Difference between revisions of "Part:BBa I13522:Experience"
Lisamsonne (Talk | contribs) |
Lisamsonne (Talk | contribs) (→iGEM team Wageningen UR) |
||
| Line 41: | Line 41: | ||
This brick was used to create GFP with a His tag at the C-terminus and the E-coil at the N-terminus to use it as reporter for the E- and K-coil [http://2012.igem.org/Team:Wageningen_UR/Coil_system Plug and Apply sytem] | This brick was used to create GFP with a His tag at the C-terminus and the E-coil at the N-terminus to use it as reporter for the E- and K-coil [http://2012.igem.org/Team:Wageningen_UR/Coil_system Plug and Apply sytem] | ||
| − | [https://parts.igem.org/wiki/index.php?title=Part:BBa_K883701 BBa_K883701] | + | see [https://parts.igem.org/wiki/index.php?title=Part:BBa_K883701 BBa_K883701]; [https://parts.igem.org/wiki/index.php?title=Part:BBa_K883700 BBa_K883700]; [https://parts.igem.org/wiki/index.php?title=Part:BBa_K883702 BBa_K883702] and [https://parts.igem.org/wiki/index.php?title=Part:BBa_K883703 BBa_K883703] |
Revision as of 17:15, 25 September 2012
For parts BBa_I13521 and BBa_I13522, their excitation and emission maxima were determined. BBa_B0034 was used as a negative control. These data were used later in following characterisation experiments as set parameters. The host was E. coli Top Ten in all experiments.
Meanwhile, the graph below shows the emission scan for GFP in part BBa_I13522. Since it was the best option in terms of fluorescence yield, 395 nm was set as the excitation wavelength. As it can be seen from the graph, maximum emission wavelength is 515 nm.

The characterization was performed by METU-TURKEY IGEM 2010 Team.
Thermostability Assay
For this BioBrick, the denaturation temperature was determined by heating the protein at a range of temperatures, and then measuring the fluorescence. This is a useful characterisation as it allows the selection of an appropriate reporter gene for the required temperature.
Stock solutions of GFP were prepared by extracting the protein from cell lysate, and then 50 μl aliquots of the solution were heated in a PCR thermocycler along a temperature gradient.
After two hours, 30 μl was removed from each aliquot and diluted with 170 μl of 20 mM Tris buffer to give 200 μl samples. The fluorescence of the samples was then measured on a 96-well plate. The corresponding curve was plotted on the graph below.

Results of the heat denaturation experiment. The temperature at which half of the protein is denatured is measured by observing its fluorescence (PTm50) mRFP1: 82.2°C; GFPmut3b: 61.6°C; Dendra2: 89.1°C; sfGFP: 75.0°C.
The sigmoidal curves that were calculated gave us the following function which also created the coefficient K which happens to relate to PTm50 (temperature at which half of the protein is denatured measured by looking at its fluorescence):

This characterisation was performed by the Imperial College London 2011 iGEM team.
Applications of BBa_I13522
Haynes Lab, 2012
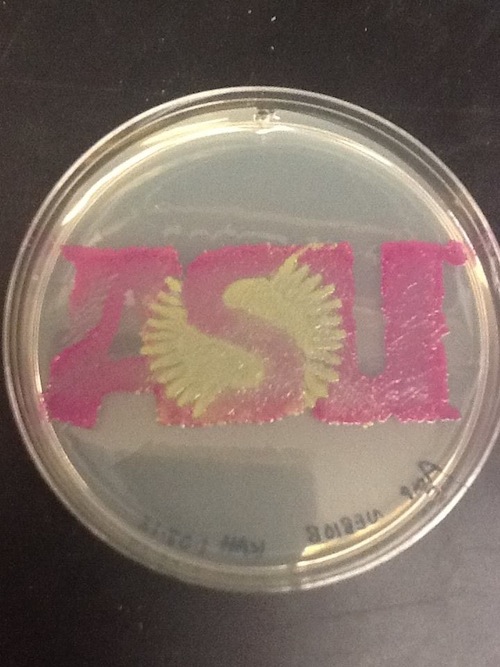
I used BBa_I13522 to demonstrate the expression of GFP in E. coli for a biomedical engineering course at Arizona State University.
- Cells: NEB10B
- Medium: LB Lennox agar, 100 ug/mL ampicillin
- The read area contains bacteria expressing RFP from BBa_J04450
iGEM team Wageningen UR
This brick was used to create GFP with a His tag at the C-terminus and the E-coil at the N-terminus to use it as reporter for the E- and K-coil [http://2012.igem.org/Team:Wageningen_UR/Coil_system Plug and Apply sytem]
see BBa_K883701; BBa_K883700; BBa_K883702 and BBa_K883703
User Reviews
UNIQd8b8554e6309d4ff-partinfo-00000002-QINU
|
Antiquity |
This review comes from the old result system and indicates that this part did not work in some test. |
|
•••••
|
We obtained this part from the Spring 2011 distribution plate. It expresses wonderfully in NEB10B cells from New England Biolabs. It does not express well in the fast-growing strain DH5-alpha Turbo, which we've found doesn't express transgenes very well in general. |
UNIQd8b8554e6309d4ff-partinfo-00000005-QINU