Difference between revisions of "Part:BBa K1761006"
(Bioluminiscence NanoBit) |
(References) |
||
| Line 5: | Line 5: | ||
* Sequence will be published later. | * Sequence will be published later. | ||
| + | <!-- --> | ||
| + | <span class='h3bb'>Sequence and Features</span> | ||
| + | <partinfo>BBa_K1761006 SequenceAndFeatures</partinfo> | ||
| + | |||
| + | |||
| + | <!-- Uncomment this to enable Functional Parameter display | ||
| + | ===Functional Parameters=== | ||
| + | <partinfo>BBa_K1761006 parameters</partinfo> | ||
| + | <!-- --> | ||
| + | |||
== Usage and Biology == | == Usage and Biology == | ||
| Line 61: | Line 71: | ||
''Figure 7: Bioluminescence of results of the NanoBit construct.'' | ''Figure 7: Bioluminescence of results of the NanoBit construct.'' | ||
| − | |||
| − | |||
| − | |||
| − | + | == References == | |
| − | + | [1] Kyle Hooper, "Applications of a smaller, brighter, more versatile luciferase: NanoLuc™ Luciferase Technology", Presentation slides Fall 2012. [https://www.promega.com/~/media/files/promega%20worldwide/north%20america/promega%20us/webinars%20and%20events/nanoluctourtalkfinal.pdf?la=en] | |
| − | == | + | |
| − | + | ||
| − | + | ||
Revision as of 21:54, 18 September 2015
SmallBit Split Luciferase
SmallBit is the small part of a Split Luciferase. Its molecular weight is 1 kDa. This construct on its own has no function since SmallBit on itself has no luminescence activity. For luminescence activity, the LargeBit of the Split Luciferace is needed.
- Sequence will be published later.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Usage and Biology
Split luciferases can be used as signaling components. This split luciferase consists of two parts, namely LargeBit and SmallBit. These parts were both inserted in a pETDuet-1 vector together with OmpX and a linker. LargeBit and SmallBit have an affinity towards each other, so when in close proximity they will come together and will give a luminescence signal. Followed is a short description of each part and of the NanoBit construct.
OmpX - SmallBit BBa_K1761006 [1]
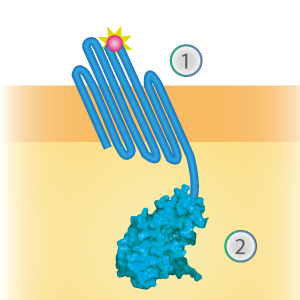
SmallBit is the small part of this Split Luciferase. Its molecular weight is 1 kDa, it is only 11 amino acids long. OmpX (1) (with a correct mutation for the amber stop codon TAG), a BsoBI-linker and SmallBit (2) together will look like Figure 1. This construct on its own has no function since SmallBit on itself has no luminescence activity.
Figure 1: Schematical overview of the OmpX - SmallBit construct.
OmpX - LargeBit BBa_K1761005
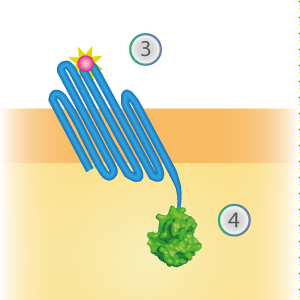
LargeBit is the big part of this Split Luciferase. Its molecular weight is 18 kDa. OmpX (1) (with a correct mutation for the amber stop codon TAG), a BamHI-linker and LargeBit (2) together will look like Figure 2. This construct on its own has no function since LargeBit on itself has no luminescence activity.
Figure 2: Schematical overview of the OmpX - LargeBit construct.
NanoBit construct
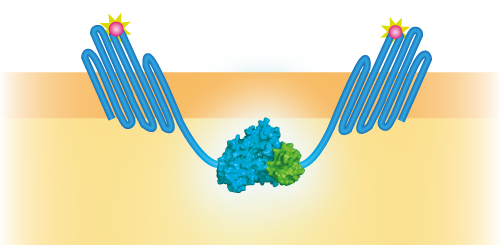
NanoBit utilizes a structural complementation-based approach to monitor protein interactions within living cells. Protein interaction promotes structural complementation and generation of a bright, luminescent enzyme. Protein dynamics can be followed in real-time in living cells following addition of the Nano-Glo Live Cell Reagent, a non-lytic detection reagent containing the cell-permeable furamizine substrate. See Figure 3 for the whole construct.
Figure 3: Schematical overview of the NanoBit construct.
Characterization
When mutated with the amber stop codon TAG, a unnatural amino acid with an azide-functionalized group can be expressed. After the expression of this amino acid, OmpX can covalently bind almost anything, as long as it contains a DBCO-functionalized group. The binding finds place by using a bio-orthogonal “click” reaction (SPAAC chemistry). To test the functionality of this “click” reaction, some experiments were done by clicking DBCO-PEG4-TAMRA at the surface. For all the experiments, the following vectors were used: pETDuet-1 with one or two construct(s) inserted (OmpX + intracellular protein) and pEVOL-pAzF (tRNA + tRNA synthetase). Both vectors were transformed into BL21(DE3). The expression was introduced by adding arabinose, IPTG and the unnatural amino acid.
DBCO-PEG4-TAMRA Confirmation
To confirm whether OmpX is in the membrane and whether or not the unnatural amino acid is being incorporated into OmpX, DBCO-PEG4-TAMRA was used. TAMRA is a fluorescent dye that can be used to verify the “click” reaction. If the unnatural amino acid is present, DBCO-PEG4-TAMRA should “click” to the transmembrane protein OmpX and stay there. This can be analyzed with FACS. For more information about how to perform FACS experiments, see our Protocol Page [http://2015.igem.org/Team:TU_Eindhoven/Project/Protocols].
To verify that OmpX is in the membrane, we used the OmpX – LgBiT and OmpX – SmBiT constructs. These gave the following results after clicking with DBCO-PEG4-TAMRA (see Figure 4, 5 and 6). From this it can be concluded that OmpX is in the membrane and that the “click” reaction works.
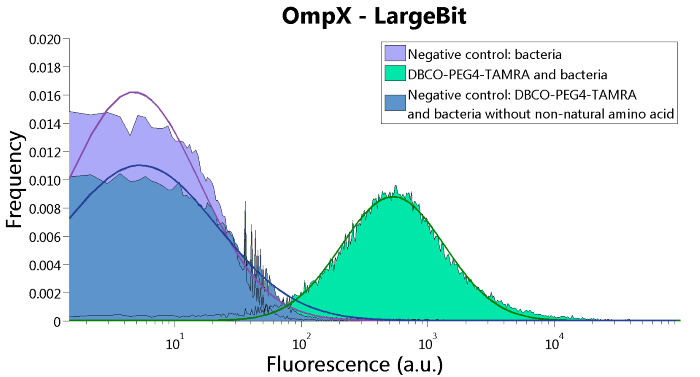
Figure 4: FACS results of OmpX - LargeBit.
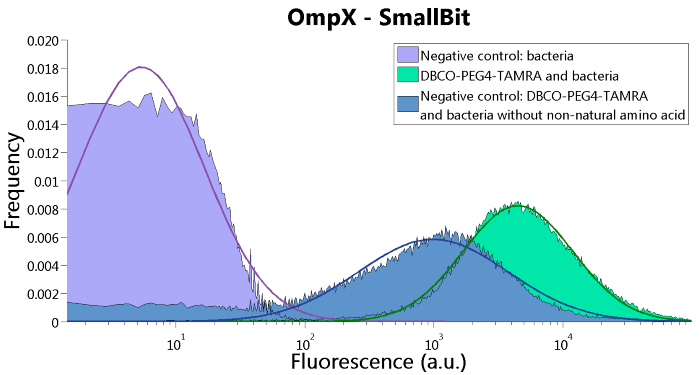
Figure 5: FACS results of OmpX - SmallBit
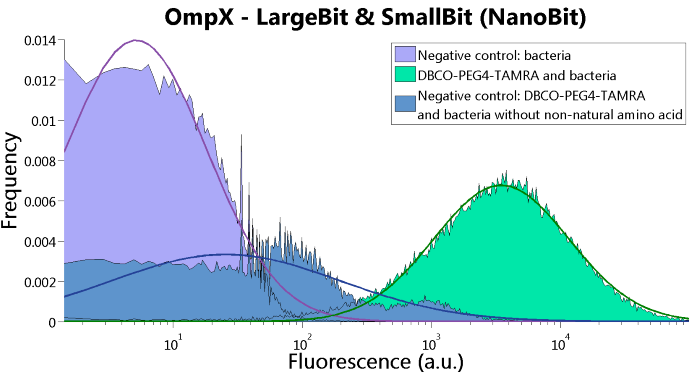
Figure 6: FACS results of OmpX - LargeBit and OmpX - SmallBit.
Bioluminescence Confirmation
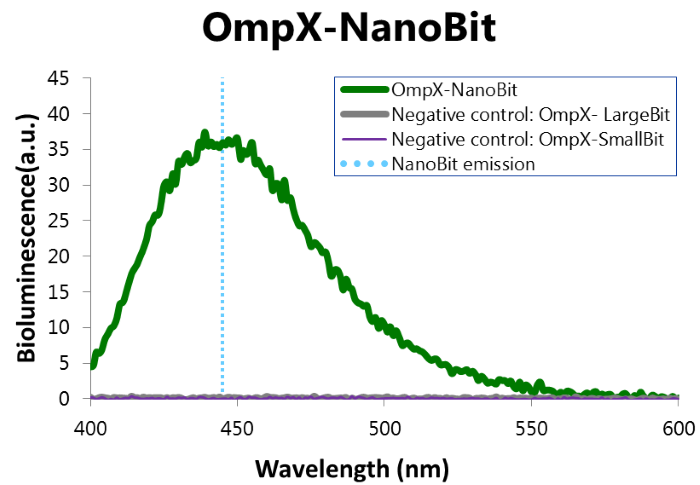
To confirm whether NanoBit is present and is working, a bioluminescence measurement was performed. The results of this experiment are shown in Figure 7.
Figure 7: Bioluminescence of results of the NanoBit construct.
References
[1] Kyle Hooper, "Applications of a smaller, brighter, more versatile luciferase: NanoLuc™ Luciferase Technology", Presentation slides Fall 2012. [2]