Difference between revisions of "Part:BBa J61051"
Sherry-Xue (Talk | contribs) |
Sherry-Xue (Talk | contribs) |
||
| Line 12: | Line 12: | ||
Improved part from Peking iGEM 2013: https://parts.igem.org/Part:BBa_K1031610 | Improved part from Peking iGEM 2013: https://parts.igem.org/Part:BBa_K1031610 | ||
| + | |||
Improved part from WHU iGEM 2017:https://parts.igem.org/Part:BBa_K2462007 | Improved part from WHU iGEM 2017:https://parts.igem.org/Part:BBa_K2462007 | ||
Revision as of 15:05, 27 October 2017
[Psal1] J61051 pSB1A2 Bca9015 Salicylate promoter
(Edited by Zheng Pu, Peking iGEM 2013)

(Please visit: http://2013.igem.org/Team:Peking/Project/BioSensors/NahR for more details)
Improved part from Peking iGEM 2013: https://parts.igem.org/Part:BBa_K1031610
Improved part from WHU iGEM 2017:https://parts.igem.org/Part:BBa_K2462007
Introduction
The nahR gene was mined from the 83 kb naphthalene degradation plasmid NAH7 of Pseudomonas putida, encoding a 34 kDa protein which binds to nah and sal promoters to activate transcription in response to the inducer salicylate. This plasmid encodes enzymes for the metabolism of naphthalene or salicylate as the sole carbon and energy source (Fig.1a) [1]. The 14 genes encoding the enzymes for this metabolism are organized in two operons: nah (nahA-F), encoding six enzymes required for metabolizing naphthalene into salicylate and pyruvate, and sal (nahG-M), encoding eight enzymes which metabolize salicylate into intermediates of TCA cycle (Fig.1b) [2].
We also did adaptor for this biosensor to improve its performance. (for more detail:http://2013.igem.org/Team:Peking/Project/Plugins)

Figure.1 Degradation pathway of naphthalene and the corresponding gene cluster in Pseudomonas putida. (a) Gene cluster on the NAH7 plasmid that degrades naphthalene: Naphthalene is degraded into salicylate by enzymes encoded by the "upper operon"; salicylate is further degraded to enter TCA cycle via the gene products of the "lower operon". Both of the operons are regulated by the transcription factor NahR in response to salicylate, the metabolite intermediate in the pathway. (b) Degradation pathway of naphthalene: Naphthalene is degraded by a series of enzymatic reactions, each catalyzed by a specific nah gene product represented by a capital letter. A through M: A, Naphthalene dioxygenase; B, cis-dihydroxy-naphthalene dioxygenase; D, 2-hydroxychromene-2-carboxylate isomerase; E, 2-hydroxybenzalpyruvate aldolase; F, salicylaldehyde dehydrogenase; G, salicylate hydroxylase; H, catechol 2,3-dioxygenase; I, 2-hydroxymuconate semialdehyde dehydrogenase; J, 2-hydroxymuconate tautomerase; K, 4-oxalcrotonate decarboxylase; L, 2-oxo-4-pentenoate hydratase; M, 2-oxo-4-hydroxypentanoate aldolase.
Protein Domain Structure

Figure.2 The organization of NahR protein domains. The domain marked by green near the N terminal accounts for DNA binding, which contains a typical helix-turn-helix motif; red domains function to bind inducer, while the orange domain is putatively involved in multimerization of NahR during the transcription activation.
Promoter Structure

Figure.3 Schematic diagram for the NahR-regulated promoters, nah and sal. Alignment of sal and nah promoter is shown and the consensus sequence motifs are highlighted in color. NahR binding sequence and RNAP binding sequence are boxed in green and yellow, respectively.
Mechanism
Several experiments conformed that NahR tightly binds to DNA in vivo in the presence or absence of salicylate. Either the amount or the affinity of NahR binding to DNA will be affected by salicylate in both E. coli and its native host Pseudomonas putida[3]. This along with the evidence from methylation protection experiments suggested a conformational change in the NahR•DNA complex before transcription activation (Fig.4)[4].

Figure.4 Schematic diagram for the transcription activation at sal (or nah) promoter by NahR in the presence of inducer salicylate. 1. The DNA structure of sal promoter: A, B, C and D represent the binding sites for the tetramer of NahR; the yellow arrow shows the direction of sal promoter. 2. RNAP and σ70 bind to the sal promoter by recognizing -35 and -10 boxes; 3. Transcription factor NahR tightly binds to sal promoter and forms a tetramer no matter whether there is salicylate or not; 4. When salicylate is present, NahR•DNA complex undergoes a conformational change. After the hydrolysis of ATP, DNA is opened and transcription is activated.
Circuit Construction
we constructed a PSal/NahR biosensor circuit using sfGFP as reporter gene (Fig.5). NahR is constitutively expressed by a constitutive promoter (Pc), and the expression of sfGFP is positively regulated by NahR in the presence of inducers.

Figure.5 Schematic diagram for the NahR biosensor circuit. The Biobrick BBa_J61051 was cloned preceding reporter sfGFP in the backbone pSB1C3. Promoters are presented in orange, RBS in light green, coding sequence in dark blue and terminators in red.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 786
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 77
Illegal NgoMIV site found at 618 - 1000COMPATIBLE WITH RFC[1000]
Characterization of Sensor
On-Off Tests

Figure.6 ON/OFF test to evaluate the induction ratios of all aromatic compounds in the aromatics spectrum. (For the full names of the compounds, Click Here). (a) The induction ratioS of various aromatic species. NahR could respond to 18 out of 78 aromatics with the induction ratio over 20. (b) The aromatics-sensing profile of NahR biosensor.The aromatic species that can elicit strong responses of NahR biosensor are highlighted in green in the aromatics spectrum. The structure formula of typical inducer is also listed around the spectrum. The induction ratio was calculated by dividing the fluorescence intensity of biosensor exposed to object inducers by the basal fluorescence intensity of the biosensor itself.
Dose-response Curve

Figure.7 Dose response curves of NahR biosensor. (a) Dose response curves for salicylate, its homologs and derivatives; (b) Dose response curves for benzoate, its derivatives and special inducers like 5-ClSaD and 2,4,6-TClPhl. The induction ratio was calculated by dividing the fluorescence intensity of biosensor exposed to inducers by the basal fluorescence intensity of the biosensor. For the full names of the compounds, Click Here.
Orthogonality of Different Sensor
If the presence of an inducer of biosensor A (not an inducer of biosensor B) doesn’t interfere with the dose response of biosensor B to any of its inducers, and vice versa, we call the B and A biosensors are "orthogonal"; namely, no synergistic/antagonistic effects happen between the inducers of A and B biosensors.(for more details, Chick Here)

| Sensor | Host | Main Inducers |
|---|---|---|
| XylS | Pseudomonas putida | BzO 2-MeBzO 3-MeBzO 2,3-MeBzO 3,4-MeBzO |
| NahR | Pseudomonas putida | 4-MeSaA 4-C1SaA 5-C1SaA SaA Aspirin |
| DmpR | Pseudomonas sp.600 | Phl 2-MePhl 3-MePhl 4-MePhl 2-ClPhl |
| HbpR | Pseudomonas azelaica | o-Phenylphenol 2,6'-DiHydroxybiphenol |
Figure.8 Summary of the orthogonality assay to evaluate the synergistic/antagonistic effects between the inducers of 4 representative biosensors. No synergistic or antagonistic effects between the inducers of 4 representative biosensors (XylS, NahR, HbpR, and DmpR) were observed. For instance, although the sensing profiles of NahR and XylS overlap to some extent, the NahR-specific and XylS-specific inducers proved to be really orthogonal.
We have confirmed the orthogonality among inducers of different biosensors, which is one of the main features we expect for our aromatics-sensing toolkit; this allowed the combination of these biosensors to profile aromatics for the ease of practical applications.
Related Parts:
XylS: https://parts.igem.org/Part:BBa_K1031911 Wiki: http://2013.igem.org/Team:Peking/Project/BioSensors/XylS
NahR: https://parts.igem.org/Part:BBa_K1031610 Wiki: http://2013.igem.org/Team:Peking/Project/BioSensors/NahR
HbpR: https://parts.igem.org/Part:BBa_K1031300 Wiki: http://2013.igem.org/Team:Peking/Project/BioSensors/HbpR
DmpR: http://2013.igem.org/Team:Peking/Project/BioSensors/DmpR
We examined the orthogonality between 4 representative biosensors (Fig.9). The orthogonality test between two biosensors, biosensor I and biosensor II, was performed in the following procedure:
1. A typical inducer A for biosensor I and a typical inducer B for biosensor II were selected.
2. The dose response of biosensor I to inducer A was measured, under the perturbation of inducer B.
3. The dose-response of biosensor II to inducer B was measured, under the perturbation of inducer A.
If biosensor I and biosensor II are orthogonal, the dose response of biosensor I to inducer A should be constant, regardless of the concentrations of inducer B; and the dose response of biosensor II to inducer B should be constant, regardless of the concentrations of inducer A. Namely, for two "orthogonal" biosensors, the perturbation of an unrelated inducer has negligible effect on the dose response of a biosensor to its related inducer (Fig.10).

Figure.9 Orthogonality test assay for inducer A (detected by biosensor I) and inducer B (detected by biosensor II). (a) Biosensor I was added into the test assay. Different mixtures of inducers were added into lane 1, 2, and 3 respectively as listed above. Effect of inducer B upon the dose-response curve of inducer A was tested by comparing the fluorescence intensity of biosensor I among lane 1 ,2, and 3. (b) Biosensor II was added into the test assay. Different mixtures of inducers were added into lane 1, 2, and 3 respectively as listed above. Effect of inducer A upon the dose-response curve of inducer B was tested by comparing the fluorescence intensity of biosensor II among lane 1 ,2, and 3.

Figure.10 Correlation of the inducer B and the dose-response of biosensor I to its inducer A. Each point on the right plot represents a concentration of inducer A. It's x coordinate represents the fluorescence when inducer B is 0 and the y coordinate represents the fluorescence when the cell is exposed to a none-zero concentration of inducer B. If the dose-response of biosensor I is invariant to the concentration of inducer B, the x coordinate of a experimental point should be equal to its y coordinate and the experimental points are supposed to be aligned in a line whose slope is one.
The orthogonality between XylS, NahR, HbpR and DmpR biosensors have been carefully evaluated using the assay discussed above (Fig.9). The data were processed by linear fitting and the slopes of the fitting curves were compared with 1 (Fig.9, Fig.10). The closer the slope was to 1, the more orthogonal the two biosensors were. Results showed that the biosensor pairs, XylS and NahR (Fig.11a, b), XylS and HbpR (Fig.11c, d), NahR and HbpR (Fig.11e, f), XylS and DmpR (Fig.11g, h), NahR and DmpR (Fig.11i, j), and HbpR and DmpR (Fig.11k, l) are all orthogonal, as summarized in Fig.8.






Figure.11 Linear fitting of the data obtained from the orthogonality assay showing that the orthogonality between the 4 representative biosensors. The experiments and data processing were performed as described in Fig.9 and Fig.10.The black dashed line denotes slope=1 as the reference line. These fittings showed the orthogonality between biosensors, (a, b) XylS and NahR; (c, d) XylS and HbpR; (e, f) NahR and HbpR, (g, h) XylS and DmpR, (i, j) NahR and DmpR, and (k, l) HbpR and DmpR. The experiment data, linear fitting curves of biosensor, and cognate inducers are in different colors: XylS in red, NahR in green, HbpR in orange and DmpR in dark cyan.
Characterization of the Inducible Promoter

The inducible promoter was induced with different concentration(0, 1.50E-07, 6.10E-07, 2.40E-06, 9.80E-06, 3.90E-05, 1.60E-04, 6.20E-04, 1.00E-03) every 30 minutes. We started the induction when the OD600 was 0.6-0.7. The fluorescence response was measured with plate reader.

The inducible promoter was induced with different concentration(0, 1.50E-07, 6.10E-07, 2.40E-06, 9.80E-06, 3.90E-05, 1.60E-04, 6.20E-04, 1.00E-03) every 30 minutes. We started the induction when the OD600 was 0.6-0.7. The fluorescence response was measured with plate reader. The red line indicate the Hill equation(P = Pmax*[SA]^n/([SA]^n+Km^n) ) fit of our dose-response data with error bar showing the standard deviation and the R square value of this fit is 0.99531. "n" indicates the hill coefficient(unitless), "Pmax" indicates the maximal promoter activity, and "K" is the salicylic acid concentration while P equals 1/2*Pmax
Characterization of the Inducible Promoter
The Uppsala igem team characterize this part as an inducible promoter with the RFP dTomato as a reporter.
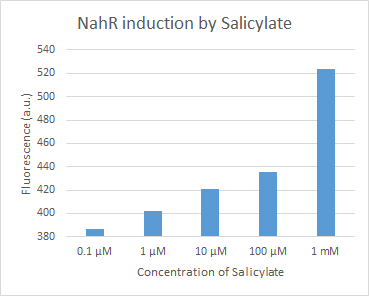
Figure 14. dTomato expressed in E. coli DH5-alpha. NahR/Psal induction test in LB medium. Concentration of salicylate from left to right: 0,1 μM, 1 μM, 10 μM, 100 μM and 1 mM.
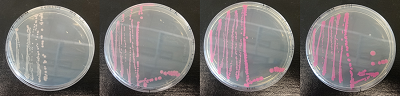
Figure 15. dTomato expressed in E. coli DH5-alpha. NahR/Psal induction test on agar plate. Concentration of salicylate from left to right: 0,1 μM, 1 μM, 10 μM, 100 μM and 1 mM.
We tested the NahR/Psal promoter system with dTomato on a pSB1C3 plasmid. The cells containing the plasmid were streaked on agar plates with different salicylate concentrations in the agar medium. Similarly, we also examined liquid cultures with different concentrations of salicylate in the solution. Both of these tests showed that the bacteria were producing the dTomato protein in presence of salicylate, independent of the salicylate being in liquid culture or in the solid growth medium. It can also be easily observed that the fluorescence increased with increasing salicylate concentrations (See figure 14 and 15). The measurements were made with the fluorometer that our team built in order to be able to characterize this system. These results were compared to MACS measurements on the same colonies and the measurements were correlated.
Reference
[1] Dunn, N. W., and I. C. Gunsalus.(1973) Transmissible plasmid encoding early enzymes of naphthalene oxidation in Pseudomonas putida. J. Bacteriol. 114:974-979 [2] M. A. Schell.(1983) Cloning and expression in Escherichia coli of the naphthalene degradation genes from plasmid NAH7. J. Bacteriol. 153(2):822 [3] M. A. Schell, and E. F. Poser.(1989) Demonstration, characterization, and mutational analysis of NahR protein binding to nah and sal promoters. J. Bacteriol. 171(2):837 [4] Jianzhong Huang and Mark A. Schell.(1991) In vivo interaction of the NahR Transcriptional Activator with its target sequences. The Journal of Biological Chemistry. 266(17):10830-10838