Difference between revisions of "Part:BBa K3584000"
(→Contribution) |
|||
| Line 8: | Line 8: | ||
===Contribution=== | ===Contribution=== | ||
| − | + | BBa_K3584000 is the coding sequence of a polyphenol oxidase (PPO) from Pleurotus ostreatus, which is different from the present enzymes showing the similar activity. PPO is a group of enzyme that are able to oxidize phenol or polyphenol by molecular oxygen to form the corresponding quinone. In a broad sense, polyphenol oxidase can be divided into three categories: monophenol monooxidase (tyrosinase, EC.1.14.18.1), bisphenol oxidase (catechol oxidase, EC. 1.10.3.2) and laccase (laccase, EC.1.10.3.1). Among these three types of polyphenol oxidase, catechol oxidase is mainly distributed in plants, and the polyphenol oxidase in microorganisms mainly includes laccase and tyrosinase. The polyphenol oxidase mentioned in most of the literature is generally the collective name of catechol oxidase and laccase. | |
[[File:T--Shanghai United--BBa K35284000 fig 0.jpg|500px|thumb|center|Figure 0]] | [[File:T--Shanghai United--BBa K35284000 fig 0.jpg|500px|thumb|center|Figure 0]] | ||
| − | |||
===Engineering Success=== | ===Engineering Success=== | ||
Latest revision as of 13:45, 27 October 2020
polyphenol oxidase (PPO)
BBa_K3584000 is a protein domain of polyphenol oxidase (PPO).
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 1242
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 194
- 1000COMPATIBLE WITH RFC[1000]
Contribution
BBa_K3584000 is the coding sequence of a polyphenol oxidase (PPO) from Pleurotus ostreatus, which is different from the present enzymes showing the similar activity. PPO is a group of enzyme that are able to oxidize phenol or polyphenol by molecular oxygen to form the corresponding quinone. In a broad sense, polyphenol oxidase can be divided into three categories: monophenol monooxidase (tyrosinase, EC.1.14.18.1), bisphenol oxidase (catechol oxidase, EC. 1.10.3.2) and laccase (laccase, EC.1.10.3.1). Among these three types of polyphenol oxidase, catechol oxidase is mainly distributed in plants, and the polyphenol oxidase in microorganisms mainly includes laccase and tyrosinase. The polyphenol oxidase mentioned in most of the literature is generally the collective name of catechol oxidase and laccase.
Engineering Success
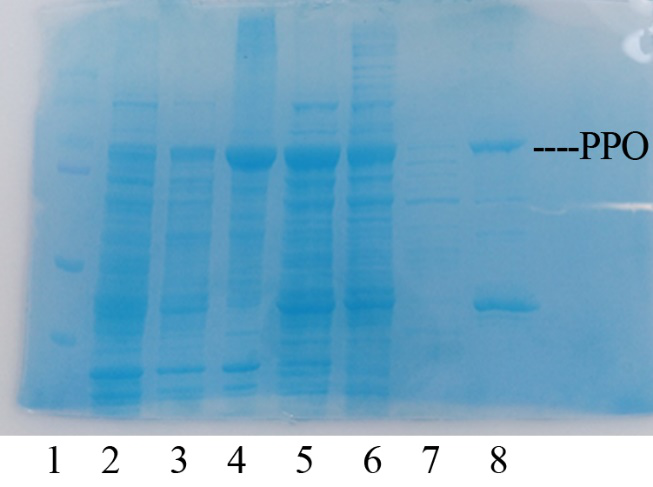
Production, purification, and SDS-PAGE analysis of recombinant PPO In order to present the function of the part, the PPO gene was expressed in E. coli BL21 (DE3) under the control of T7 promoter. Then the bacterial cells are collected and crushed, and the PPO enzyme solution is purified by further confirmation by the SDS-PAGE method, which is found in the corresponding protein band of approximately 57 kDa (Figure 1).
Figure 1. Production, purification, and SDS-PAGE analysis of recombinant PPO. Lane: protein molecular weight standard; lane 8: purified PPO.
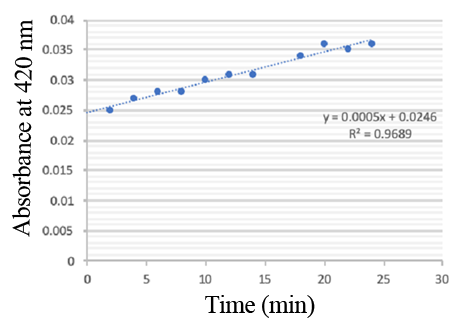
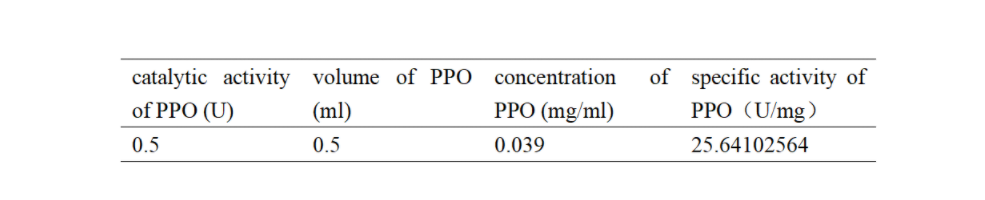
Enzymatic activity of the PPO PPO activity was determined using an analogue of p-cresol (catechol) as a substrate. In proper reaction mixture, PPO and 10 mM catechol was incubated at 50 ℃, pH 7.0 for 3 min, and the change in absorbance at 420 nm was measured spectrophotometrically. One unit of enzyme (PPO) activity was defined as the amount of enzyme that increased absorbance of 0.001 per minute. We found that as the extension of reaction time, reaction liquid of absorbance at 420 nm increased linear (Figure 2), we obtained by catalytic activity of PPO enzyme fluid, and divided by the corresponding protein concentration, the specific activity of PPO (Table 1).
Figure 2. PPO activity was determined using catechol as a substrate. The reaction mixture contained PPO protein and 10 mM catechol which was incubated at 50 ℃, pH 7.0 for 3 min, and the change in absorbance at 420 nm was recorded.
Table 1. Specific activity of recombinant PPO
Improvement
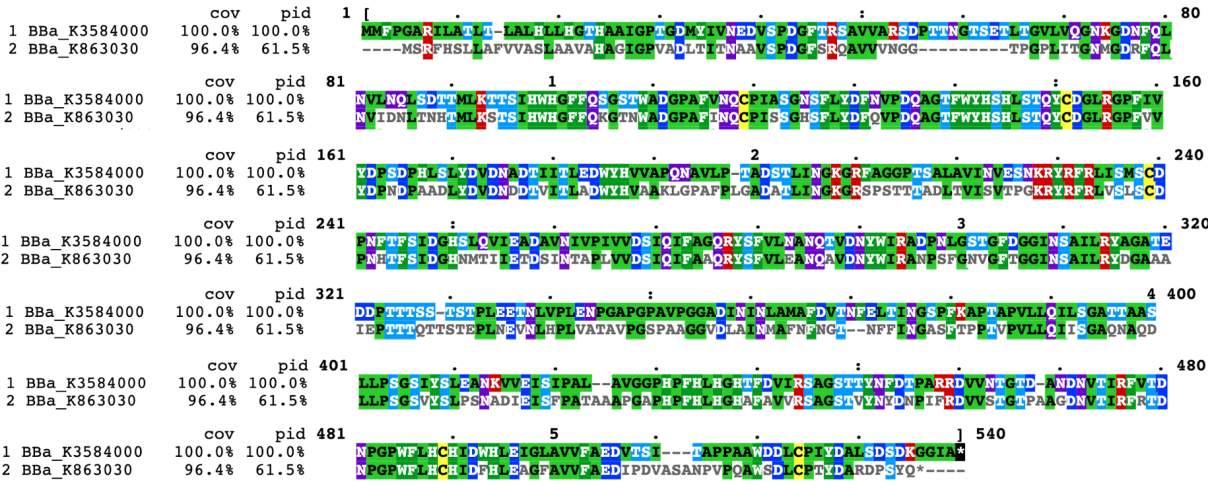
Compared with the old part BBa_K863030, expressing laccase (EC1.10.3.2), we design a new part BBa_K3584000, expressing polyphenol oxidase (EC 1.14.18.1, PPO). Both of the two enzymes can catalyze phenol or polyphenol to the corresponding quinone. The team iGEM12_Bielefeld-Germany aimed to use laccase to degrade polycyclic aromatic hydrocarbons for filtering populated water. They successfully identified one of the laccases, part BBa_K863030. They purified and detected the activity of this laccase protein.
In contrast, our team aimed to degrade p-cresol sulfate, on the basis of which we hope to provide chronic kidney disease patients with potential health supplements or even treatment. We chose PPO, whose sequence is different from laccase(Fig1), to construct our desired engineering strain.
Initially, we constructed a composite part BBa_K3584002 and transformed it into E.coli BL21(DE3). We successfully expressed and purified PPO protein, and detected its enzyme activity.
Furthermore, in order to optimize the function of BBa_K3584000, we constructed an improved composite part BBa_K3584008, which was designed as tyrosine-induced regulation system. This system allowed us to control the expression of PPO by adding or removing tyrosine.
In addition, we select a probiotics E.coli Nissle 1917 as the host. Therefore, our engineering strain would be a competitive health supplement or drug candidate for degrading p-cresol and hence bring about benefits to chronic kidney patients.