Difference between revisions of "Part:BBa K4274009"
| Line 13: | Line 13: | ||
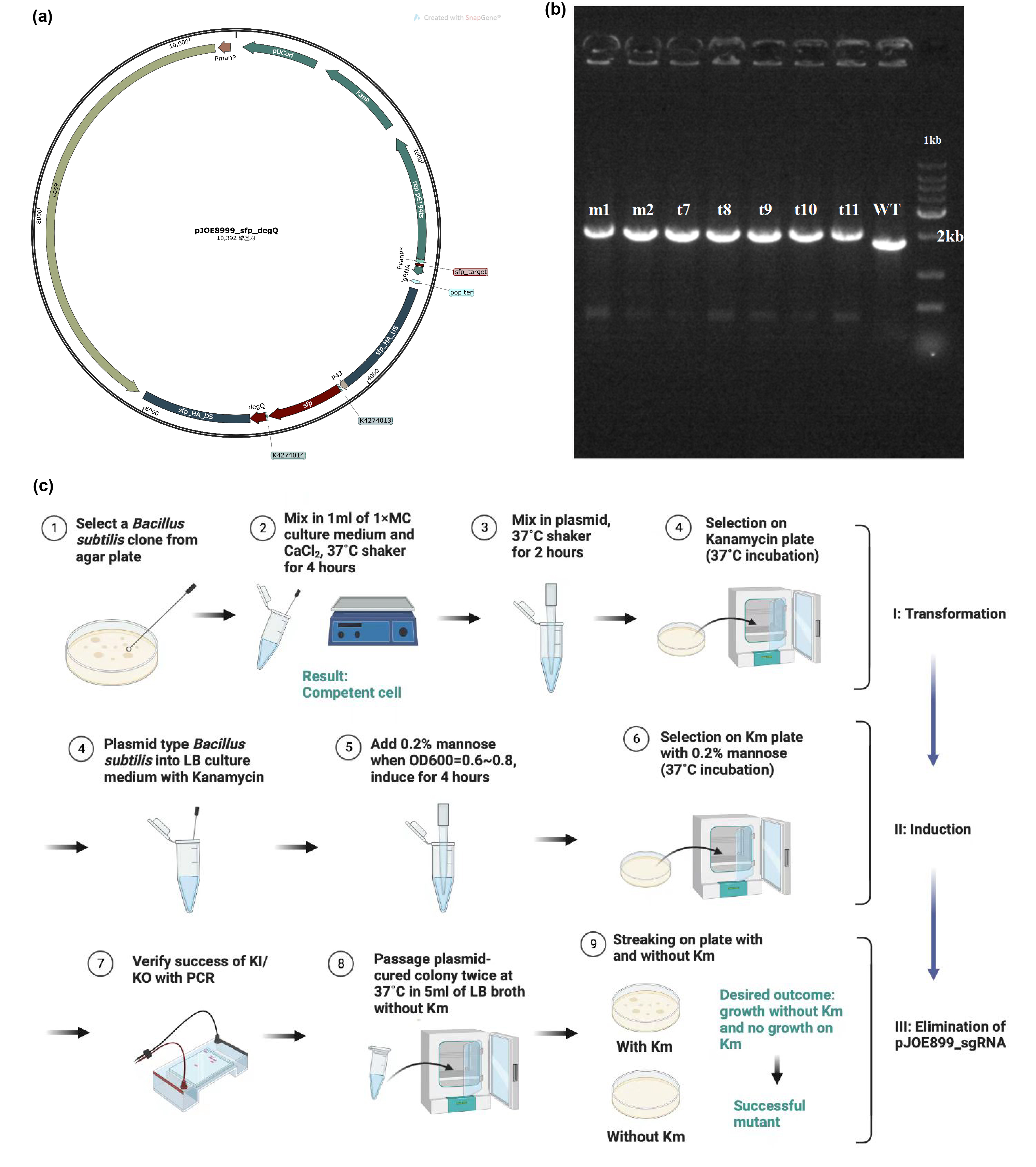
Edting the genome of <i>Bacillus subtilis</i> 168 to enable it to produce fengycins. (a) Plasmid design for knocking out the invalid sfp gene of <i>Bacillus subtilis</i> 168 and knocking in the sfp gene from B. amyloliquefaciens FZB42 and degQ gene from <i>Bacillus subtilis</i> 168. (b) Electrophoresis results show that gene editing is successful. (c) Protocol about transformation, induction and elimination of pJOE8999 plasmid in <i>Bacillus subtilis</i> 168. ]] | Edting the genome of <i>Bacillus subtilis</i> 168 to enable it to produce fengycins. (a) Plasmid design for knocking out the invalid sfp gene of <i>Bacillus subtilis</i> 168 and knocking in the sfp gene from B. amyloliquefaciens FZB42 and degQ gene from <i>Bacillus subtilis</i> 168. (b) Electrophoresis results show that gene editing is successful. (c) Protocol about transformation, induction and elimination of pJOE8999 plasmid in <i>Bacillus subtilis</i> 168. ]] | ||
| − | [[Image:Parts-keystone- | + | [[Image:Parts-keystone-fengycin2.tiff|thumbnail|750px|center|'''Figure 2:''' |
Quantification analysis of fengycins production. (a) The sequencing result of engineering <i>Bacillus subtilis</i> 168 strain for production. (b) The standard curve of peak area obtained by HPLC (y) and its corresponding fengycins' concentration (x). (c) HPLC results of sample and various concentrations of fengycins standard. ]] | Quantification analysis of fengycins production. (a) The sequencing result of engineering <i>Bacillus subtilis</i> 168 strain for production. (b) The standard curve of peak area obtained by HPLC (y) and its corresponding fengycins' concentration (x). (c) HPLC results of sample and various concentrations of fengycins standard. ]] | ||
==Source== | ==Source== | ||
Revision as of 08:17, 12 October 2022
sfp
sfp gene is a biobrick part from B. amyloliquefaciens FZB42 encoding 4'-phosphopantetheinyl transferase which functions as a primer of nonribosomal peptide synthesis via phosphopantetheinylation of thiotemplates. However, compared to the sfp gene in Bacillus subtilis 168, they only had an amino acid homology of 70%. And the mutation existed in Bacillus subtilis 168 resulted in its unable to produce any lipopeptide biosurfactant. Hence, to allow B. subtilis to produce fengycins, it is essential to first knock-out the mutant gene and then knock-in the correct sfp (Part: BBa_K4274009) and degQ (Part: BBa_K4274010).
Usage and Biology
Our sfp gene (Gene ID: 45022253) is a biobrick part derived from B. amyloliquefaciens FZB42 encoding 4'-phosphopantetheinyl transferase. As mentioned above, the correct sfp gene is the core of lipopeptide biosurfactant production, thus the mutant sfp gene in B. subtilis is invalid for producing fengycins.
To construct a strain of B. subtilis to produce fengycins, we will use sfp_target (gRNA) (part: BBa_K4274008) to target the region of mutant sfp in B. subtilis for knock-out, simultaneously sfp (Part: BBa_K4274009) and degQ (Part: BBa_K4274010) gene were knocked-in in situ. It was used in the composite part PvanP*-sfp_target-sfp_HA_US-p43-K4274013-sfp-K4274014-degQ-sfp_HA_DS (Part: BBa_K4274035) to realize fengycins’ production in B. subtilis.
Characterization
File:Parts-keystone-fengycin2.tiff
Source
B. amyloliquefaciens FZB42
Sequence and Features
- 10INCOMPATIBLE WITH RFC[10]Illegal PstI site found at 38
- 12INCOMPATIBLE WITH RFC[12]Illegal PstI site found at 38
- 21COMPATIBLE WITH RFC[21]
- 23INCOMPATIBLE WITH RFC[23]Illegal PstI site found at 38
- 25INCOMPATIBLE WITH RFC[25]Illegal PstI site found at 38
Illegal NgoMIV site found at 95 - 1000INCOMPATIBLE WITH RFC[1000]Illegal SapI.rc site found at 661
References
[1]Chen X.H., Koumoutsi A., Scholz R., et al. More than Anticipated – Production of Antibiotics and Other Secondary Metabolites by Bacillus amyloliquefaciens FZB42. Mircrobial Biotech. 16 (2), 14-24 (2009). https://doi.org/10.1159/000142891.
[2]Jin P., Wang H., Liu W., et al. Characteriztion of IpaH2 gene corresponding to lipopeptide synthesis in Bacillus amyloliquefaciens HAB-2. BMC Microbio. 17 (2), 227 (2017). https://doi.org/10.1186/s12866-017-1134-z.
[3]Tsuge K., Ano T., Hirai M., et al. The Genes degQ, pps, and Ipa-8(sfp) Are Responsible for Conversion of Bacillus subtilis 168 to Plipastin Production. Antimicrobial Agents and Chemo. 43(9), 2183-2192 (1999). https://doi.org/10.1128/AAC.43.9.2183.
