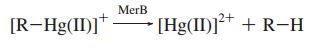Difference between revisions of "Part:BBa K1420002"
| Line 59: | Line 59: | ||
==MIT MAHE 2020== | ==MIT MAHE 2020== | ||
| − | ''' | + | '''Usage and Biology''' |
The merB gene is expressed from the mer operon of all bacteria resistant to organomercurial compounds, but it occurs more commonly in gram-positive bacteria than in gram-negative bacteria.9 The first reports of bacterial degradation of organomercurial compounds were from the gram-negative mercury-resistant Pseudomonas strain K62. Subsequent studies demonstrated that this particular strain contained two distinct forms of the MerB protein and both forms were capable of degrading a broad range of organomercurial compounds. This is a highly conserved sequence and the most significant differences occur at the amino-terminal end of the protein. | The merB gene is expressed from the mer operon of all bacteria resistant to organomercurial compounds, but it occurs more commonly in gram-positive bacteria than in gram-negative bacteria.9 The first reports of bacterial degradation of organomercurial compounds were from the gram-negative mercury-resistant Pseudomonas strain K62. Subsequent studies demonstrated that this particular strain contained two distinct forms of the MerB protein and both forms were capable of degrading a broad range of organomercurial compounds. This is a highly conserved sequence and the most significant differences occur at the amino-terminal end of the protein. | ||
Revision as of 10:39, 20 October 2020
merB, Organomercurial Lyase from Serratia marcescens
Summary
• MerB is a lyase that catalyzes the breaking of carbon-mercury bonds through protonolysis of toxic mercury compounds, such as methylmercury. This produces the less toxic and less mobile Hg2+ which is then completely volatilized to Hg0 by the enzyme MerA.
• MerB can catalyze the protonolysis of carbon mercury bonds for a number of organomercurials, accelerating the rate of this reaction up to 107 times the rate of spontaneous abiotic decay.
• In our methylmercury degradation activity assay, we observed a significant increase in the rate degradation of methylmercury was seen in bacterial cultures containing pBBRBB::mer compared to pBBRBB::GFP control cells.
Overview
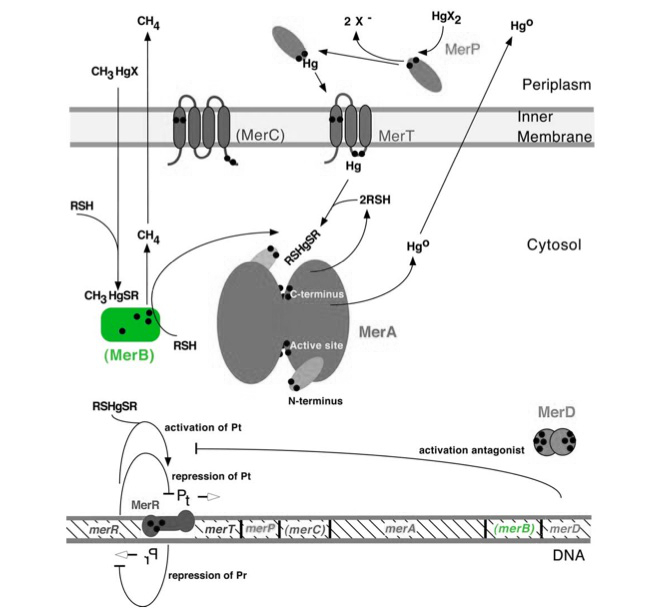
The gene merB (0.6Kb) encodes the carbon-mercury bond lyase MerB. MerB, along with its corresponding gene, are highlighted in green in Figure 1. The lower portion of Figure 1 shows the arrangement of mer genes in the operon, with merB located downstream of the transport and reductase genes, merP, merT and merA, respectively. Two mechanisms for mercury resistance are prevelant in nature. Resistance to inorganic Hg(II) is conferred by MerA (narrow spectrum) while additional resistance to organic mercury compounds (such as methylmercury) is conferred by addition of MerB (broad spectrum) to the operon.
Figure 1. Model of carbon-mercury lyase operon. The symbol • indicates a cysteine residue. RSH indicates cytosolic thiol redox buffers such as glutathione. Figure 1 shows the interactions of MerB, in green, with mercury compounds and other gene products of mer operon. (Figure adapted from "Bacterial mercury resistance from atoms to ecosystems." 1)
Molecular Function
The merB gene is often found immediately downstream of merA, and is essential for the detoxification and bioremediation of organic toxic mercury compounds in congruence with merA.1 MerB is a lyase that catalyzes the breaking of carbon-mercury bonds through protonolysis of toxic mercury compounds, such as methylmercury (Scheme 1).2 This produces the less toxic and less mobile Hg2+ which is then completely volatilized to Hg0 by the enzyme MerA.1
Scheme 1. MerB Catalyzed Reaction (R= alkyl, aryl)
Structure and Mechanism
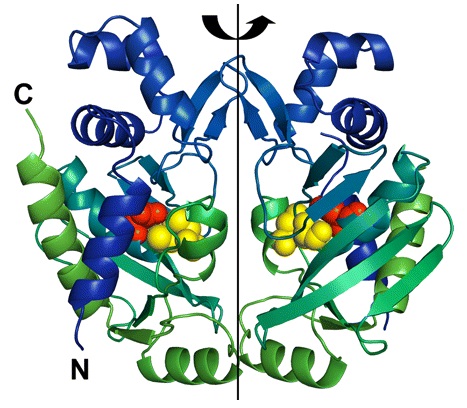
MerB forms a dimer by a 2-fold pseudosymmetry around the depicted rotation axis in Figure 2.3 The two subunits are color coded from blue to green distinguishing the amino-terminal end of the protein from the carboxy-terminal, respectively. Active site residues are color coded as well using van der Waals sphere representation: residues Cys-96 and Cys-159 are in yellow, and residue Asp-99 is in red. The structure of mercury-bound MerB and free MerB is extremely similar with the exception of the mercury ion.3 The mercury is bound to MerB by two sulfurs from the Cys-96 and Cys-159. An oxygen from a water molecule is also involved, binding to the mercury atom.3 On the other hand, the active site residues for the free MerB are completely inaccessible.3
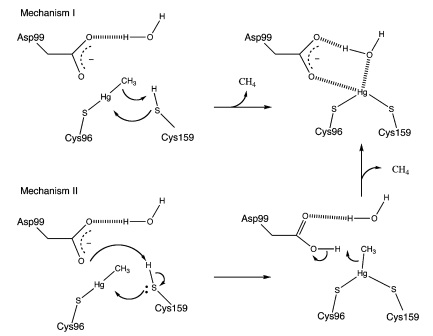
MerB can catalyze the protonolysis of carbon mercury bonds for a number of organomercurials, accelerating the rate of this reaction up to 107 times the rate of spontaneous abiotic decay.4 Two proposed mechanisms for how this is accomplished is displayed in Figure 3.4 Both mechanisms assume that the methylmercury substrate forms an initial bond with Cys96 and that the Cys159 site is protonated. In Mechanism 1, Cys159 protonates the methyl carbon and forms a covalent bond with Hg(II). In Mechanism 2 however, Cys159 donates a proton to Asp99 before forming a bond with Hg(II). Asp99 is then responsible for protonating the methyl group.
Figure 2. Structure of MerB enzyme. MerB is a dimer with 2-fold pseudosymmetry. The amino-terminal and carboxy-terminal are color coded blue and green, respectively. In addition, the active site residues Cys-96 and Cys-159 are in yellow, and residue Asp-99 is in red. (Figure adapted from "Enzyme Catalysis and Regulation: Crystal Structures of the Organomercurial Lyase MerB in Its Free and Mercury-bound Forms: Insights into the Mechanism of Methylmercury Degredation." 3)
Figure 3. Proposed mechanisms of MerB catalysis for the Hg-C protonolysis reaction. Both of these proposed mechanisms display how methyl group of methylmercury is protonated, producing Hg(II). Mechanism 2 differs from Mechanism 1 by utilizing Asp99 for protonation of the methyl group with the hydrogen attached to the sulfur from Cys159. (Figure adapted from "Mechanism of Hg-C Protonolysis in the Organomercurial Lyase MerB." 4)
merB Experimental Results
The function of MerB is to convert methylmercury to Hg(II). MerB functionality was demonstrated in the context of the entire mer operon using both silica encapsulated Escherichia coli K12 and unencapsulated E. coli . Both encapsulated and unencapsulated E. coli K12 containing pBBRBB::mer were compared to three negative controls; encapsulated and unencapsulated E. coli K12 containing pBBRBB::gfp (vector control) and abiotic LB medium. Negative controls were used to determine the amount of methylmercury absorbed by beads and the abiotic degradation rate of methylmercury. The experiment was conducted by adding methylmercury chloride to LB at a starting concentration of 1 mg/L. Prior to addition of methylmercury, tubes were inoculated either with 0.5 mL of silica bead-encapsulated cells or inoculated to an OD600 = 0.08 with unencapsulated bacteria from LB overnights. Time points were then taken to determine a rate of degradation of methylmercury. Samples were diluted one-million fold and were analyzed using a Tekran model 2700 Automated Methyl Mercury Analyzer using EPA method 1630 without distillation. This is a highly sensitive and ultra-stable cold vapor atomic fluorescence spectrometry (CVAFS) Hg detector. All quality assurance and quality control measures were taken as outlined in EPA method 1630. All MeHg standards (ongoing precision recoveries) were within the acceptable range averaging 96%.(We were aided in methylmercury measurements by Sona Parska of Ed Nater's Lab - University of Minnesota) </p>
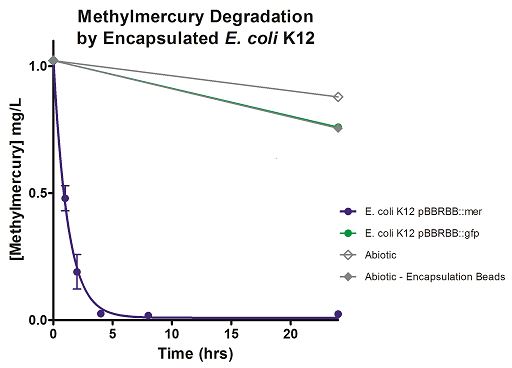
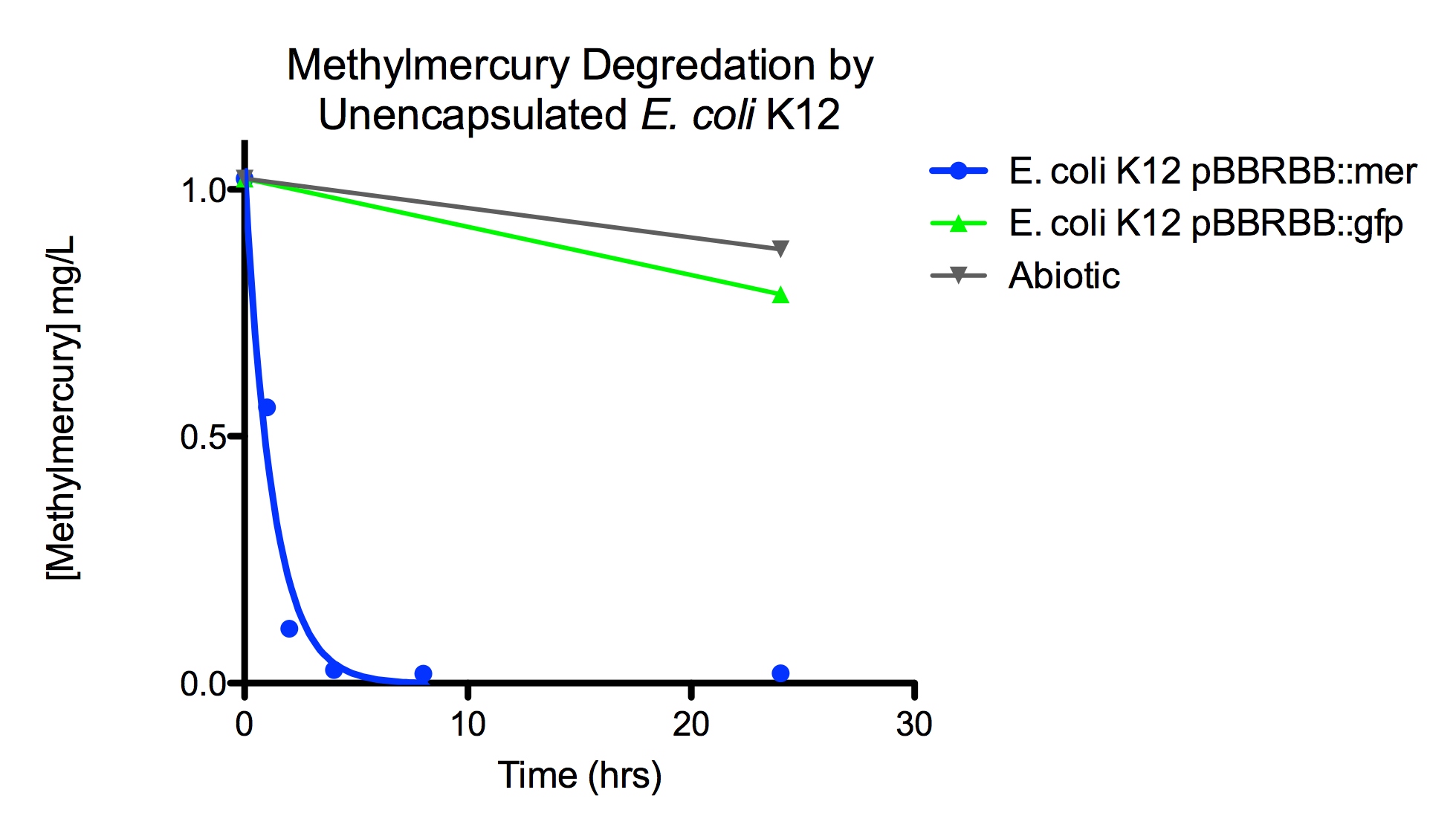
Degradation rates for methylmercury are depicted in Figure 4 for encapsulated bacteria and “Figure 5” using unencapsulated bacteria. The data show that only very small amounts of methylmercury are degraded in abiotic solutions over our time course. Data shows that the negative control, K12 strain containing pBBRBB::gfp, had a more significant decrease in concentration of methylmercury over time when compared to the abiotic control, but is still negligible in comparison to E. coli K12 containing pBBRBB::mer. The mer operon, is seen to remove methylmercury from the system almost completely after approximately 4 hours.At the 8 hour sampling point, methylmercury levels were below detection limits for the Tekran 2700.
From the data, it can be extrapolated that the presence of merB in our pBBRBB::mer construct facilitates the bioremediation and degradation of methylmercury. These results confirm the proposed mechanism of MerB and its function in the cleavage of carbon groups from methylmercury reducing it to Hg(II).In the future, we would like to express merB in trans on a plasmid with a different origin of replication than pBBRBB alongside pBBRBB containing the mer operon with merB deleted. This would allow us to determine if our part could complement a merB deletion strain, and hence test the MerB part more directly.
Figure 4. Monitoring the levels of mer operon-mediated methylmercury degradation. The graph displays that abiotic, abiotic with encapsulation beads, and the E. coli K12 strain containing the pBBRBB::gfp control degrade methylmercury from the environment at a much slower rate than the E. coli K12 containing the pBBRBB::mer, which can remove methylmercury from the system almost completely in approximately 4 hours.
Figure 5. Monitoring the levels of mer operon-mediated methylmercury degradation. The graph displays that abiotic and E. coli K12 strain containing the pBBRBB::gfp control degrade methylmercury from solution at much slower rates than the E. coli K12 containing the pBBRBB::mer, which can degrade 1 mg/L methylmercury to below detection limits after approximately 4 hours.
MIT MAHE 2020
Usage and Biology
The merB gene is expressed from the mer operon of all bacteria resistant to organomercurial compounds, but it occurs more commonly in gram-positive bacteria than in gram-negative bacteria.9 The first reports of bacterial degradation of organomercurial compounds were from the gram-negative mercury-resistant Pseudomonas strain K62. Subsequent studies demonstrated that this particular strain contained two distinct forms of the MerB protein and both forms were capable of degrading a broad range of organomercurial compounds. This is a highly conserved sequence and the most significant differences occur at the amino-terminal end of the protein.
The activity of MerB has been directly assayed by measuring either the release of the hydrocarbon products from the organomercurial or the generation of volatilized elemental mercury following the reduction of the ionic mercury product. Hydrocarbon product formation has been determined by a number of techniques including gas chromatographic analysis, UV analysis, and high-performance liquid chromatography (HPLC) analysis.
Paradoxically for a protonolysis catalyst, MerB’s pH optimum was >9. MerB has no bound cofactors, however, it has been shown that for full in vitro activity the enzyme requires the presence of thiols in solution, but the nature and concentration of the buffer thiols can have dramatically different effects. It is thought that the thiols play a fundamental role both in substrate binding and mercury release.
Relevant chemical model studies demonstrated a 1000-fold acceleration of aryl-Hg protonolysis by a compound capable of biscoordination, whereas a monothiol reagent provided only a 50-fold rate acceleration, suggesting that in MerB two protein thiol groups might be involved in stabilizing a reaction intermediate.
Interestingly, in many occurrences, the operon bearing the merB gene in broad-spectrum-resistant Enterobacteriaceae has undergone deletions of varying lengths removing most of the intervening genes between merB and the operonic promoter resulting in the merB gene being much closer to that promoter. In these internally deleted operons, the associated merR gene is retained, possibly because the MerR protein of the co-resident narrow-spectrum resistance operon will not respond to organomercurials.
In a recent development, the efficiency of Hg(0) evolution from hydrophobic organomercury was improved 10-70-fold by targeting recombinant MerB to the endoplasmic reticulum or the cell wall in Arabidopsis compared to activities of plants carrying cytoplasmic MerB.
To better understand MerB's unique enzymatic activity, nuclear magnetic resonance (NMR) spectroscopy was used to determine the structure of the free enzyme. MerB is characterized by a novel protein fold consisting of three noninteracting antiparallel beta-sheets surrounded by six alpha-helices. By comparing the NMR data of free MerB and the MerB/Hg/DTT complex, a set of residues were identified that likely define a Hg/DTT binding site. These residues cluster around two cysteines (C(96) and C(159)) that are crucial to MerB's catalytic activity. A detailed analysis of the structure revealed the presence of an extensive hydrophobic groove adjacent to this Hg/DTT binding site. This extensive hydrophobic groove has the potential to interact with the hydrocarbon moiety of a wide variety of substrates and may explain the broad substrate specificity of MerB.
An enzyme buffering test was used to determine that the MerB/Hg/DTT complex acts as a substrate for the mercuric reductase MerA. The observed MerA activity is higher than the expected activity assuming free diffusion of the mercuric ion from MerB to MerA. This suggests that the mercuric ion can be transferred between the two enzymes by a direct transfer mechanism.
MerB could be utilized for bioremediation applications, but certain organolead and organotin compounds may present an obstacle by inhibiting the enzyme.
References
1. T. Barkay et al (2003). "Bacterial mercury resistance from atoms to ecosystems." FEMS Microbiology Reviews 27: 355-384.
2. V. B. Mathema et al (2011). "Bacterial mer operon-mediated detoxification of mercurial compounds: a short review". Archives of Mictobiology 193: 837-844.
3. J. Lafrance-Vanasse et al (2009). "Enzyme Catalysis and Regulation: Crystal Structures of the Organomercurial Lyase MerB in Its Free and Mercury-bound Forms: Insights into the Mechanism of Methylmercury Degredation."The Journal of Biological Chemistry 284: 938-944.
4. J. M. Parks et al (2009). "Mechanism of Hg-C Protonolysis in the Organomercurial Lyase MerB."Journal of American Chemical Society 131: 13278-13285.
5. Paola Di Lello, Julien Lafrance‐Vanasse, James G Omichinski. The Organomercurial Lyase MerB. Part 12. Mercury. https://doi.org/10.1002/0470028637.met278
6. Wahba, H. M., Stevenson, M. J., Mansour, A., Sygusch, J., Wilcox, D. E., & Omichinski, J. G. (2017). Structural and Biochemical Characterization of Organotin and Organolead Compounds Binding to the Organomercurial Lyase MerB Provide New Insights into Its Mechanism of Carbon-Metal Bond Cleavage. Journal of the American Chemical Society, 139(2), 910–921. https://doi.org/10.1021/jacs.6b11327
7. Moore, M. J., Distefano, M. D., Zydowsky, L. D., Cummings, R. T., & Walsh, C. T. (1990). Organomercurial lyase and mercuric ion reductase: nature’s mercury detoxification catalysts. Accounts of Chemical Research, 23(9), 301–308. https://doi.org/10.1021/ar00177a006
8. Di Lello, P., Benison, G. C., Valafar, H., Pitts, K. E., Summers, A. O., Legault, P., & Omichinski, J. G. (2004). NMR structural studies reveal a novel protein fold for MerB, the organomercurial lyase involved in the bacterial mercury resistance system. Biochemistry, 43(26), 8322–8332. https://doi.org/10.1021/bi049669z
9. Benison, G. C., Di Lello, P., Shokes, J. E., Cosper, N. J., Scott, R. A., Legault, P., & Omichinski, J. G. (2004). A stable mercury-containing complex of the organomercurial lyase MerB: catalysis, product release, and direct transfer to MerA. Biochemistry, 43(26), 8333–8345. https://doi.org/10.1021/bi049662h
Structure:
https://www.rcsb.org/structure/3F0O
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 415
- 1000COMPATIBLE WITH RFC[1000]