Difference between revisions of "Part:BBa K1639000"
| Line 21: | Line 21: | ||
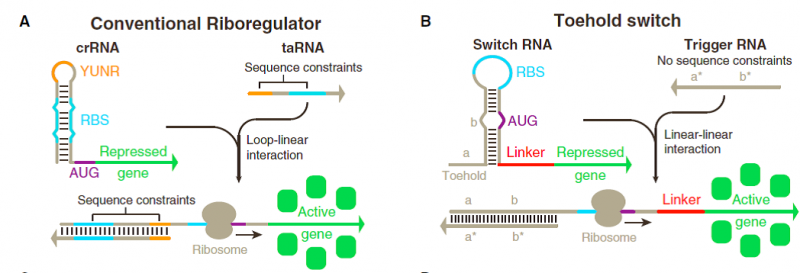
Toehold switch systems are composed of two RNA strands referred to as the switch and trigger (Figure 1B). The switch RNA contains the coding sequence of the gene being regulated. Upstream of this coding sequence is a hairpin-based processing module containing both a strong RBS and a start codon that is followed by a common 21 nt linker sequence coding for low-molecular-weight amino acids added to the N terminus of the gene of interest. A single-stranded toehold sequence at the 50 end of the hairpin module provides the initial binding site for the trigger RNA strand. This trigger molecule contains an extended single stranded region that completes a branch migration process with the hairpin to expose the RBS and start codon, thereby initiating translation of the gene of interest. The hairpin processing unit functions as a repressor of translation in the absence of the trigger strand. Unlike previous riboregulators, the RBS sequence is left completely unpaired within the 11 nt loop of the hairpin. Instead, the bases immediately before and after the start codon are sequestered within RNA duplexes that are 6 bp and 9 bp long, respectively. The start codon itself is left unpaired in the switches we tested, leaving a 3 nt bulge near the midpoint of the 18 nt hairpin stem. Because the repressing domain b (Figure 1B) does not possess complementary bases to the start codon, the cognate trigger strand in turn does not need to contain corresponding start codon bases, thereby increasing the number of potential trigger sequences. The sequence in the hairpin added after the start codon was also screened for the presence of stop codons, as they would prematurely terminate translation of the gene of interest when the riboregulator was activated. We employed a 12 nt Toehold domain at the 50 end of the hairpin to initiate its interaction with the cognate trigger strand. The trigger RNA contains a 30 nt single-stranded RNA sequence that is complementary to the toehold and stem of the switch RNA. | Toehold switch systems are composed of two RNA strands referred to as the switch and trigger (Figure 1B). The switch RNA contains the coding sequence of the gene being regulated. Upstream of this coding sequence is a hairpin-based processing module containing both a strong RBS and a start codon that is followed by a common 21 nt linker sequence coding for low-molecular-weight amino acids added to the N terminus of the gene of interest. A single-stranded toehold sequence at the 50 end of the hairpin module provides the initial binding site for the trigger RNA strand. This trigger molecule contains an extended single stranded region that completes a branch migration process with the hairpin to expose the RBS and start codon, thereby initiating translation of the gene of interest. The hairpin processing unit functions as a repressor of translation in the absence of the trigger strand. Unlike previous riboregulators, the RBS sequence is left completely unpaired within the 11 nt loop of the hairpin. Instead, the bases immediately before and after the start codon are sequestered within RNA duplexes that are 6 bp and 9 bp long, respectively. The start codon itself is left unpaired in the switches we tested, leaving a 3 nt bulge near the midpoint of the 18 nt hairpin stem. Because the repressing domain b (Figure 1B) does not possess complementary bases to the start codon, the cognate trigger strand in turn does not need to contain corresponding start codon bases, thereby increasing the number of potential trigger sequences. The sequence in the hairpin added after the start codon was also screened for the presence of stop codons, as they would prematurely terminate translation of the gene of interest when the riboregulator was activated. We employed a 12 nt Toehold domain at the 50 end of the hairpin to initiate its interaction with the cognate trigger strand. The trigger RNA contains a 30 nt single-stranded RNA sequence that is complementary to the toehold and stem of the switch RNA. | ||
| + | |||
| + | '''Sources:''' | ||
| + | |||
| + | [1]: Montecucco C, Rappuoli R. Living dangerously: how Helicobacter pylori survives in the human stomach. Nat Rev Mol Cell Biol. 2001;2(6):457-66. | ||
| + | |||
| + | [2]: Vendeville A, Winzer K, Heurlier K, Tang CM, Hardie KR. Making 'sense' of metabolism: autoinducer-2, LuxS and pathogenic bacteria. Nat Rev Microbiol. 2005;3(5):383-96. | ||
| + | |||
| + | [3]: Loh JT, Forsyth MH, Cover TL. Growth phase regulation of flaA expression in Helicobacter pylori is luxS dependent. Infect Immun. 2004;72(9):5506-10. | ||
| + | |||
| + | [4]: Brown SW, Sonenshein AL. Autogenous regulation of the Bacillus subtilis glnRA operon. J Bacteriol. 1996;178(8):2450-4. | ||
| + | |||
| + | [5]: Commichau FM, Wacker I, Schleider J, et al. Characterization of Bacillus subtilis mutants with carbon source-independent glutamate biosynthesis. J Mol Microbiol Biotechnol. 2007;12(1-2):106-13. | ||
| + | |||
| + | [6]: Wray LV, Zalieckas JM, Fisher SH. Bacillus subtilis glutamine synthetase controls gene expression through a protein-protein interaction with transcription factor TnrA. Cell. 2001;107(4):427-35. | ||
| + | |||
| + | [7]: Yoshida K, Yamaguchi H, Kinehara M, Ohki YH, Nakaura Y, Fujita Y. Identification of additional TnrA-regulated genes of Bacillus subtilis associated with a TnrA box. Mol Microbiol. 2003;49(1):157-65. | ||
| + | |||
| + | [8]: Ahmer BM. Cell-to-cell signalling in Escherichia coli and Salmonella enterica. Mol Microbiol 2004; 52:933-945 | ||
| + | |||
| + | [9]: Federle MJ, Bassler BL. Interspecies communication in bacteria. J Clin Invest 2003; 112:1291-1299. | ||
| + | |||
| + | [10]: Vendeville A, Winzer K, Heurlier K, Tang CM, Hardie KR. Making ‘sense’ of metabolism: autoinducer-2, LuxS and pathogenic bacteria. Nat Rev Microbiol 2005; 3:383-396. | ||
| + | |||
| + | [11]: Taga ME, Miller ST, Bassler BL. Lsr-mediated transport and processing of AI-2 in Salmonella typhimurium. Mol Microbiol 2003; 50:1411-1427. | ||
| + | |||
| + | [12]: Xavier KB, Miller ST, Lu W, et al. Phosphorylation and processing of the quorum-sensing molecule autoinducer-2 in enteric bacteria. ACS Chem Biol 2007; 2:128-136. | ||
| + | |||
| + | [13]: Forsyth, M. H., and T. L. Cover. 2000. Intercellular communication in Helicobacter pylori: luxS is essential for the production of an extracellular signaling molecule. Infect. Immun. 68:3193–3199. | ||
| + | |||
| + | [14]: Joyce, E. A., B. L. Bassler, and A. Wright. 2000. Evidence for a signaling system in Helicobacter pylori: detection of a luxS-encoded autoinducer. J. Bacteriol. 182:3638–3643 | ||
| + | |||
| + | [15]:Green, A., Silver, P., Collins, J., & Yin, P. (n.d.). Toehold Switches: De-Novo-Designed Regulators of Gene Expression. Cell, 925-939. | ||
<span class='h3bb'>Sequence and Features</span> | <span class='h3bb'>Sequence and Features</span> | ||
Revision as of 20:15, 20 September 2015
Toehold-GFP
Toehold-GFP reporter device. This part produces reporter flourescence protein GFP, only when a specific mRNA present in the cell called Trigger RNA(BBa_K1639001).
Usage and Biology
Toehold switches provide a high level of orthogonality and can be forward engineered to provide average dynamic range above 400. Toehold switches, with their wide dynamic range, orthogonality, and programmability, represent a versatile and powerful platform for regulation of translation, offering diverse applications in molecular biology, synthetic biology, and biotechnology.New classes of regulatory components that offer wide dynamic range, low system crosstalk, and design flexibility represent a much-needed, enabling step toward fully realizing the potential of synthetic biology in areas such as biotechnology and medicine. (Khalil and Collins, 2010).
Engineered riboregulators consist of cognate pairs of RNAs: a transducer strand that regulates translation or transcription and a trans-acting RNA that binds to the transducer to modulate its biological activity. Riboregulator designs can be classified according to the initial RNA-RNA interaction that drives hybridization between the transducer and trans-acting RNAs. Reactions initiated between loop sequences in both RNAs are termed loop-loop interactions, whereas those that occur between a loop sequence and an unstructured RNA are termed loop-linear(Takahashi and Lucks, 2013).
A common limitation for riboregulators has been their dynamic range (Liu et al., 2012). Previous prokaryotic translational riboregulators have typically modulated biological signals by up to a maximum of 55-fold for activators (Callura et al., 2012) and up to 10-fold for repressors (Mutalik et al., 2012). In contrast, protein-based transcriptional regulators have demonstrated dynamic ranges over an order of magnitude higher, with widely-used inducible promoters regulating protein expression over 350-fold (Lutz and Bujard, 1997) and sigma factor-promoter pairs providing up to 480-fold modulation (Rhodius et al., 2013).Despite the inherent programmability of RNA-based systems, efforts at constructing large sets of orthogonal riboregulators have been limited to libraries of at most seven parts with crosstalk levels of 20% (Takahashi and Lucks, 2013). Typical RNA-based regulators employ interaction domains consisting of30 nts, which corresponds to a sequence space of over 1018 potential regulatory elements. Thus, the sheer diversity of possible RNA-based parts suggests that previous devices have not come close to realizing the potential of highly orthogonal regulation.
Much of this discrepancy arises from the significant sequence constraints imposed on riboregulators engineered thus far (Figure1A). Like natural riboregulators, engineered riboregulators of translation have invariably used base pairing to the ribosome binding site (RBS) to prevent ribosome binding, thereby preventing translation (Callura et al., 2012; Isaacs et al., 2004; Mutaliket al., 2012; Rodrigo et al., 2012). Because repression is caused by RBS binding, trigger RNAs that activate translation are engineered to contain an RBS sequence to displace the repressing sequence, which in turn reduces the potential sequence space for the riboregulator.
Previous riboregulators have also relied on U-turn loop structures to drive loop-loop and loop-linear interactions between RNAs (Figure 1A) (Callura et al., 2012; Isaacs et al., 2004; Luckset al., 2011; Takahashi and Lucks, 2013). U-turn loops are common RNA structural motifs formed by tertiary interactions that have been identified in ribozymes, ribosomal RNAs, and transfer RNA anticodon loops (Gutell et al., 2000). Although recent work has begun to show that loops with canonical U-turn sequences are not essential for riboregulators (Mutalik et al., 2012; Rodrigoet al., 2012), the engineered systems reported to date have continued their reliance on the loop-mediated RNA interactions from natural systems. Although these loop interactions have been selected by evolution in nature, alternative approaches employing linear-linear RNA interactions are amenable to rational engineering and exhibit more favorable reaction kinetics and thermodynamics, factors that could be exploited to increase riboregulator dynamic range.
A riboregulator that activates gene expression must switch from a secondary structure that prevents translation to a configuration that promotes translation upon binding of a cognate trans-acting RNA. Although the Shine-Dalgarno sequence is an important factor in determining the efficiency of translation from a given mRNA, studies have found that secondary structure in regions near by the start codon also plays a critical role (Kudla et al., 2009).Furthermore, genome-wide analyses have revealed strong biases toward low secondary structures around the start codon of mRNAs from a panel of hundreds of bacterial genomes.
Toehold switch systems are composed of two RNA strands referred to as the switch and trigger (Figure 1B). The switch RNA contains the coding sequence of the gene being regulated. Upstream of this coding sequence is a hairpin-based processing module containing both a strong RBS and a start codon that is followed by a common 21 nt linker sequence coding for low-molecular-weight amino acids added to the N terminus of the gene of interest. A single-stranded toehold sequence at the 50 end of the hairpin module provides the initial binding site for the trigger RNA strand. This trigger molecule contains an extended single stranded region that completes a branch migration process with the hairpin to expose the RBS and start codon, thereby initiating translation of the gene of interest. The hairpin processing unit functions as a repressor of translation in the absence of the trigger strand. Unlike previous riboregulators, the RBS sequence is left completely unpaired within the 11 nt loop of the hairpin. Instead, the bases immediately before and after the start codon are sequestered within RNA duplexes that are 6 bp and 9 bp long, respectively. The start codon itself is left unpaired in the switches we tested, leaving a 3 nt bulge near the midpoint of the 18 nt hairpin stem. Because the repressing domain b (Figure 1B) does not possess complementary bases to the start codon, the cognate trigger strand in turn does not need to contain corresponding start codon bases, thereby increasing the number of potential trigger sequences. The sequence in the hairpin added after the start codon was also screened for the presence of stop codons, as they would prematurely terminate translation of the gene of interest when the riboregulator was activated. We employed a 12 nt Toehold domain at the 50 end of the hairpin to initiate its interaction with the cognate trigger strand. The trigger RNA contains a 30 nt single-stranded RNA sequence that is complementary to the toehold and stem of the switch RNA.
Sources:
[1]: Montecucco C, Rappuoli R. Living dangerously: how Helicobacter pylori survives in the human stomach. Nat Rev Mol Cell Biol. 2001;2(6):457-66.
[2]: Vendeville A, Winzer K, Heurlier K, Tang CM, Hardie KR. Making 'sense' of metabolism: autoinducer-2, LuxS and pathogenic bacteria. Nat Rev Microbiol. 2005;3(5):383-96.
[3]: Loh JT, Forsyth MH, Cover TL. Growth phase regulation of flaA expression in Helicobacter pylori is luxS dependent. Infect Immun. 2004;72(9):5506-10.
[4]: Brown SW, Sonenshein AL. Autogenous regulation of the Bacillus subtilis glnRA operon. J Bacteriol. 1996;178(8):2450-4.
[5]: Commichau FM, Wacker I, Schleider J, et al. Characterization of Bacillus subtilis mutants with carbon source-independent glutamate biosynthesis. J Mol Microbiol Biotechnol. 2007;12(1-2):106-13.
[6]: Wray LV, Zalieckas JM, Fisher SH. Bacillus subtilis glutamine synthetase controls gene expression through a protein-protein interaction with transcription factor TnrA. Cell. 2001;107(4):427-35.
[7]: Yoshida K, Yamaguchi H, Kinehara M, Ohki YH, Nakaura Y, Fujita Y. Identification of additional TnrA-regulated genes of Bacillus subtilis associated with a TnrA box. Mol Microbiol. 2003;49(1):157-65.
[8]: Ahmer BM. Cell-to-cell signalling in Escherichia coli and Salmonella enterica. Mol Microbiol 2004; 52:933-945
[9]: Federle MJ, Bassler BL. Interspecies communication in bacteria. J Clin Invest 2003; 112:1291-1299.
[10]: Vendeville A, Winzer K, Heurlier K, Tang CM, Hardie KR. Making ‘sense’ of metabolism: autoinducer-2, LuxS and pathogenic bacteria. Nat Rev Microbiol 2005; 3:383-396.
[11]: Taga ME, Miller ST, Bassler BL. Lsr-mediated transport and processing of AI-2 in Salmonella typhimurium. Mol Microbiol 2003; 50:1411-1427.
[12]: Xavier KB, Miller ST, Lu W, et al. Phosphorylation and processing of the quorum-sensing molecule autoinducer-2 in enteric bacteria. ACS Chem Biol 2007; 2:128-136.
[13]: Forsyth, M. H., and T. L. Cover. 2000. Intercellular communication in Helicobacter pylori: luxS is essential for the production of an extracellular signaling molecule. Infect. Immun. 68:3193–3199.
[14]: Joyce, E. A., B. L. Bassler, and A. Wright. 2000. Evidence for a signaling system in Helicobacter pylori: detection of a luxS-encoded autoinducer. J. Bacteriol. 182:3638–3643
[15]:Green, A., Silver, P., Collins, J., & Yin, P. (n.d.). Toehold Switches: De-Novo-Designed Regulators of Gene Expression. Cell, 925-939.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 117
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 772