Difference between revisions of "Part:BBa K3398001"
| Line 5: | Line 5: | ||
This part is a T7 promoter derived from BBa_I712074 with some point mutation according to previous research [1], and as a higher expression efficiency relative to BBa_I712074 by the fluorescence intensity when expressing EGFP. To express BioBricks under the control of this part, a T7 polymerase gene is needed in the chassis. | This part is a T7 promoter derived from BBa_I712074 with some point mutation according to previous research [1], and as a higher expression efficiency relative to BBa_I712074 by the fluorescence intensity when expressing EGFP. To express BioBricks under the control of this part, a T7 polymerase gene is needed in the chassis. | ||
| − | |||
===Usage and Biology=== | ===Usage and Biology=== | ||
T7 promoter is very specific promoter which is transcribed only by specific T7 RNA polymerase. Usually this promoter is used in expression systems where T7 promoter is cotransfected with T7 RNA polymerase. That ensures strong transcription of desired genes. | T7 promoter is very specific promoter which is transcribed only by specific T7 RNA polymerase. Usually this promoter is used in expression systems where T7 promoter is cotransfected with T7 RNA polymerase. That ensures strong transcription of desired genes. | ||
| + | |||
| + | |||
| + | ===Characterization=== | ||
| + | ====Result==== | ||
| + | To test the function of mutant promoters, we chose eGFP as our reporter, and added a lac operator behind the promotor to control transcription starts simultaneously. When the E.coli BL21(DE3) is cultured at the stage of logarithmic phase, we added 0.5 mM IPTG to induce the expression of GFP in strains BL21(DE3) for 4 hours. By assessing the absolute fluorescence and OD600, we can conclude the relative strength of all promoters. We screened out four mutants with higher intensity (<partinfo>K2965011</partinfo>, <partinfo>K2965012</partinfo>, <partinfo>K2965013</partinfo>, <partinfo>K2965014</partinfo>). | ||
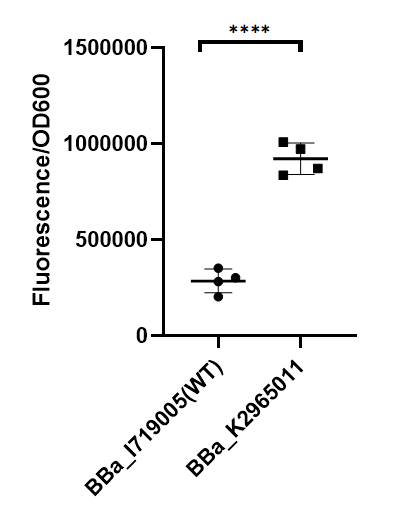
| + | [[File:BBa K2965011 measurement.png|center|350px|thumb|'''Figure 1. Strength of wildtype T7 promoter and mutant promoter.'''Relative fluorescent intensity is standardized with fluorescence (excitation wavelength: 485 nm; detection wavelength: 528 nm) per OD600. The intensity of the mutant was significantly higher than that of the wild type.]] | ||
| + | As we can see from the figure, our mutant promoter showed largely increased strength compared with wild type T7 promoter. Therefore, our mutant promoter offer users more opportunity to control the expression of protein using T7 promoter and permit higher levels of target protein expression to be obtained. | ||
| + | |||
| + | ====Protocol==== | ||
| + | 1. Use BamHI and XhoI to insert eGFP fragment into pET28 plasmid. | ||
| + | |||
| + | 2. Linearize pET28-eGFP plasmid by PCR. | ||
| + | (WT: F-CCCGCTGCAGTAATACGACTCACTATAGGGAGAGGAATTGTGAGCGGATAACAA / R- GCGGTGGACTGCAGCAACTCAGCTTCCTTTCGGGCT ; | ||
| + | Mu: F-CCCGCTGCAGTAATACGACTCACWNNNGSRRNNGGAATTGTGAGCGGATAACAA / R- GCGGTGGACTGCAGCAACTCAGCTTCCTTTCGGGCT ) | ||
| + | |||
| + | 3. Cyclization the plasmid by PstI and ligase. | ||
| + | |||
| + | 4. Transform the plasmids into E. coli BL21(DE3), add 20μl 100mM IPTG to the plate before coating the plate. | ||
| + | |||
| + | 5. Incubate at 37℃ and observe colony color to screen for strong mutants preliminarily. | ||
| + | |||
| + | 6. Pick greener single colonies, add the colonies into 3ml Lb, incubate at 37℃ in a shaker for 6-8h to dilute the IPTG introduced when picking single colonies. | ||
| + | |||
| + | 7. Add 10μl germ solution from last step into 3ml Lb, incubate at 37℃ in a shaker until OD600 reaches 0.6. | ||
| + | |||
| + | 8. Add IPTG to final concentrations 0.5mM, and induce for 4h at 18℃. | ||
| + | |||
| + | 9. Measure the fluorescence (excitation wavelength: 485 nm; detection wavelength: 528 nm) and OD600. | ||
| + | |||
| + | ===Reference=== | ||
| + | [1] Ikeda R A, Ligman C M, Warshamana S, et al. T7 promoter contacts essential for promoter activity in vivo[J]. Nucleic Acids Research, 1992, 20(10): 2517-2524. | ||
| + | [2] Paul S, Stang A, Lennartz K, Tenbusch M, Uberla K. Selection of a T7 promoter mutant with enhanced in vitro activity by a novel multi-copy bead display approach for in vitro evolution[J]. Nucleic Acids Research, 2013, 41(1):e29. | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
<!-- --> | <!-- --> | ||
<span class='h3bb'>Sequence and Features</span> | <span class='h3bb'>Sequence and Features</span> | ||
| − | <partinfo> | + | <partinfo>BBa_K2965011 SequenceAndFeatures</partinfo> |
<!-- Uncomment this to enable Functional Parameter display | <!-- Uncomment this to enable Functional Parameter display | ||
===Functional Parameters=== | ===Functional Parameters=== | ||
| − | <partinfo> | + | <partinfo>BBa_K2965011 parameters</partinfo> |
<!-- --> | <!-- --> | ||
Revision as of 07:56, 8 October 2020
Enhanced T7 promoter 2
This part is a T7 promoter derived from BBa_I712074 with some point mutation according to previous research [1], and as a higher expression efficiency relative to BBa_I712074 by the fluorescence intensity when expressing EGFP. To express BioBricks under the control of this part, a T7 polymerase gene is needed in the chassis.
Usage and Biology
T7 promoter is very specific promoter which is transcribed only by specific T7 RNA polymerase. Usually this promoter is used in expression systems where T7 promoter is cotransfected with T7 RNA polymerase. That ensures strong transcription of desired genes.
Characterization
Result
To test the function of mutant promoters, we chose eGFP as our reporter, and added a lac operator behind the promotor to control transcription starts simultaneously. When the E.coli BL21(DE3) is cultured at the stage of logarithmic phase, we added 0.5 mM IPTG to induce the expression of GFP in strains BL21(DE3) for 4 hours. By assessing the absolute fluorescence and OD600, we can conclude the relative strength of all promoters. We screened out four mutants with higher intensity (BBa_K2965011, BBa_K2965012, BBa_K2965013, BBa_K2965014).
As we can see from the figure, our mutant promoter showed largely increased strength compared with wild type T7 promoter. Therefore, our mutant promoter offer users more opportunity to control the expression of protein using T7 promoter and permit higher levels of target protein expression to be obtained.
Protocol
1. Use BamHI and XhoI to insert eGFP fragment into pET28 plasmid.
2. Linearize pET28-eGFP plasmid by PCR. (WT: F-CCCGCTGCAGTAATACGACTCACTATAGGGAGAGGAATTGTGAGCGGATAACAA / R- GCGGTGGACTGCAGCAACTCAGCTTCCTTTCGGGCT ; Mu: F-CCCGCTGCAGTAATACGACTCACWNNNGSRRNNGGAATTGTGAGCGGATAACAA / R- GCGGTGGACTGCAGCAACTCAGCTTCCTTTCGGGCT )
3. Cyclization the plasmid by PstI and ligase.
4. Transform the plasmids into E. coli BL21(DE3), add 20μl 100mM IPTG to the plate before coating the plate.
5. Incubate at 37℃ and observe colony color to screen for strong mutants preliminarily.
6. Pick greener single colonies, add the colonies into 3ml Lb, incubate at 37℃ in a shaker for 6-8h to dilute the IPTG introduced when picking single colonies.
7. Add 10μl germ solution from last step into 3ml Lb, incubate at 37℃ in a shaker until OD600 reaches 0.6.
8. Add IPTG to final concentrations 0.5mM, and induce for 4h at 18℃.
9. Measure the fluorescence (excitation wavelength: 485 nm; detection wavelength: 528 nm) and OD600.
Reference
[1] Ikeda R A, Ligman C M, Warshamana S, et al. T7 promoter contacts essential for promoter activity in vivo[J]. Nucleic Acids Research, 1992, 20(10): 2517-2524. [2] Paul S, Stang A, Lennartz K, Tenbusch M, Uberla K. Selection of a T7 promoter mutant with enhanced in vitro activity by a novel multi-copy bead display approach for in vitro evolution[J]. Nucleic Acids Research, 2013, 41(1):e29.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]