Difference between revisions of "Part:BBa K1150034"
| Line 44: | Line 44: | ||
[[File:Multiple_targeting_Freiburg_2013_%281%29.png|800px|]] <br> | [[File:Multiple_targeting_Freiburg_2013_%281%29.png|800px|]] <br> | ||
'''Figure 1.:''' Scheme of the RNAimer. Two RNAs and an insertion site. | '''Figure 1.:''' Scheme of the RNAimer. Two RNAs and an insertion site. | ||
| + | <br> <br> <br> | ||
| + | === Proof of functionality === | ||
| + | To test the functionality of this plasmid we used the dCas9-VP16 construct in combination with several targets. We tested if we still can activate transcription of a SEAP reporter with this RNA plasmid. If this is functional we expect an incerase in SEAP activity in the supernatant. | ||
| + | Cells were seeded in 24-well plates at a density of 65.000 cells. 24 hrs later cells were transfected with a CMV<sub>min</sub>:SEAP and RNAimer plasmids containing different target sites in front of the promoter. After 48 hours SEAP was assayed as descirbed in our standard protocol. | ||
| + | |||
| + | |||
| + | [[File:Multiple_Targeting_Freiburg_2013_%285%29.png|800px|]] <br> | ||
| + | '''Figure 2.:''' Activation via dCas9-VP16 and RNAimer. It is clearly visible that activation is depending on which target site is ligated into the RNAimer. <br> <br> | ||
==References== | ==References== | ||
[1]Cong, L., Ran, F.A., Cox, D., Lin, S., Barretto, R., Habib, N., Hsu, P.D., Wu, X., Jiang, W., Marraffini, L.A., Zhang, F. (2013). Multiplex Genome Engineering Using CRISPR/Cas Systems. Science 339 (6121), 819-23 <br> | [1]Cong, L., Ran, F.A., Cox, D., Lin, S., Barretto, R., Habib, N., Hsu, P.D., Wu, X., Jiang, W., Marraffini, L.A., Zhang, F. (2013). Multiplex Genome Engineering Using CRISPR/Cas Systems. Science 339 (6121), 819-23 <br> | ||
Revision as of 19:32, 2 October 2013
dCas RNA plasmid: "uniCAS RNAimer"
| pH1:tracrRNA pU6:DR_dummy:DR | |
|---|---|
| Function | two small RNAs giving rise to
specific binding of dCas9 to DNA loci |
| Use in | Mammalian cells |
| RFC standard | RFC 25 |
| Backbone | pSB1C3 |
| Organism | Streptococcus pyogenes |
| Source | Feng Zhang, Addgene |
| Submitted by | [http://2013.igem.org/Team:Freiburg Freiburg 2013] |
This device contains loci that can code for small RNAs that are able to target the dCas9 protein [1] to any desired locus containg a so-called PAM seqeunce. It consists of a human H1 promoter driving the expression of the tracrRNA which mediates the interaction between the dCas9 protein and the second RNA, the crRNA. This crRNA gives rise to the specific binding of dCAS9 to DNA and is under control of the human U6 promoter. The 30bp target site can be inserted by simply opening the plasmid with BbsI and ligating it into the plasmid. The flanking direct repeats (DR) will mediate interaction with the tracrRNA. [1]
Multiple target sites can be combined on one plasmid by using the BioBrick standard and enables to target multiple loci at once with a small amount of plasmids.
The very minimalistic pSB1C3 backbone guarantees strong expression of the RNAs, which are suspected to be the limiting factor of dCAS9 interaction.
This plasmid should be used in combination of dCAS9 effector proteins, that can be found in the registry, e.g. [2] or [3].
An efficient tool for designing target sites can be found at Freiburg 2013 teams website. [http://2013.igem.org/Team:Freiburg/Project/crrna]
Assembly
On this plasmid are two RNAs encoded. the tracrRNA is mediating the interaction between dCas9 and the crRNA. the direct repeats (DRs) of the crRNA are complementary to part of tracrRNA. The inserted locus n the crRNA is complementary to the locus at which the dCas9 should be targeted. On this plasmid a 10bp long dummy DNA is inserted, which has two BbsI cleavage sites. When digesting with BbsI the locus is opened and the DRs do have sticky ends, so new targets can be ligated easily. For target sites annealed oligos are working fine.
Once single loci are inserted it is simple to combine several loci on one plasmid using iGEM idempotent cloning.
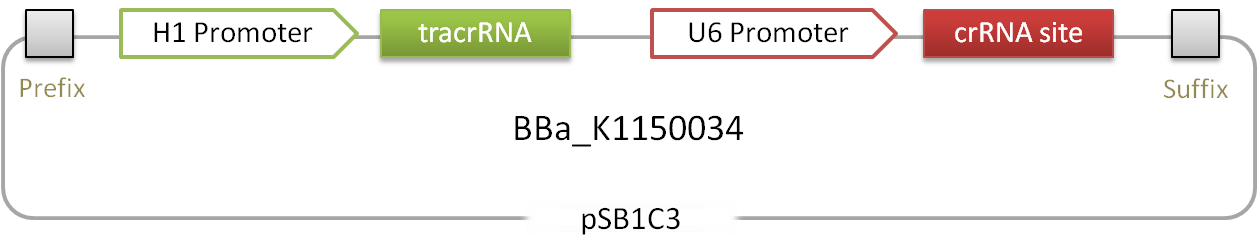
Figure 1.: Scheme of the RNAimer. Two RNAs and an insertion site.
Proof of functionality
To test the functionality of this plasmid we used the dCas9-VP16 construct in combination with several targets. We tested if we still can activate transcription of a SEAP reporter with this RNA plasmid. If this is functional we expect an incerase in SEAP activity in the supernatant. Cells were seeded in 24-well plates at a density of 65.000 cells. 24 hrs later cells were transfected with a CMVmin:SEAP and RNAimer plasmids containing different target sites in front of the promoter. After 48 hours SEAP was assayed as descirbed in our standard protocol.
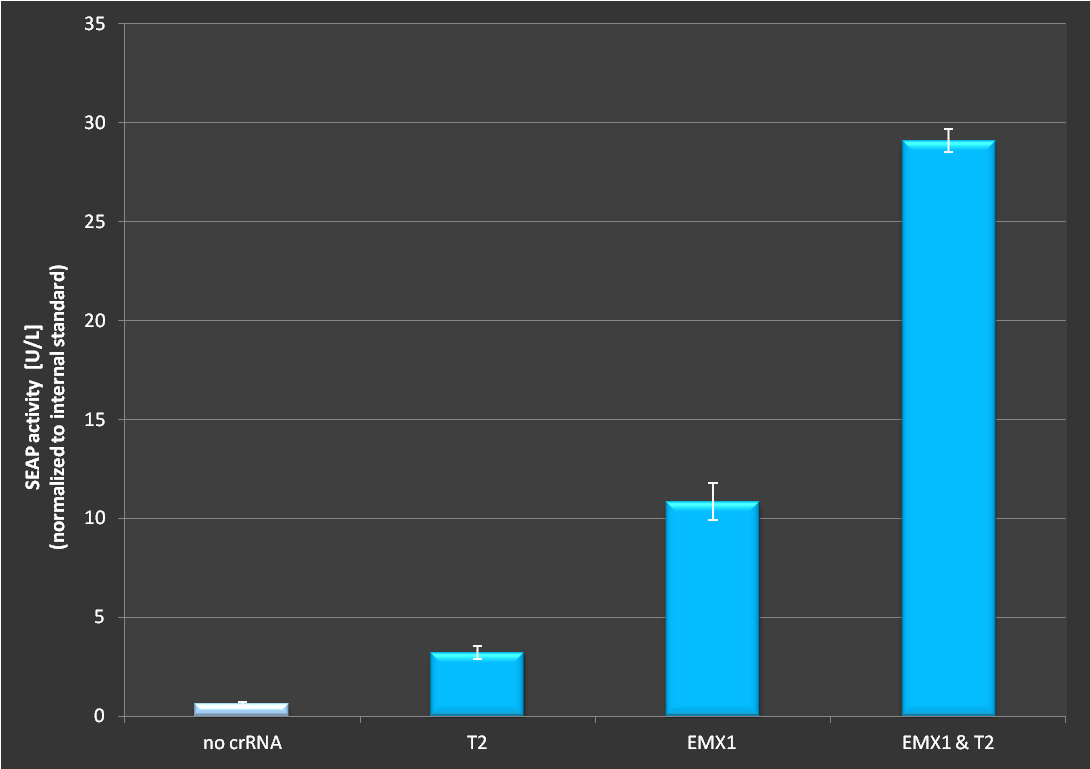
Figure 2.: Activation via dCas9-VP16 and RNAimer. It is clearly visible that activation is depending on which target site is ligated into the RNAimer.
References
[1]Cong, L., Ran, F.A., Cox, D., Lin, S., Barretto, R., Habib, N., Hsu, P.D., Wu, X., Jiang, W., Marraffini, L.A., Zhang, F. (2013). Multiplex Genome Engineering Using CRISPR/Cas Systems. Science 339 (6121), 819-23
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 224
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 216
Literature
