Difference between revisions of "Part:BBa K530000"
(→Characterization of performance) |
(→Sequencing) |
||
| Line 30: | Line 30: | ||
===Sequencing=== | ===Sequencing=== | ||
| − | [[Image:CrtYB Sequencing.png]] | + | [[Image:CrtYB Sequencing 2.png]] |
This is the sequencing for colony 2. Colony 2 had areas of poor match between both forward and reverse regions. We attribute this to the fact that our gene is far too big for two primers to cover on their own. The middle regions are due to this, and the initial and final regions are due to the trace taking some time to start up. | This is the sequencing for colony 2. Colony 2 had areas of poor match between both forward and reverse regions. We attribute this to the fact that our gene is far too big for two primers to cover on their own. The middle regions are due to this, and the initial and final regions are due to the trace taking some time to start up. | ||
Revision as of 21:06, 28 October 2011
CRTYB
Enzyme in the pathway required for B-Carotene Synthesis. This enzyme is a combination, it is the full Phytoene Synthase enzyme spliced with a Lycopene B-Cyclase enzymatic domain. This sequence was taken from a WT strain of xanthophyllomyces dendrorhous. It catalyzes both the conversion of Geranylgeranyl diphosphate to Phytoene and Lycopene to B-Carotene.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 1068
Illegal BsaI.rc site found at 1399
Plasmid Map
Characterization of performance
Below we can see the beta-carotene production levels on a per cell basis for different combinations of parts involved in beta-carotene synthesis. These are compared with a wild type control. tHMG1 is used to funnel more initial substrate into the pathway and although useful, was not submitted as a part. 2I indicates that two copies of crtI were used.
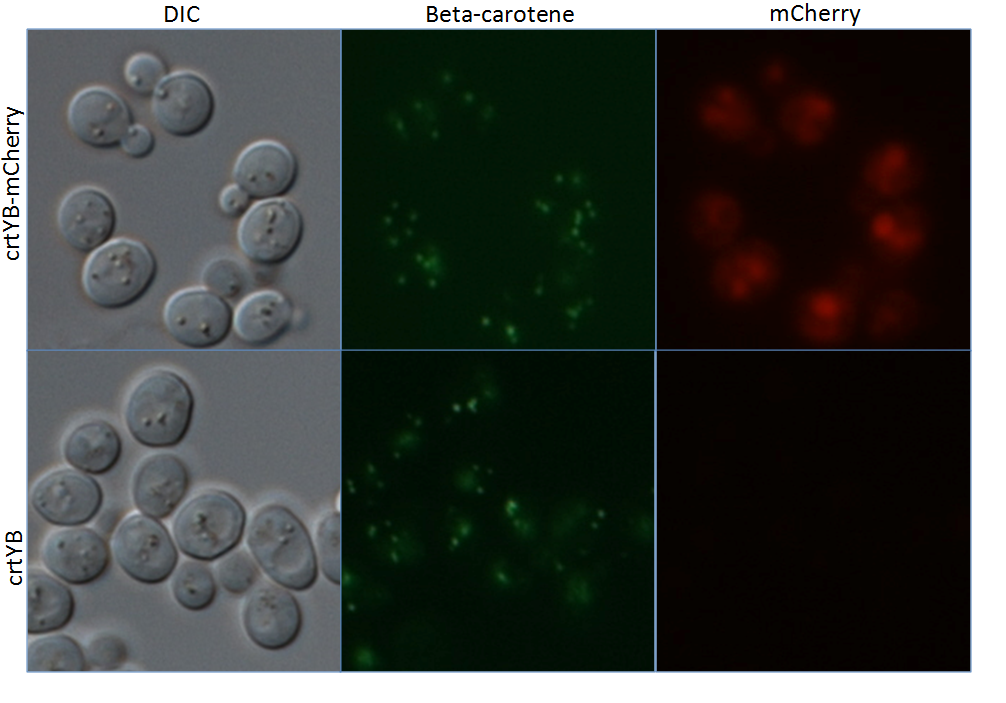
Localization of crtYB-mCherry and beta-carotene in yeast
Yeast cells, previously engineered to express all genes in the beta carotene biosynthetic pathway were subsequently transformed with a copy of crtYB C-terminally tagged with mCherry. The subcellular localization pattern of mCherry (crtYB-mCherry) was then assessed by fluorescence microscopy as compared to the parental, untagged control strain (crtYB) under identical conditions. Autofluorescence of beta carotene using a filter set for fluorescein fluorescence was also performed. Yeast cells are shown using differential interference contrast (DIC). Microscopy was performed at 1000X magnification.
Sequencing
This is the sequencing for colony 2. Colony 2 had areas of poor match between both forward and reverse regions. We attribute this to the fact that our gene is far too big for two primers to cover on their own. The middle regions are due to this, and the initial and final regions are due to the trace taking some time to start up.