Difference between revisions of "Part:BBa K3739011"
| Line 23: | Line 23: | ||
===Characterization=== | ===Characterization=== | ||
====Characterization of signal peptides==== | ====Characterization of signal peptides==== | ||
| − | Since some proteins in our project are intended to function in extracellular environment, how to get out of the cells is vital for realization of their functions. Signal peptide is a short amino acid sequence on secreted protein, which is a natural helper of protein to get out of cytoplasm. However, signal peptides can lead protein go to not only the extracellular environment but also the periplasm and membrane. There are several signal peptides selected for ''V.natriegens'', but these are all directed into periplasm. <sup>1</sup> Although there is a signal peptide of alginate lyase from ''V.natriegens'' able to direct heterogenous protein to extracellular environment in E.coli chasis, It is not clear that there is a signal peptide conducting protein to extracellular environment in ''V.natriegens''. 2 | + | Since some proteins in our project are intended to function in extracellular environment, how to get out of the cells is vital for realization of their functions. Signal peptide is a short amino acid sequence on secreted protein, which is a natural helper of protein to get out of cytoplasm. However, signal peptides can lead protein go to not only the extracellular environment but also the periplasm and membrane. There are several signal peptides selected for ''V.natriegens'', but these are all directed into periplasm. <sup>1</sup> Although there is a signal peptide of alginate lyase from ''V.natriegens'' able to direct heterogenous protein to extracellular environment in E.coli chasis, It is not clear that there is a signal peptide conducting protein to extracellular environment in ''V.natriegens''. <sup>2<sup/> |
=====Selected procedure===== | =====Selected procedure===== | ||
Here, we use a tip to select the signal peptide. Download the translated CDS information of ''V.natriegens'' from NCBI Assembly. Through analysis of SignalP 5.0, we found out several candidate signal peptides which have significant likelihood to work with corresponding secretion system. According to the function of proteins containing proposed signal peptide, we can exclude the signal peptides targeting to the periplasm and cell membrane. Finally, we find out the best candidate signal peptide, LMT signal peptide, from our chassis | Here, we use a tip to select the signal peptide. Download the translated CDS information of ''V.natriegens'' from NCBI Assembly. Through analysis of SignalP 5.0, we found out several candidate signal peptides which have significant likelihood to work with corresponding secretion system. According to the function of proteins containing proposed signal peptide, we can exclude the signal peptides targeting to the periplasm and cell membrane. Finally, we find out the best candidate signal peptide, LMT signal peptide, from our chassis | ||
| Line 49: | Line 49: | ||
[[File:T--XMU-China--waimi-6.png|300px]]<br/> | [[File:T--XMU-China--waimi-6.png|300px]]<br/> | ||
The data shows that after the protein synthesis was inhibited, the change in the amount of secreted protein over time was well matched with the model. | The data shows that after the protein synthesis was inhibited, the change in the amount of secreted protein over time was well matched with the model. | ||
| + | ===Reference=== | ||
| + | 1. Eichmann, J.; Oberpaul, M.; Weidner, T.; Gerlach, D.; Czermak, P., Selection of High Producers From Combinatorial Libraries for the Production of Recombinant Proteins in Escherichia coli and Vibrio natriegens. ''Frontiers in Bioengineering and Biotechnology'' '''2019''', 7.<br/> | ||
| + | 2. Meng, Q.; Tian, X.; Jiang, B.; Zhou, L.; Chen, J.; Zhang, T., Characterization and enhanced extracellular overexpression of a new salt‐activated alginate lyase. Journal of the Science of Food and Agriculture 2021.<br/> | ||
| + | 3. Haitjema, C. H.; Boock, J. T.; Natarajan, A.; Dominguez, M. A.; Gardner, J. G.; Keating, D. H.; Withers, S. T.; Delisa, M. P., Universal Genetic Assay for Engineering Extracellular Protein Expression. ''ACS Synthetic Biology'' '''2014''', 3 (2), 74-82.<br/> | ||
| + | |||
<!-- --> | <!-- --> | ||
Latest revision as of 03:16, 22 October 2021
LMT-his-CBM-hutH
LMT here represents a signal peptide used to secrect the fusion protein outside the cell. The fusion protein CBM-hutH works on the surface of single cell Phaeocystis. globosa and the enzyme catalyzes the reaction of converting histidine to form toxic urocanic acid. His-tag is added to purify the protein.
Biology
LMT
Lytic murein transglycosylase (LMT) is an enzyme which is able to degrade murein, a component of cell wall of bacteria. There are two kind of LMTs existing in E.coli: the membrane-binding one and the soluble one. The gene coding LMT homolog is also incorporated in the genome of Vibrio natriegens. The LMT signal peptide (named LMT in our parts) is from the LMT homolog, which can lead the fused protein secreted out of Vibrio natriegens.
CBM
Cellulose enzymes have two domains, and the one that helps bind to cellulose is called cellulose binding module (CBM), and therefore it helps our fusion protein bind to cellulose-rich cell wall. Here we choose the CBM of CenA from Cellulomonas fimi, which has been successfully expressed in Escherichia coli.
hutH
The HutH comes from Pseudomonas putida. Under natural conditions, many microorganisms can use the histidine ammonia-lyase (HutH) to change L-histidine into urocanic acid. HutH catalyzes the first step in the degradation of histidine, and the product urocanic acid is further metabolized to glutamate. This enzyme could be found in the liver of vertebrates and in bacteria such as Escherichia coli, Salmonella and Pseudomonas. It is specific for L-histidine and can be inhibited by D-histidine or imidazole. The active center of the enzyme is thought to be dehydroalanine.
Usage
Used to construct the composite part BBa_K3739039 and BBa_K3739109.
Characterization
Characterization of signal peptides
Since some proteins in our project are intended to function in extracellular environment, how to get out of the cells is vital for realization of their functions. Signal peptide is a short amino acid sequence on secreted protein, which is a natural helper of protein to get out of cytoplasm. However, signal peptides can lead protein go to not only the extracellular environment but also the periplasm and membrane. There are several signal peptides selected for V.natriegens, but these are all directed into periplasm. 1 Although there is a signal peptide of alginate lyase from V.natriegens able to direct heterogenous protein to extracellular environment in E.coli chasis, It is not clear that there is a signal peptide conducting protein to extracellular environment in V.natriegens. 2<sup/>
Selected procedure
Here, we use a tip to select the signal peptide. Download the translated CDS information of V.natriegens from NCBI Assembly. Through analysis of SignalP 5.0, we found out several candidate signal peptides which have significant likelihood to work with corresponding secretion system. According to the function of proteins containing proposed signal peptide, we can exclude the signal peptides targeting to the periplasm and cell membrane. Finally, we find out the best candidate signal peptide, LMT signal peptide, from our chassis
Qualification results
To clarify whether the signal peptides we selected are able to direct the fused proteins to extracellular environment, we constructed Aly01 and LMT single peptides with BBa_K2560063 which is from 2018 Marhurg iGEM program and transformed the pET28a(+) plasmid with composite parts BBa_K3739038 and BBa_K3739041 into V.natriegens. Then we use the anti-GFP antibody to detect secreted GFP in the filtered supernatant of V.natriegens liquid culture by western blotting after V.natriegens is cultivated for .16 hours. The concentration of his-GFP in the supernatant of BBa_K3739041 is significantly higher than that in J23100-B0030-GFP. (Fig. 1A, 1B) Furthermore, less expression of BBa_K3739041 than that of GFP conforms that LMT-his-GFP is secreted rather than from lysed cells. (Fig. 1C) However, the concentration of Aly01-his-GFP is less than that of GFP. (Fig. 1A, 1B) It is probably because that there are less expression of Aly01-his-GFP in V.natriegens. (Fig. 1C) Thus, we use another method to prove our hypothesis.
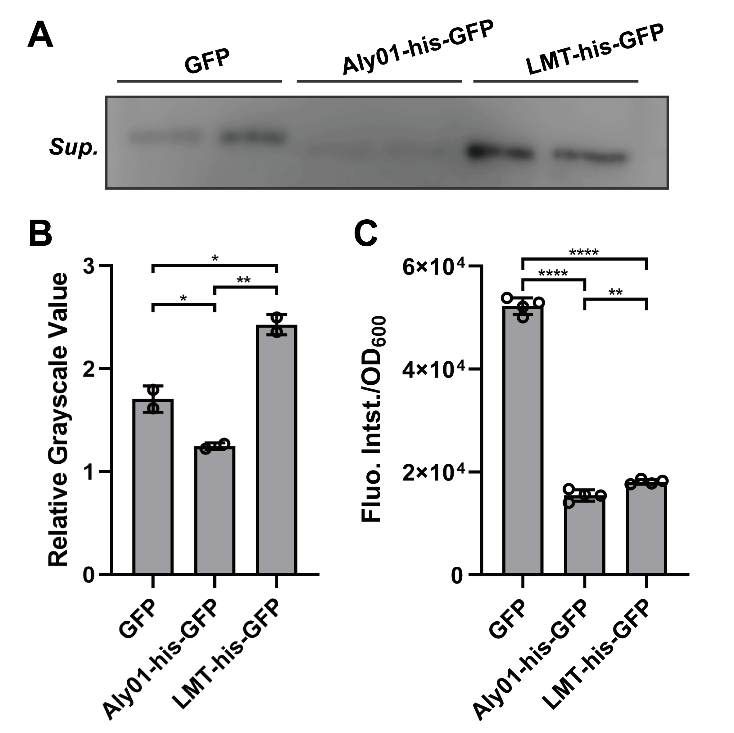
Fig. 1. (A) Western Blotting detection of the V.natriegens supernatants of J23100-B0030-GFP, BBa_K3739038 and BBa_K3739041 after 16h cultivation in LBv2 liquid culture, 37℃. Sup. means supernatant. (B) Grayscale analysis of (A). (C) Fluorescence intensity/OD600 value of V.natriegens strain of J23100-B0030-GFP, BBa_K3739038 and BBa_K3739041 in LBv2 liquid culture after 16h cultivation in 37℃.
Quantitation results
In order to exclude the difference of expression among J23100-B0030-GFP, BBa_K3739038 and BBa_K3739041 and quantify secretion effect of signal peptide, we treated V.natriegens liquid cultures after 12.5 h cultivation in 37℃ with 12.5 μg/mL chloramphenicol. Then we sampled the cultures every 6 minutes and terminated secretion with 20 mM NaN3. As time goes, the his-GFP concentration of culture supernatant were increased in both BBa_K3739038 and BBa_K3739041. This is indicated that both LMT and Aly01 signal peptide are able to translocate fused protein to extracellular environment. (Fig. 2A, 2B) We made a grayscale analysis for the modelling of secretion, and the total GFP is detected for confirmation that the translation is completely repressed. (See our Model https://2021.igem.org/Team:XMU-China/Model pages)
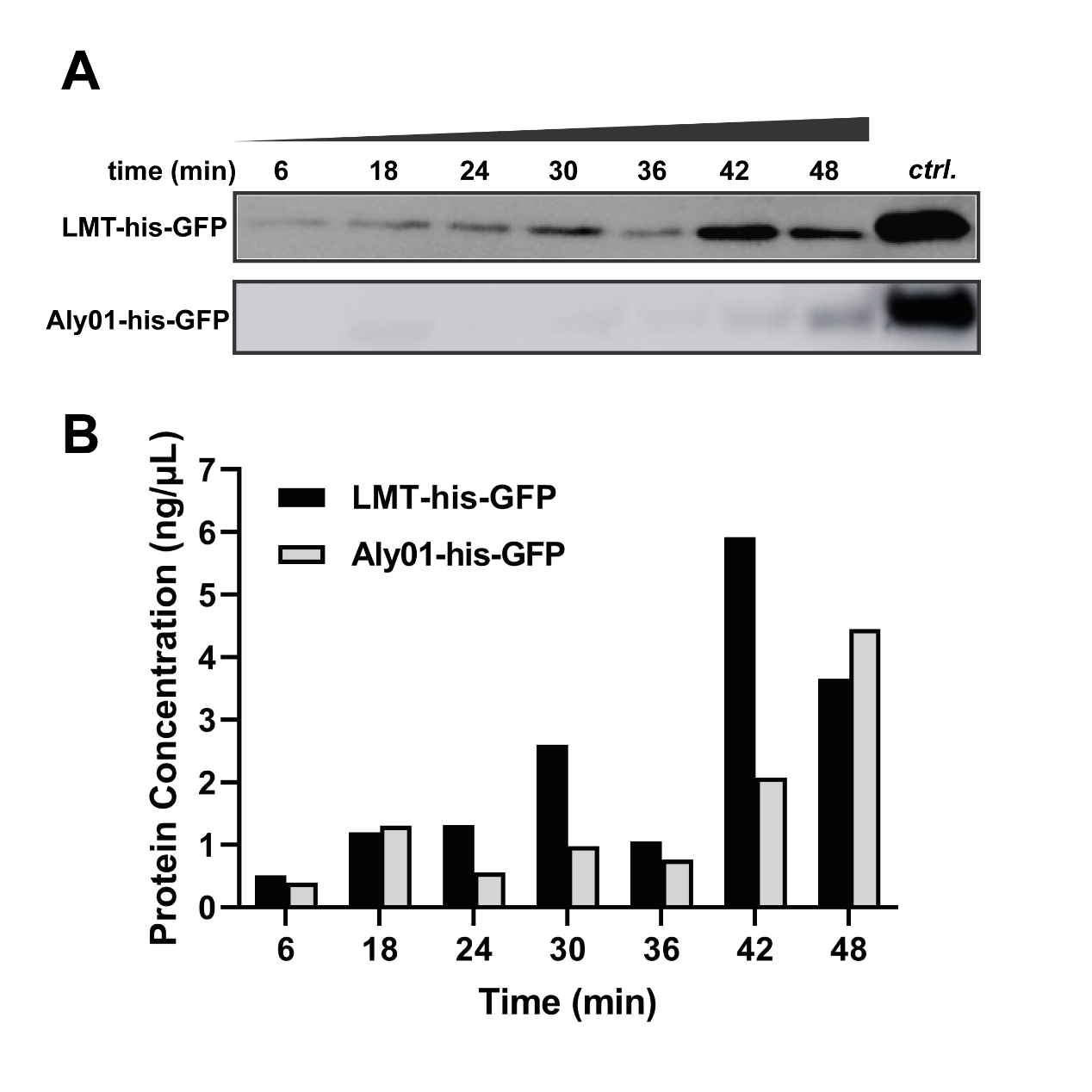
Fig. 2. (A) Western blotting detection of sampled supernatants treated with chloramphenicol in different times. (B) GFP concentration of treated supernatant of BBa_K3739038 and BBa_K3739041 in (A)
Secretion in E.coli
Besides conducting protein secretion in V.natriegens, Aly01 signal peptide is reported that it is able to conduct protein secretion in E.coli as well. 2We supposed that LMT signal peptide may also have conducting function in E.coli. Thus, we used filtered culture of BL21 strain transformed with pET28a(+)-BBa_K3739043 and pET28a(+)-BBa_K3739041 for SDS-PAGE. Then the SDS-PAGE gel is stained by silver staining. Contrasting to BBa_K3739043, his-GFP is detected in the supernatant of pET28a(+)-BBa_K3739041.(Fig. 3) It is indicated that LMT signal peptide can also work for conducting ====secretion in E.coli====
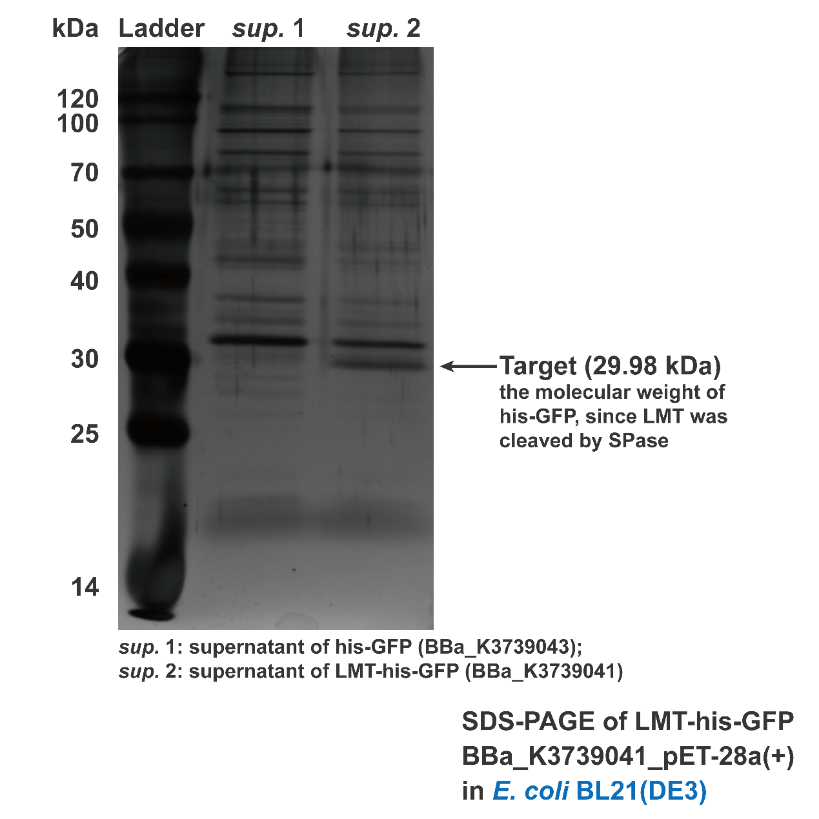
Fig. 3. Silver staining result of supernatant of BL21(DE3) transformed with BBa_K3739043 and BBa_K3739041, respectively.
Then we used an alternative way to verify the secretion function of signal peptides in E.coIi. The proteins labeled of tetracysteine motif tag (FlAsH tag, FT in short) could be detected with the biarsenical compound FlAsH-EDT2. 3
To quantify and qualify the secretion effect, FT was added into the C-terminal of BBa_K3739000, BBa_K3739006 and BBa_K3739009. Composite parts BBa_K3739084, BBa_K3739085, BBa_K3739087 were constructed (Fig. 4) on plasmid backbone pSB1C3 or pET-28a(+) and then the recombinant plasmid was transformed into E. coli BL21 (DE3). Positive colonies selected by colony PCR and sequencing were cultivated for secretion analysis.
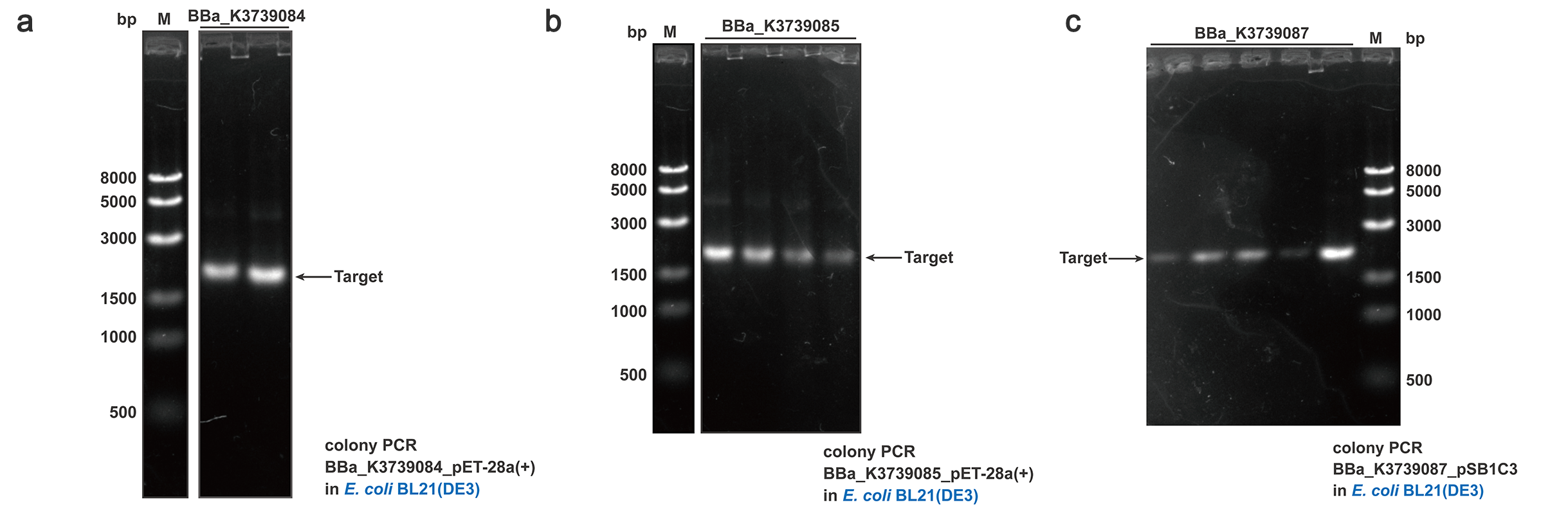
Fig. 4. Colony PCR results of BBa_K3739084, BBa_K3739085, BBa_K3739087
After cultivation for 5 hours at 37℃ and induction for 2 hours with 0.1 mM IPTG, the cell supernatant was separated by centrifugation. The supernatant along with 2 μM FlAsH-EDT2 and 1 mM DTT were added to wells in a 96-well plate. Following incubation in the dark for 1 h at 37 °C, fluorescence was measured by 503 nm excitation and 528 nm emission, and each value was normalized by OD600. The result shows that the fluorescence intensity/OD600 of the group Aly01 and LMT is significantly higher than native control group, which proves that these 2 signal peptides can function well. (Fig. 5)
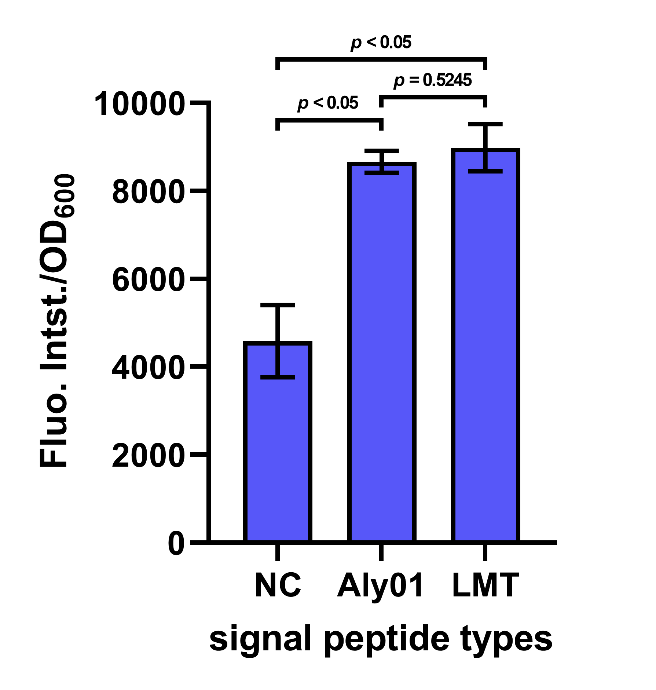
Fig. 5. Fluorescence intensity/OD600 of cell supernatant after incubation in the dark for 1h.
Moreover, to promote our understanding on secretion process mediated by signal peptides, we built a model about relationship between the amount of secreted recombinant protein and time. (See our Model https://2021.igem.org/Team:XMU-China/Model pages) In order to get the value of P_T (the total amount of recombinant protein per unit volume of reactor), P_M (the amount of the recombinant protein in the medium compartment in unit volume) and γ(extracellular secretion rate constant), a sets of characterization experiments were done.
BBa_K3739087 were constructed on plasmid backbone pET-28a(+) and then the recombinant plasmid was transformed into E. coli BL21 (DE3). After cultivation for 5 hours and induction for 3 hours with 0.1 mM IPTG, 50mg/L chloroamphenicol was added to bog down the synthesis of protein in the engineered bacteria in order to make the experimental operations more convenient. Then, the cell supernatant was collected in 0, 6, 12, 18, 24, 30, 40 min to test the amount of secreted recombinant protein by fluorescence detection of FlAsH-EDT2 and FT. (Fig. 6)
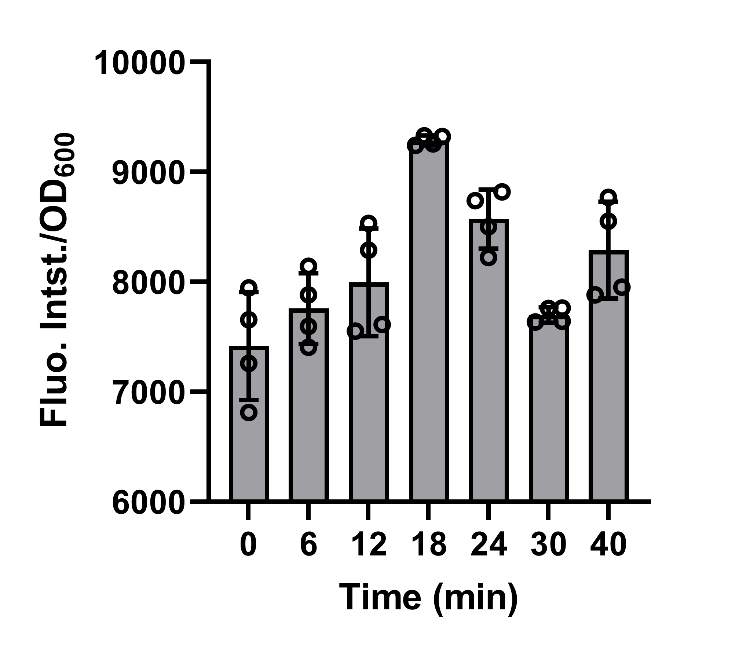
The data shows that after the protein synthesis was inhibited, the change in the amount of secreted protein over time was well matched with the model.
Reference
1. Eichmann, J.; Oberpaul, M.; Weidner, T.; Gerlach, D.; Czermak, P., Selection of High Producers From Combinatorial Libraries for the Production of Recombinant Proteins in Escherichia coli and Vibrio natriegens. Frontiers in Bioengineering and Biotechnology 2019, 7.
2. Meng, Q.; Tian, X.; Jiang, B.; Zhou, L.; Chen, J.; Zhang, T., Characterization and enhanced extracellular overexpression of a new salt‐activated alginate lyase. Journal of the Science of Food and Agriculture 2021.
3. Haitjema, C. H.; Boock, J. T.; Natarajan, A.; Dominguez, M. A.; Gardner, J. G.; Keating, D. H.; Withers, S. T.; Delisa, M. P., Universal Genetic Assay for Engineering Extracellular Protein Expression. ACS Synthetic Biology 2014, 3 (2), 74-82.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 597
Illegal NgoMIV site found at 1033
Illegal NgoMIV site found at 1768
Illegal NgoMIV site found at 2004
Illegal AgeI site found at 343
Illegal AgeI site found at 433
Illegal AgeI site found at 670
Illegal AgeI site found at 1497
Illegal AgeI site found at 1993 - 1000COMPATIBLE WITH RFC[1000]
