Difference between revisions of "Part:BBa K1351028"
| Line 50: | Line 50: | ||
The coverage of RBS strength as calculated by the [https://salis.psu.edu/software/ RBS calculator] is depicted in the Figure 1 shown below. It shows an evenly distributed strength spanning approxiamately three orders of magnitude. | The coverage of RBS strength as calculated by the [https://salis.psu.edu/software/ RBS calculator] is depicted in the Figure 1 shown below. It shows an evenly distributed strength spanning approxiamately three orders of magnitude. | ||
| − | [[File:LMU_14_RBSlibrary.png|thumb| | + | [[File:LMU_14_RBSlibrary.png|thumb|center|400px|Fig.1: Coverage of the translation initiation rate of the RBS library as calculated by the [https://salis.psu.edu/software/ RBS calculator]]] |
Revision as of 06:30, 1 November 2014
LMU Bacillus RBS collection 1
This part was generated in RFC10, and has the following prefix and suffix:
Prefix Suffix
5' - GAATTC GCGGCCGC T TCTAGA G aaggagggata T ACTAGT A GCGGCCG CTGCAG - 3'
EcoRI NotI XbaI SpeI NotI PstI
This part is used in the 2014 LMU-Munich iGEM project [http://2014.igem.org/Team:LMU-Munich BaKillus].
This library was generated utilizing the Salis lab RBS calculator to find RBS that cover a high space of translation intitiation rate in 'B. subtilis'. It includes seven RBS with the following sequences:
| Number | BBa_K1310 .. | Sequence |
|---|---|---|
| 1 | 28 | aaggagggata |
| 2 | 29 | agaggaggata |
| 3 | 30 | aaggagagata |
| 4 | 31 | aggagaggata |
| 5 | 32 | aagaggagata |
| 6 | 33 | agaaagggata |
| 7 | 34 | aagaagagata |
The coverage of RBS strength as calculated by the RBS calculator is depicted in the Figure 1 shown below. It shows an evenly distributed strength spanning approxiamately three orders of magnitude.
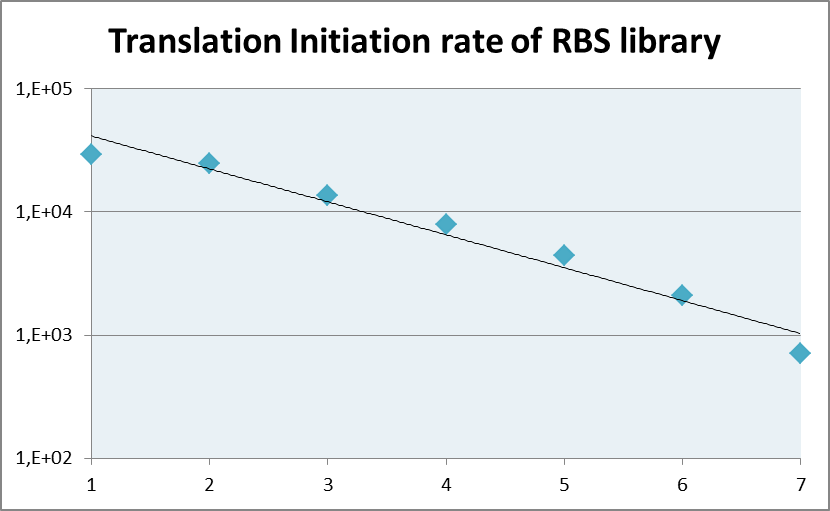
Fig.1: Coverage of the translation initiation rate of the RBS library as calculated by the RBS calculator
Sequence and Features
Assembly Compatibility:
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
