | 1.623 ± 0.481 | |
| E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2(66-575) | 4.035 ± 0.051 | |
| E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2(31-575) | 3.865 ± 0.008 | |
| E. coli BL21(DE3) pSB1C3-PlacUV5-pelB-SP-PtNTT2 | 4.110 ± 0.005 | |
| E. coli BL21(DE3) pSB1C3-PlacUV5-TAT-SP-PtNTT2 | 2.280 ± 0.337 |
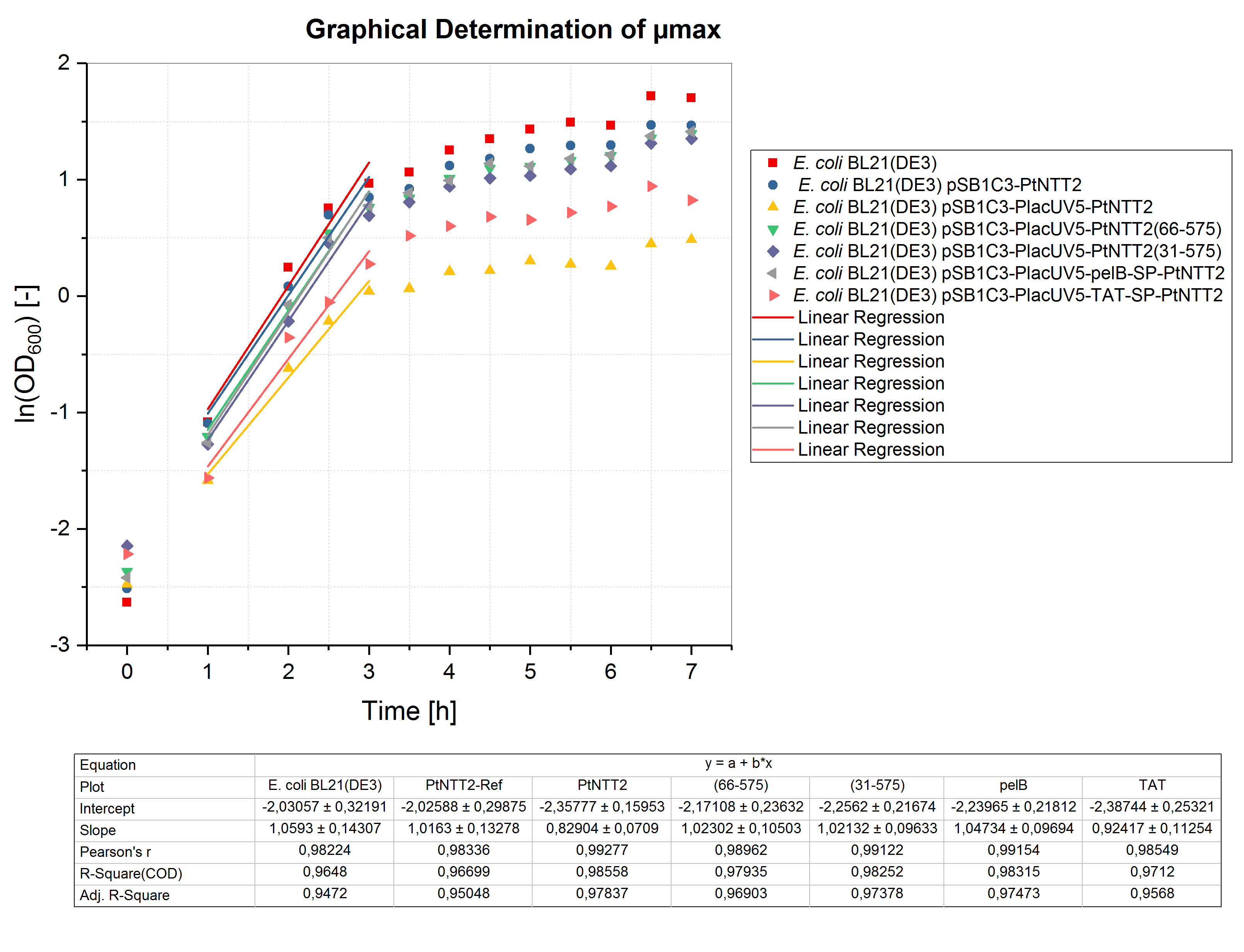
Like for the shake flask cultivation, µmax was determined graphically (figure 5). Bases on the obtained values, the minimum doubling time was calculated. The results are summarized in table (5).
All cultures show extended doubling times when compared to the shake flask cultivations. This is not surprising, since the oxygen transfer is much better in shake flasks compared to well plates. Nonetheless, similar final OD600 values were reached in the micro cultivations. Furthermore, the same differences in growth between the different strains can be observed, with E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2 and E. coli BL21(DE3) pSB1C3-PlacUV5-TAT-SP-PtNTT2 growing the weakest.
Table (5): Maximum specific growth rate and minimum doubling time for all cultures cultivated in 12 well plates.
| Strain | µmax [h-1] | td [h] |
|---|---|---|
| E. coli BL21(DE3) | 1.059 ± 0.143 | 0.655 ± 0.135 |
| E. coli BL21(DE3) pSB1C3-PtNTT2 | 1.016 ± 0.133 | 0.682 ± 0.131 |
| E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2 | 0.829 ± 0.071 | 0.836 ± 0.086 |
| E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2(66-575) | 1.023 ± 0.105 | 0.678 ± 0.103 |
| E. coli BL21(DE3) pSB1C3-PlacUV5-PtNTT2(31-575) | 1.021 ± 0.096 | 0.679 ± 0.094 |
| E. coli BL21(DE3) pSB1C3-PlacUV5-pelB-SP-PtNTT2 | 1.047 ± 0.097 | 0.662 ± 0.093 |
| E. coli BL21(DE3) pSB1C3-PlacUV5-TAT-SP-PtNTT2 | 0.924 ± 0.113 | 0.750 ± 0.122 |