Difference between revisions of "Part:BBa K1980010"
| Line 11: | Line 11: | ||
==Description== | ==Description== | ||
<p>This part contains the bacterial copper chelator Csp1 (Copper Storage Protein 1) with a C-terminal sfGFP tag (connected with short hydrophilic flexible linker) behind the pCopA promoter with constitutively expressed CueR (a bacterial copper sensitive repressor/activator) in the biobrick. </p> | <p>This part contains the bacterial copper chelator Csp1 (Copper Storage Protein 1) with a C-terminal sfGFP tag (connected with short hydrophilic flexible linker) behind the pCopA promoter with constitutively expressed CueR (a bacterial copper sensitive repressor/activator) in the biobrick. </p> | ||
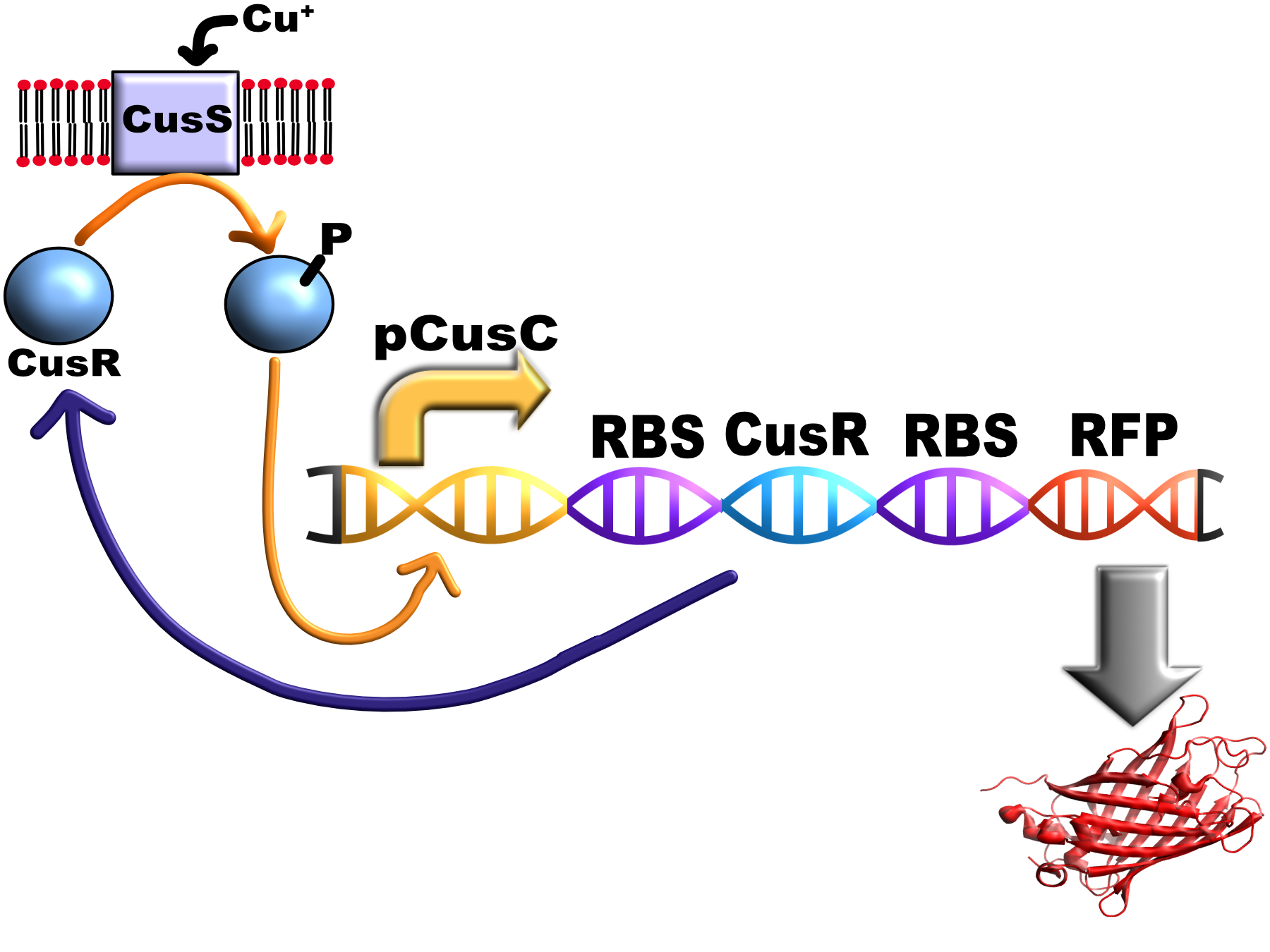
| + | [[Image:T--Oxford--FCK_diagram_v2.jpg|400px|thumb|center|Schematic of the system]] | ||
<!-- --> | <!-- --> | ||
| Line 16: | Line 17: | ||
<p><i>E. coli</i> cells use a protein called CueR to regulate the cytoplasmic copper concentration. CueR is a MerR-type regulator with an interesting mechanism of action whereby it can behave as a net activator or a net repressor under different copper concentrations through interaction with RNA polymerase<sup>(1)</sup>. CueR forms dimers consisting of three functional domains (a DNA-binding, a dimerisation and a metal-binding domain). The DNA binding domains bind to DNA inverted repeats called CueR boxes with the sequence:</p> | <p><i>E. coli</i> cells use a protein called CueR to regulate the cytoplasmic copper concentration. CueR is a MerR-type regulator with an interesting mechanism of action whereby it can behave as a net activator or a net repressor under different copper concentrations through interaction with RNA polymerase<sup>(1)</sup>. CueR forms dimers consisting of three functional domains (a DNA-binding, a dimerisation and a metal-binding domain). The DNA binding domains bind to DNA inverted repeats called CueR boxes with the sequence:</p> | ||
| − | + | ||
CCTTCCNNNNNNGGAAGG | CCTTCCNNNNNNGGAAGG | ||
| − | + | ||
<p>This box is present at the promoter regions of the copper exporting ATPase CopA, some molybdenum cofactor synthesis genes and the periplasmic copper oxidase protein CueO.<sup>(2)</sup></p> | <p>This box is present at the promoter regions of the copper exporting ATPase CopA, some molybdenum cofactor synthesis genes and the periplasmic copper oxidase protein CueO.<sup>(2)</sup></p> | ||
| Line 26: | Line 27: | ||
<p>Likely due to the expression problems of the Csp1 chelator in <i>E. coli</i>, it responded worse than the similar part with our MymTsfGFP fusion protein (https://parts.igem.org/Part:BBa_K1980012)</p> | <p>Likely due to the expression problems of the Csp1 chelator in <i>E. coli</i>, it responded worse than the similar part with our MymTsfGFP fusion protein (https://parts.igem.org/Part:BBa_K1980012)</p> | ||
| + | |||
| + | [[Image:T--Oxford--FCKDATA.jpeg|400px|thumb|center|Absolute fluorescence for the construct at four hours at various copper concentrations.]] | ||
==References== | ==References== | ||
Revision as of 15:39, 20 October 2016
pCopA TAT Csp1 sfGFP with divergent CueR
pCopA TAT Csp1 sfGFP with divergent expressed CueR
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 438
Illegal NheI site found at 461 - 21INCOMPATIBLE WITH RFC[21]Illegal XhoI site found at 1466
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 1028
- 1000COMPATIBLE WITH RFC[1000]
Description
This part contains the bacterial copper chelator Csp1 (Copper Storage Protein 1) with a C-terminal sfGFP tag (connected with short hydrophilic flexible linker) behind the pCopA promoter with constitutively expressed CueR (a bacterial copper sensitive repressor/activator) in the biobrick.
Usage and Biology
E. coli cells use a protein called CueR to regulate the cytoplasmic copper concentration. CueR is a MerR-type regulator with an interesting mechanism of action whereby it can behave as a net activator or a net repressor under different copper concentrations through interaction with RNA polymerase(1). CueR forms dimers consisting of three functional domains (a DNA-binding, a dimerisation and a metal-binding domain). The DNA binding domains bind to DNA inverted repeats called CueR boxes with the sequence:
CCTTCCNNNNNNGGAAGG
This box is present at the promoter regions of the copper exporting ATPase CopA, some molybdenum cofactor synthesis genes and the periplasmic copper oxidase protein CueO.(2)
Experience
We cloned pCopA TAT Csp1 sfGFP with divergent expressed CueR from a gBlock into the shipping plasmid pSB1C3. E. coli strain MG1655 was transformed using the specific recombinant plasmid and a 5ml culture of a transformed colony was grown overnight. A plate with four repeats at 10 different copper concentrations (ranging from 0mM to 2mM) plus a negative control was made. The absorbance and fluorescence of each well was measured over time in order to assess the absolute fluorescence of the construct.
Likely due to the expression problems of the Csp1 chelator in E. coli, it responded worse than the similar part with our MymTsfGFP fusion protein (https://parts.igem.org/Part:BBa_K1980012)
References
(1) Danya J. Martell, Chandra P. Joshi, Ahmed Gaballa, Ace George Santiago, Tai-Yen Chen, Won Jung, John D. Helmann, and Peng Chen (2015) “Metalloregulator CueR biases RNA polymerase’s kinetic sampling of dead-end or open complex to repress or activate transcription” Proc Natl Acad Sci U S A. 2015 Nov 3; 112(44): 13467–13472.
(2) Yamamoto K, Ishihama A. (2005) “Transcriptional response of Escherichia coli to external copper.” Mol Microbiol. 2005 Apr;56(1):215-27.