Difference between revisions of "Part:BBa K1689009"
| Line 7: | Line 7: | ||
| − | 2015 Peking iGEM fused Cluc to N terminus of dCas9 (Figure 1). Guided by sgRNA, it binds to target DNA sequence. Together with Nluc-dCas9 [https://parts.igem.org/wiki/index.php?title=Part:BBa_K1689010 (BBa_K1689010)]:sgRNA complex, our paired dCas9 (PC) reporter system was constructed (Figure | + | 2015 Peking iGEM fused Cluc to N terminus of dCas9 (Figure 1) and purified it (Figure 2). Guided by sgRNA, it binds to target DNA sequence. Together with Nluc-dCas9 [https://parts.igem.org/wiki/index.php?title=Part:BBa_K1689010 (BBa_K1689010)]:sgRNA complex, our paired dCas9 (PC) reporter system was constructed (Figure 3) which converts invisible sequence information into measurable bioluminescence. |
[[File:Peking-Cluc-dCas9.png|400px|]] | [[File:Peking-Cluc-dCas9.png|400px|]] | ||
'''Figure 1. Configuration of Cluc-dCas9 fusion protein.''' | '''Figure 1. Configuration of Cluc-dCas9 fusion protein.''' | ||
| + | |||
| + | [[File:Peking-CD-SDS.png|400px|]] | ||
| + | |||
| + | '''Figure 2. SDS-PAGE of the purified Cluc-dCas9 fusion protein.''' | ||
| + | |||
[[File:Peking-CRISPR-Figure2.png|700px|]] | [[File:Peking-CRISPR-Figure2.png|700px|]] | ||
| − | '''Figure | + | '''Figure 3. Schematic of the paired dCas9 (PC) reporter system.''' |
| − | We invented a new protocol for testing our PC reporter system (see Methods). Provided that initial binding of dCas9 depends on the protospacer adjacent motif (PAM, a short 3’ motif adjacent to target sequence), four sets of sgRNA orientation settings were also tested (Figure | + | We invented a new protocol for testing our PC reporter system (see Methods). Provided that initial binding of dCas9 depends on the protospacer adjacent motif (PAM, a short 3’ motif adjacent to target sequence), four sets of sgRNA orientation settings were also tested (Figure 4a). To find how split luciferase-dCas9 fusion architecture influences our PC reporter system, we constructed and tested Nluc-dCas9[https://parts.igem.org/wiki/index.php?title=Part:BBa_K1689010 (BBa_K1689010)], dCas9-Nluc [https://parts.igem.org/wiki/index.php?title=Part:BBa_K1689008 (BBa_K1689008) ]fusion protein to respectively pair with Cluc-dCas9 (Figure 4b). |
a | a | ||
| Line 28: | Line 33: | ||
[[File:Peking-Cluc.png|700px|]] | [[File:Peking-Cluc.png|700px|]] | ||
| − | '''Figure | + | '''Figure 4. Test on Cluc-dCas9 fusion protein across four different sgRNA orientations.''' (a) 4 different sgRNA orientation settings. In orientation PAM-out, the pair of PAM sequences are distal from the spacer sequence, with the 5' end of the sgRNA adjacent to the spacer; in orientation PAM-in, the pair of PAM sequences are adjacent to the spacer sequence, with the 3' end of the sgRNA in proximity to the spacer; in orientation PAM-direct 1 and PAM-direct 2, one PAM sequence is adjacent to and another distal from the spacer. (b) Test on Cluc-dCas9 fusion protein paired with Nluc-dCas9 and dCas9-Nluc across four different sgRNA orientations. |
| − | We chose Nluc-dCas9/ Cluc-dCas9 fusion proteins with sgRNA orientation PAM-out for our PC reporter system because it presents the highest signal. Then we further validate its specificity and sensitivity. We carried out specificity test to distinguish the target plasmid [https://parts.igem.org/Part:BBa_K909009 (BBa_K909009)] from different kinds of non-specific controls (Figure | + | We chose Nluc-dCas9/ Cluc-dCas9 fusion proteins with sgRNA orientation PAM-out for our PC reporter system because it presents the highest signal. Then we further validate its specificity and sensitivity. We carried out specificity test to distinguish the target plasmid [https://parts.igem.org/Part:BBa_K909009 (BBa_K909009)] from different kinds of non-specific controls (Figure 5a,b,c). The results showed that the luminescence intensity using the positive target is significantly higher than that of non-specific controls (Figure 5d). |
[[File:Peking-CRISPR-Figure7.png|700px|]] | [[File:Peking-CRISPR-Figure7.png|700px|]] | ||
| − | '''Figure | + | '''Figure 5. Specificity test of PC reporter system.''' (a) Positive target with two sites recognized simultaneously by a pair of split luciferase-dCas9:sgRNA complexes. (b) Control with only one site recognized by "half pair". (c) Negative control with no site to be recognized. (d) Experimental data. Error bars denote s.d.; n=3. |
| − | Then, we tested the sensitivity of PC reporter system. As shown in Figure 5, PC reporter can still work at the concentration of ~0.1nM target for both plasmid (Figure | + | Then, we tested the sensitivity of PC reporter system. As shown in Figure 5, PC reporter can still work at the concentration of ~0.1nM target for both plasmid (Figure 6a) and crude PCR product (Figure 6b). |
[[File:Peking-CRISPR-Figure8.png|700px|]] | [[File:Peking-CRISPR-Figure8.png|700px|]] | ||
| − | '''Figure | + | '''Figure 6. Exploring the sensitivity of PC reporter using plasmid and PCR product as the target.''' (a) Bioluminescence intensity at different concentrations of plasmid as the target. (b) Bioluminescence intensity at different concentrations of crude PCR product as the target. Both showed that PC reporter was able to detect target DNA at a low concentration (0.1 nM). |
==References== | ==References== | ||
| − | 1. Lei S. Qi, Matthew H. Larson, Luke A. Gilbert et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell, 2013, 152: 1173-1183. | + | 1. Lei S. Qi, Matthew H. Larson, Luke A. Gilbert et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell, 2013, 152: 1173-1183. |
| − | 2. Kathryn E. Luker, Matthew C. P. Smith, et al. Kinetics of regulated protein–protein interactions revealed with firefly luciferase complementation imaging in cells and living animals. PNAS, 2004, 101: 12288-12293. | + | 2. Kathryn E. Luker, Matthew C. P. Smith, et al. Kinetics of regulated protein–protein interactions revealed with firefly luciferase complementation imaging in cells and living animals. PNAS, 2004, 101: 12288-12293. |
<!-- --> | <!-- --> | ||
Revision as of 02:46, 19 September 2015
C-luc-dCas9
Cluc-dCas9 fusion protein ORF
A catalytically dead Cas9 (dCas9), when coexpressed with a guide RNA, generates a DNA recognition complex which can binds to any targeted gene [1]. Firefly luciferase, widely used as a reporter, is split into two fragments, namely Nluc and Cluc [2]. Each protein fragment by itself is inactive, when two fragments are reassembled, the enzymatic activity of the original protein would be reconstituted, providing easily measurable read out.
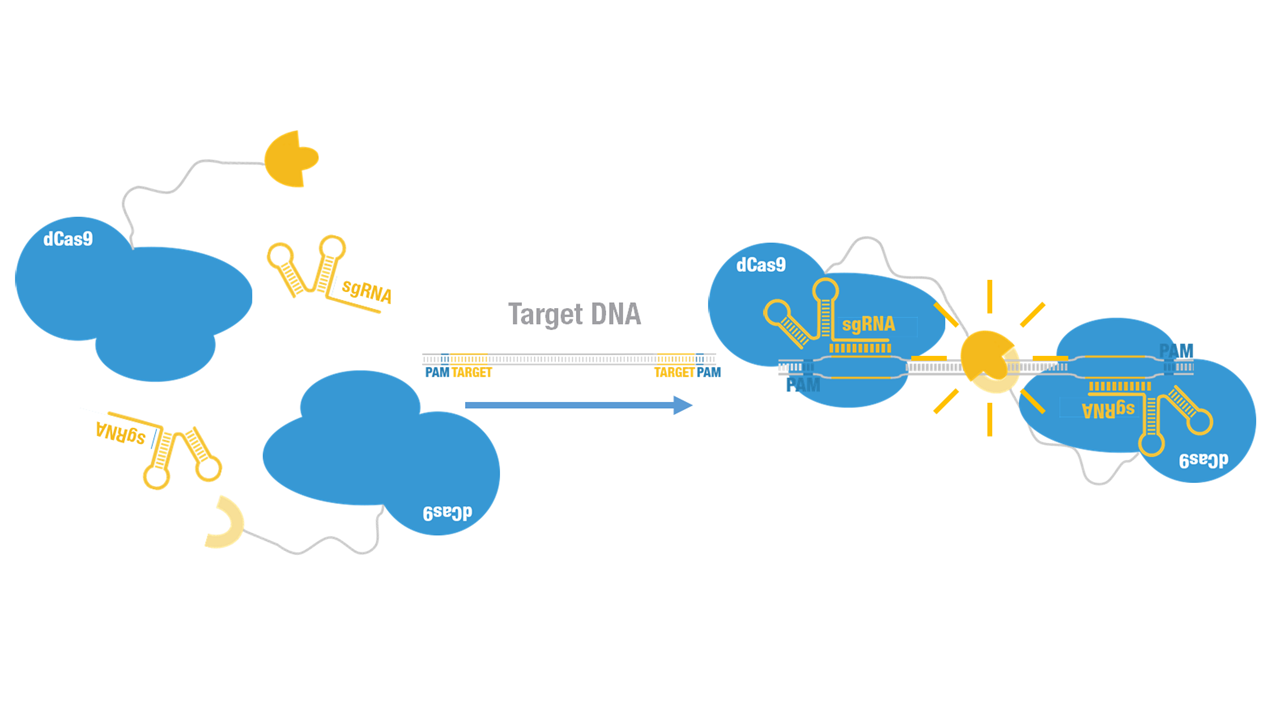
2015 Peking iGEM fused Cluc to N terminus of dCas9 (Figure 1) and purified it (Figure 2). Guided by sgRNA, it binds to target DNA sequence. Together with Nluc-dCas9 (BBa_K1689010):sgRNA complex, our paired dCas9 (PC) reporter system was constructed (Figure 3) which converts invisible sequence information into measurable bioluminescence.
Figure 1. Configuration of Cluc-dCas9 fusion protein.
Figure 2. SDS-PAGE of the purified Cluc-dCas9 fusion protein.
Figure 3. Schematic of the paired dCas9 (PC) reporter system.
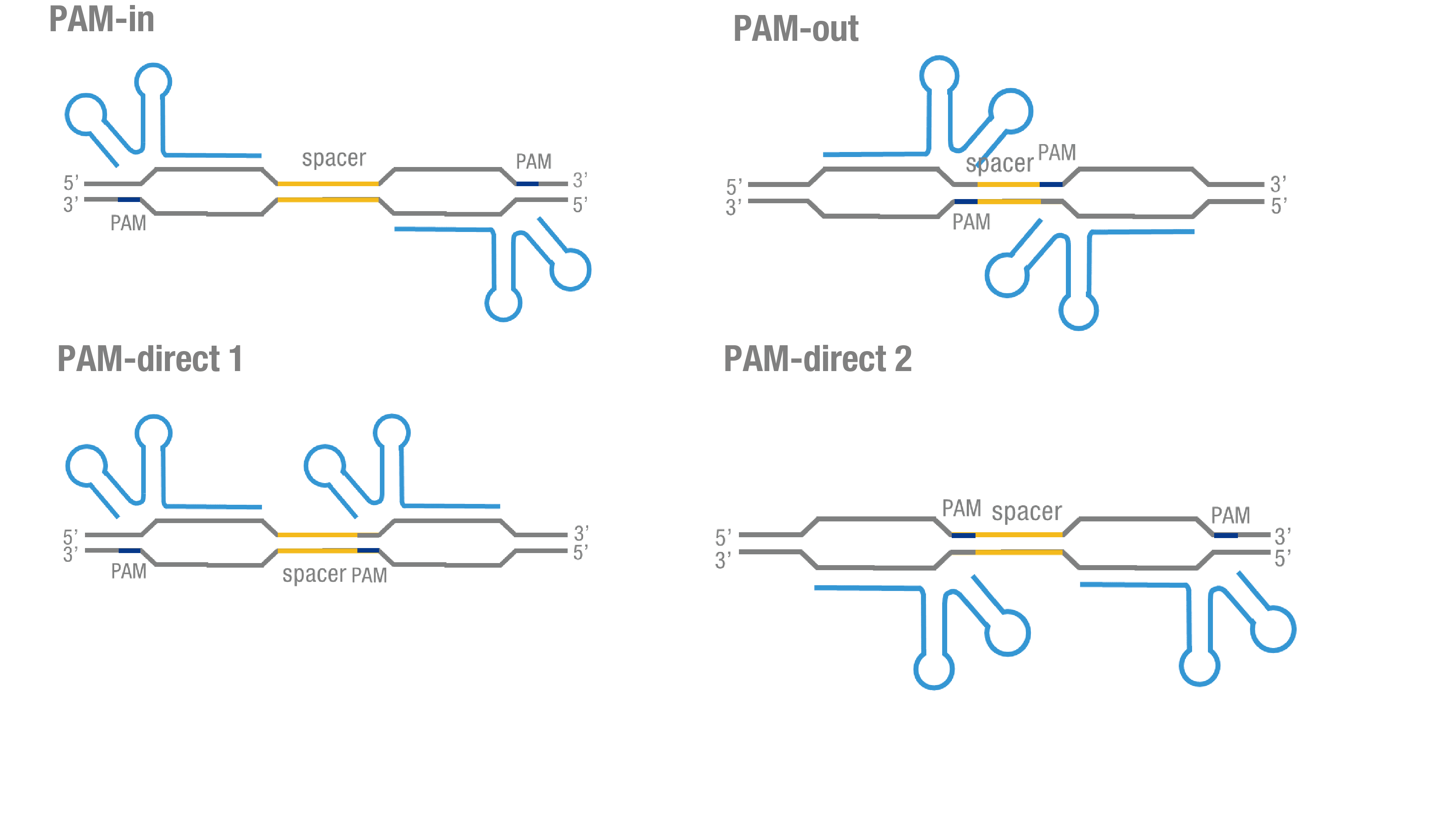
We invented a new protocol for testing our PC reporter system (see Methods). Provided that initial binding of dCas9 depends on the protospacer adjacent motif (PAM, a short 3’ motif adjacent to target sequence), four sets of sgRNA orientation settings were also tested (Figure 4a). To find how split luciferase-dCas9 fusion architecture influences our PC reporter system, we constructed and tested Nluc-dCas9(BBa_K1689010), dCas9-Nluc (BBa_K1689008) fusion protein to respectively pair with Cluc-dCas9 (Figure 4b).
a
b
Figure 4. Test on Cluc-dCas9 fusion protein across four different sgRNA orientations. (a) 4 different sgRNA orientation settings. In orientation PAM-out, the pair of PAM sequences are distal from the spacer sequence, with the 5' end of the sgRNA adjacent to the spacer; in orientation PAM-in, the pair of PAM sequences are adjacent to the spacer sequence, with the 3' end of the sgRNA in proximity to the spacer; in orientation PAM-direct 1 and PAM-direct 2, one PAM sequence is adjacent to and another distal from the spacer. (b) Test on Cluc-dCas9 fusion protein paired with Nluc-dCas9 and dCas9-Nluc across four different sgRNA orientations.
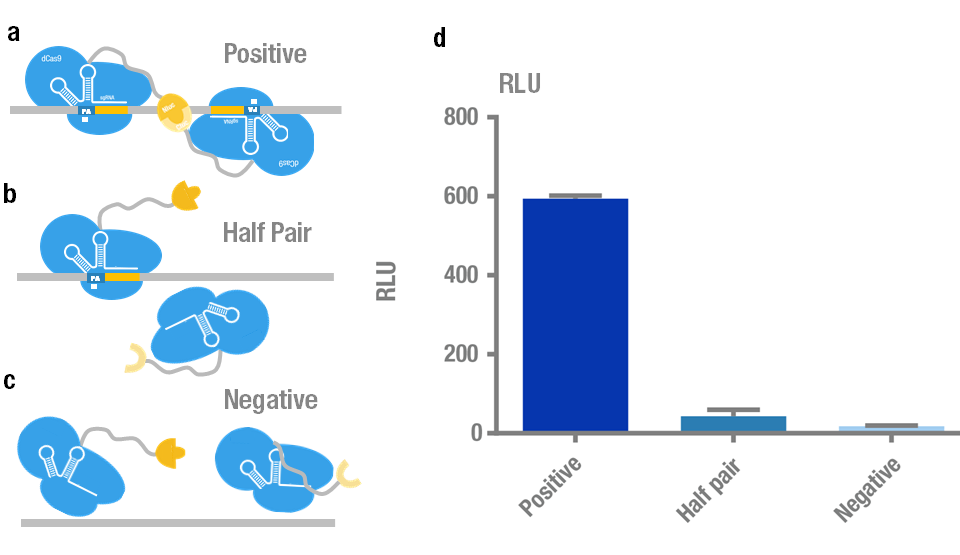
We chose Nluc-dCas9/ Cluc-dCas9 fusion proteins with sgRNA orientation PAM-out for our PC reporter system because it presents the highest signal. Then we further validate its specificity and sensitivity. We carried out specificity test to distinguish the target plasmid (BBa_K909009) from different kinds of non-specific controls (Figure 5a,b,c). The results showed that the luminescence intensity using the positive target is significantly higher than that of non-specific controls (Figure 5d).
Figure 5. Specificity test of PC reporter system. (a) Positive target with two sites recognized simultaneously by a pair of split luciferase-dCas9:sgRNA complexes. (b) Control with only one site recognized by "half pair". (c) Negative control with no site to be recognized. (d) Experimental data. Error bars denote s.d.; n=3.
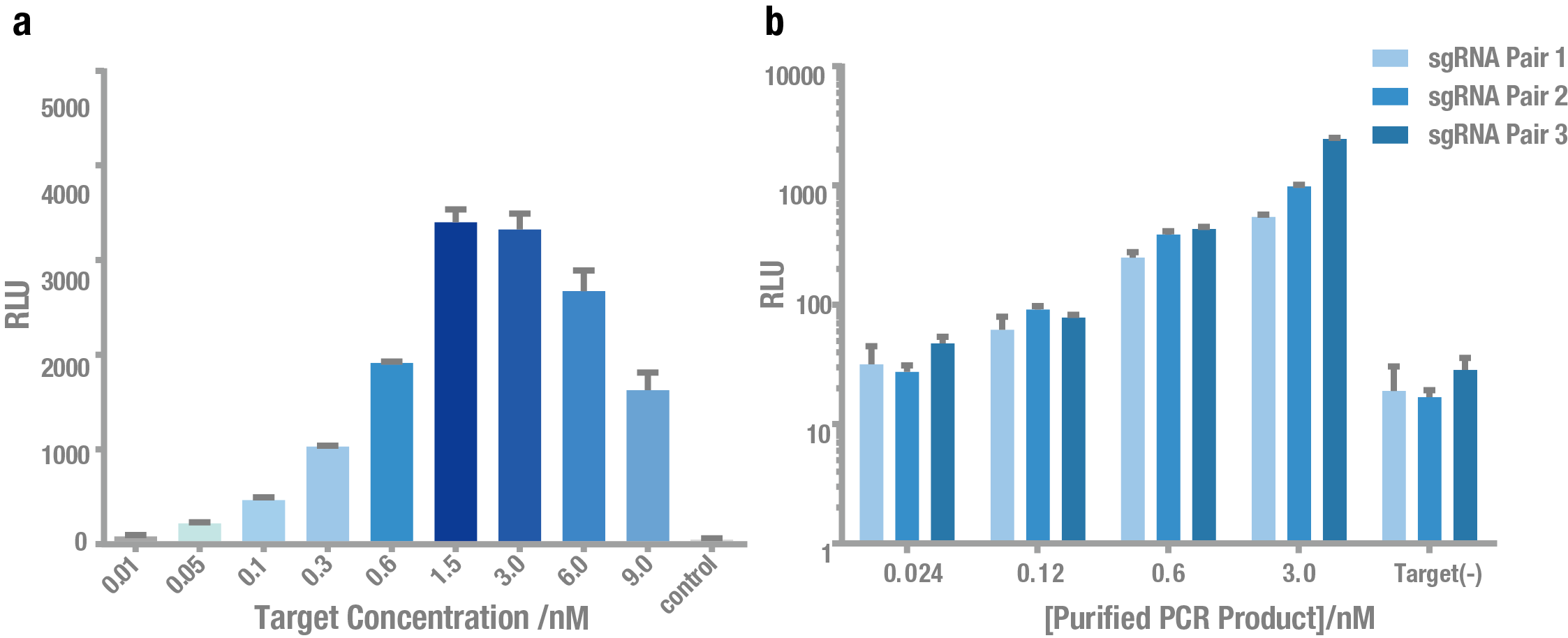
Then, we tested the sensitivity of PC reporter system. As shown in Figure 5, PC reporter can still work at the concentration of ~0.1nM target for both plasmid (Figure 6a) and crude PCR product (Figure 6b).
Figure 6. Exploring the sensitivity of PC reporter using plasmid and PCR product as the target. (a) Bioluminescence intensity at different concentrations of plasmid as the target. (b) Bioluminescence intensity at different concentrations of crude PCR product as the target. Both showed that PC reporter was able to detect target DNA at a low concentration (0.1 nM).
References
1. Lei S. Qi, Matthew H. Larson, Luke A. Gilbert et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell, 2013, 152: 1173-1183.
2. Kathryn E. Luker, Matthew C. P. Smith, et al. Kinetics of regulated protein–protein interactions revealed with firefly luciferase complementation imaging in cells and living animals. PNAS, 2004, 101: 12288-12293.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 1662
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 3
Illegal BamHI site found at 3941 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 326
Illegal AgeI site found at 465 - 1000COMPATIBLE WITH RFC[1000]