Difference between revisions of "Part:BBa K1041000:Experience"
(→Team NRP-UEA_Norwich 2013) |
(→Team NRP-UEA_Norwich 2013) |
||
| Line 7: | Line 7: | ||
Team NRP-UEA_Norwich 2013 added an NdeI restriction site at the start of the RFP coding region of [[BBa_J04450]] using mutagenesis. This was to allow either the promoter region or RFP gene to be excised and exchanged for a different promoter or gene and provide restriction sites for further cloning such as part [[Bba_K1041002]]. | Team NRP-UEA_Norwich 2013 added an NdeI restriction site at the start of the RFP coding region of [[BBa_J04450]] using mutagenesis. This was to allow either the promoter region or RFP gene to be excised and exchanged for a different promoter or gene and provide restriction sites for further cloning such as part [[Bba_K1041002]]. | ||
| − | + | Characterisation of this biobrick involved comparisons with the original Bba_J04450 biobrick by performing transformations, restriction digests and BLAST analysis. We also had our biobrick sequenced. | |
| + | ===Transformation=== | ||
| + | |||
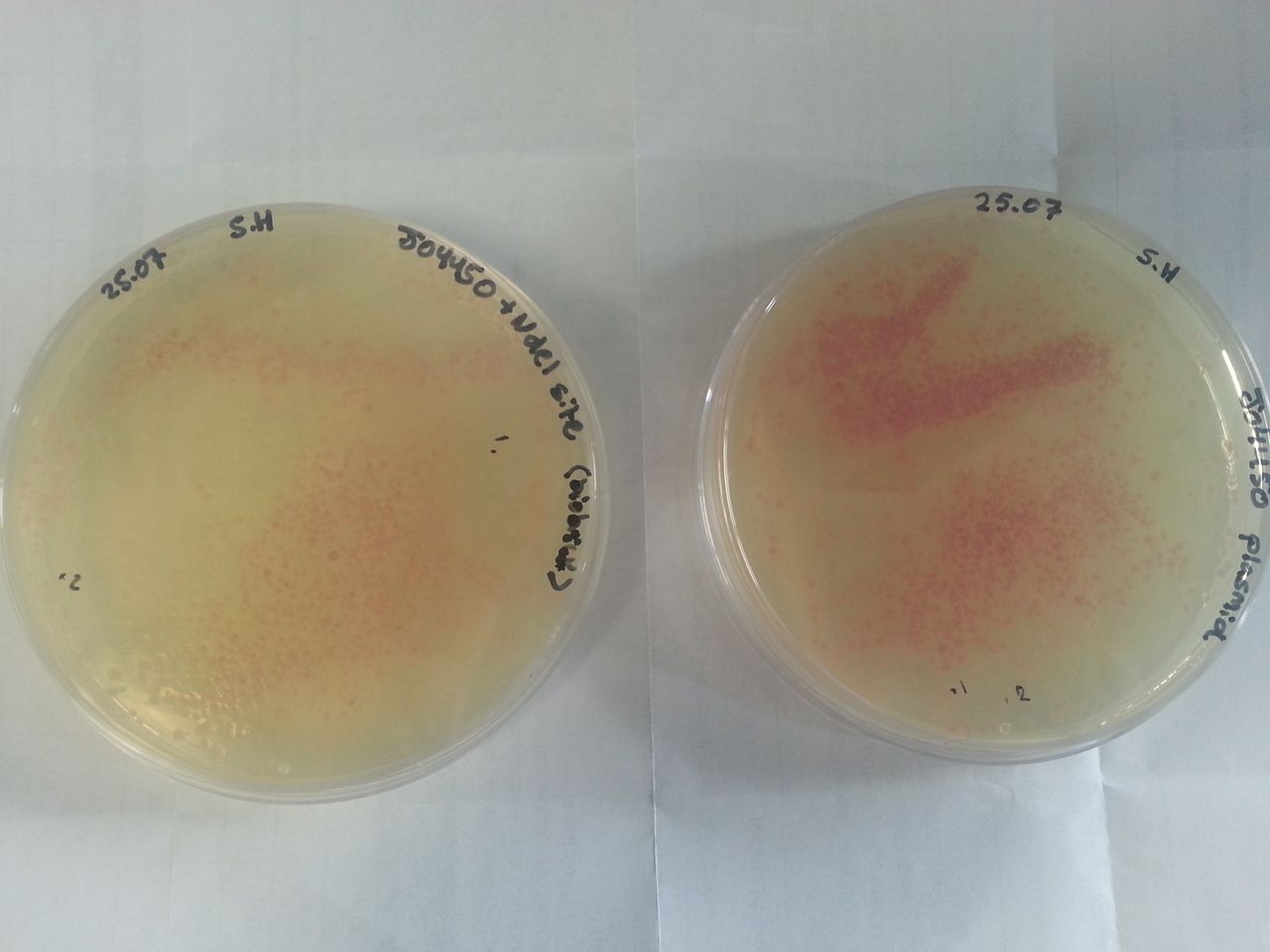
| + | Parts BBa_K1041000 and BBa_J04450 were transformed into Alpha-Select competent ''E.Coli'' cells and plated onto LB Agar plates which contained the antibiotic ampicillin. Both plates have colonies that appear red under natural light ''fig.1''. Therefore the addition of a NdeI site has not effected the ability for the RFP gene to be transcribed. | ||
| + | |||
| + | [[Image:WP 000051.jpg|thumb|Fig 1: Plates containing (left) BBa_K1041000 and (right) BBa_J04450 visualized under non-UV lightbox]] | ||
| + | |||
| + | ===Sequencing=== | ||
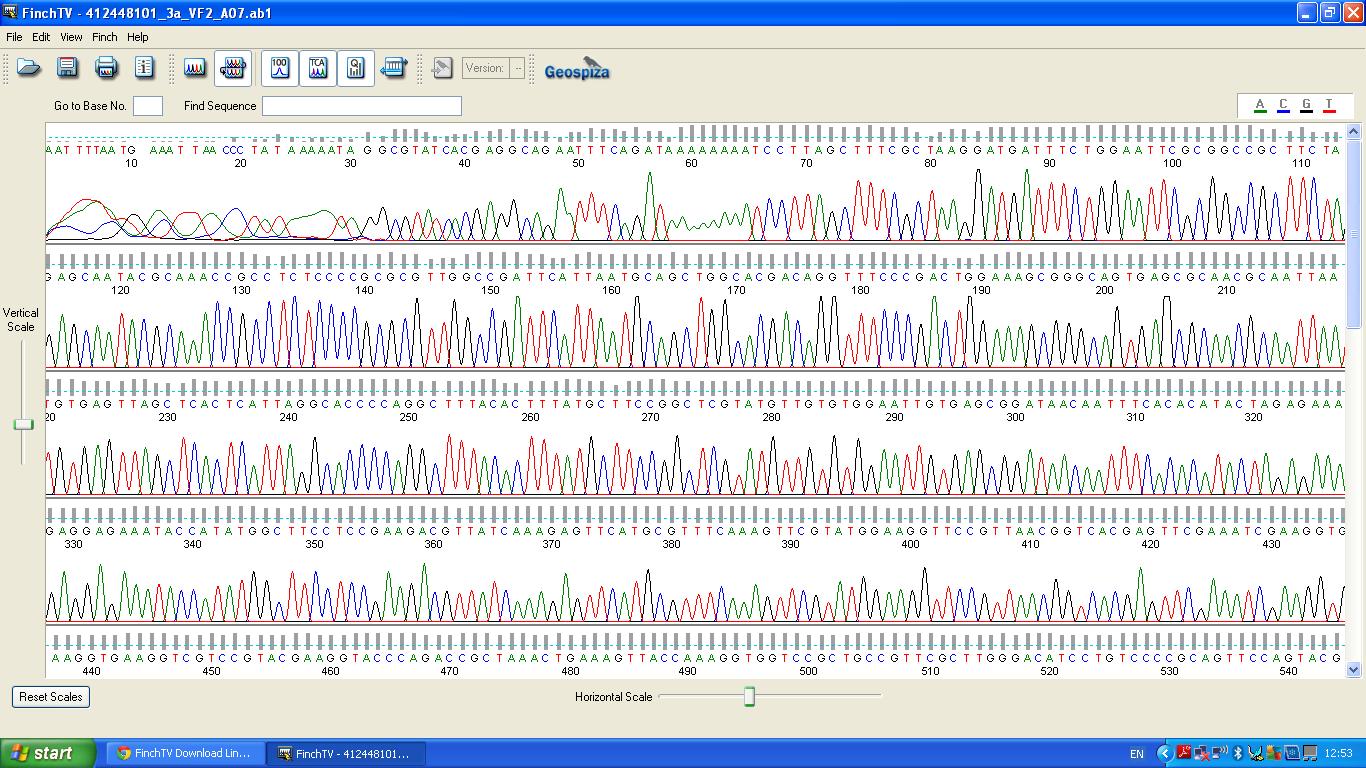
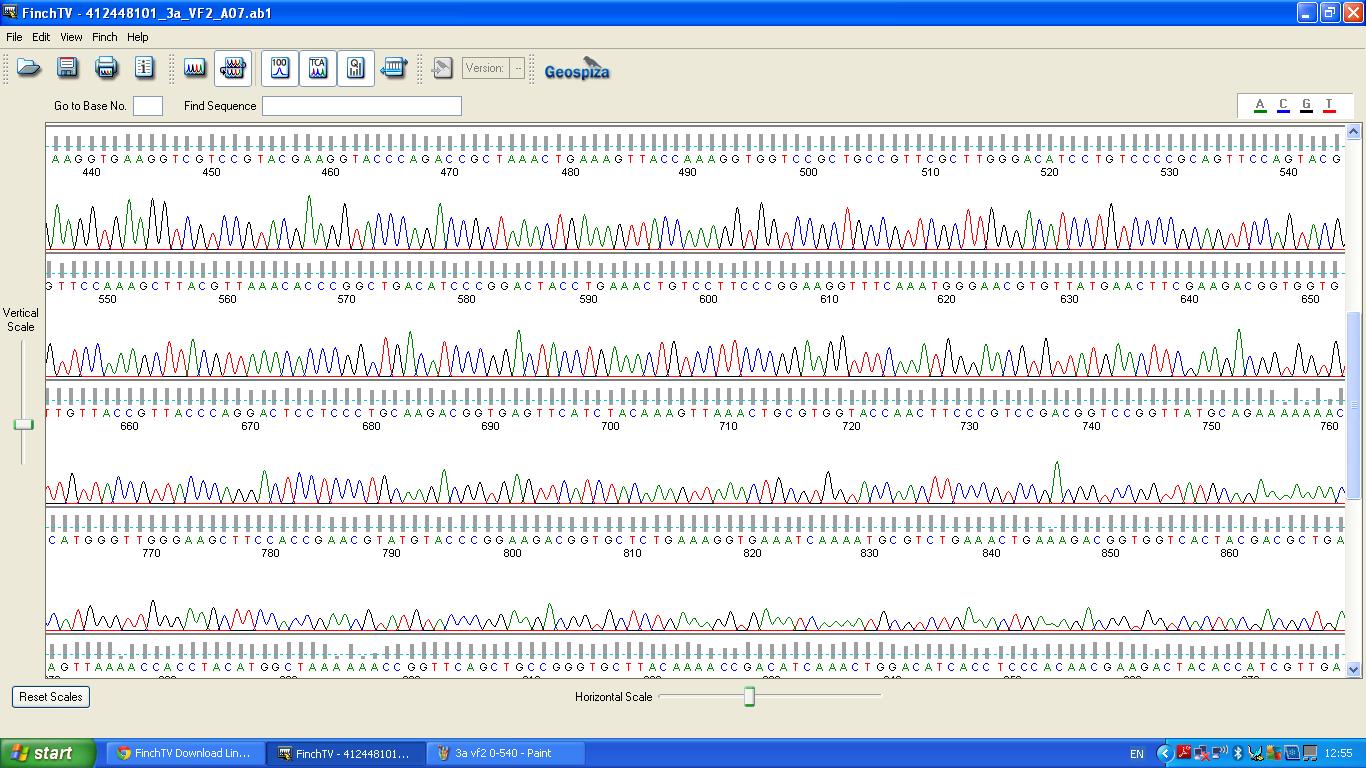
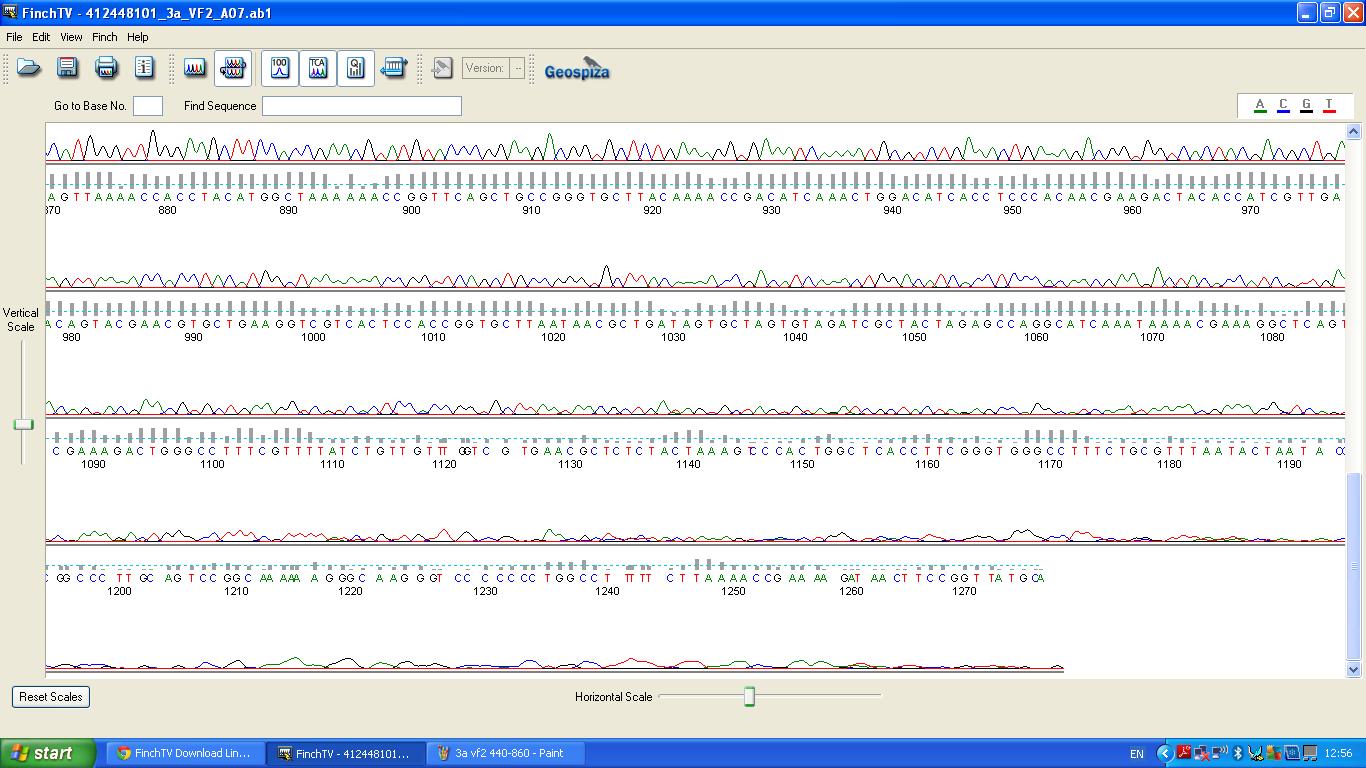
| + | The biobrick was sent off to a company for sequencing. | ||
| + | |||
| + | [[image:3a vf2 0-540.JPG|thumb|left|K1041000 sequencing data part 1]] | ||
| + | [[image:3a vf2 440-860.JPG|thumb|left|K1041000 sequencing data part 2 ]] | ||
| + | [[image:3a vf2 860-.JPG|thumb|left|K1041000 sequencing data part 3]] | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | ===BLAST Analysis=== | ||
| + | The data we recieved back from the sequencing company was aligned using BLAST with the expected DNA sequence ''fig.2'' | ||
===User Reviews=== | ===User Reviews=== | ||
Revision as of 11:04, 17 September 2013
This experience page is provided so that any user may enter their experience using this part.
Please enter
how you used this part and how it worked out.
Team NRP-UEA_Norwich 2013
Team NRP-UEA_Norwich 2013 added an NdeI restriction site at the start of the RFP coding region of BBa_J04450 using mutagenesis. This was to allow either the promoter region or RFP gene to be excised and exchanged for a different promoter or gene and provide restriction sites for further cloning such as part Bba_K1041002.
Characterisation of this biobrick involved comparisons with the original Bba_J04450 biobrick by performing transformations, restriction digests and BLAST analysis. We also had our biobrick sequenced.
Transformation
Parts BBa_K1041000 and BBa_J04450 were transformed into Alpha-Select competent E.Coli cells and plated onto LB Agar plates which contained the antibiotic ampicillin. Both plates have colonies that appear red under natural light fig.1. Therefore the addition of a NdeI site has not effected the ability for the RFP gene to be transcribed.
Sequencing
The biobrick was sent off to a company for sequencing.
BLAST Analysis
The data we recieved back from the sequencing company was aligned using BLAST with the expected DNA sequence fig.2
User Reviews
UNIQ24551b1bf3c0df12-partinfo-00000000-QINU UNIQ24551b1bf3c0df12-partinfo-00000001-QINU