Difference between revisions of "Part:BBa K3187028"
JonathanFu (Talk | contribs) |
JonathanFu (Talk | contribs) |
||
| Line 1: | Line 1: | ||
| + | |||
| + | __NOTOC__ | ||
| + | <partinfo>BBa_K3187028 short</partinfo> | ||
| + | |||
ext install esbenp.prettier-vscode | ext install esbenp.prettier-vscode | ||
__NOTOC__ | __NOTOC__ | ||
| Line 1,990: | Line 1,994: | ||
</html> | </html> | ||
| + | |||
| + | <!-- Add more about the biology of this part here | ||
| + | ===Usage and Biology=== | ||
| + | |||
| + | <!-- --> | ||
| + | <span class='h3bb'>Sequence and Features</span> | ||
| + | <partinfo>BBa_K3187028 SequenceAndFeatures</partinfo> | ||
| + | |||
| + | |||
| + | <!-- Uncomment this to enable Functional Parameter display | ||
| + | ===Functional Parameters=== | ||
| + | <partinfo>BBa_K3187028 parameters</partinfo> | ||
| + | <!-- --> | ||
Revision as of 03:58, 21 October 2019
Sortase A7M (Ca2+-independent variant)
ext install esbenp.prettier-vscode
Sortase A7M (Ca2+-independent variant)
Profile
| Name | Sortase A7M |
| Base pairs | 470 |
| Molecular weight | 17.85 kDa |
| Origin | Staphylococcus aureus, synthetic |
| Parts | Basic part |
| Properties | Ca2+-independent, transpeptidase, linking sorting motif LPXTG to poly-glycine Tag |
Usage and Biology
Transpeptidase: Sortase
Sortases belong to the class of transpeptidases and are mostly found in gram-positive bacteria.
The high rate of resistance to several antibiotics targeting gram-positive bacteria is also based on the
property of this enzyme class. Sortases can non-specifically attach virulence and
adhesion‐associated proteins to the peptidoglycans of the cell-surface.
[21]
[22]
In general, sortases are divided into six groups (A-F) that have slightly different properties and
perform three tasks in cells. Group A and B attach proteins to the cell-surface while Group C and D help
building pilin-like structures. Group E and F are not properly investigated yet which is why their exact
function is not known.
[23]
For our project we are especially interested in the sortases of the group A since they
covalently attach various proteins or peptides on the cell membrane as long as their targeting
motif is at the C-terminus of the corresponding protein. In comparison to other transpeptidases
Sortase A has the advantage that it is rather stable regarding variations in pH
[24]
and temperature.
[22]
Sortase A catalyzes the formation and cleavage of a peptide bond between the C-terminal
LPXTG amino acid motif and an N-terminal poly-glycine motif. The enzyme originates from
Staphylococcus aureus and is able to connect any two proteins as long as they possess those
matching target sequences.
[25]
In the pentapeptide motif LPXTG, X can be any amino acid except cysteine.
[21]
Sortase A is rather promiscuous with regard to the amino acid sequence directly upstream of
this motif, a fact that makes it optimal for labeling applications. Even better, amino acids C-terminal
of the poly-glycine motif are not constrained to a certain sequence.
[26]
The reaction works with any primary amine.
[30]
Reaction
To better understand how the enzymatic reaction works it is necessary to look at the crystal structure of Sortase A. The enyzme consists of an eight-stranded β‐barrel fold structure. The active site is hydrophobic and contains the catalytic cysteine residue Cys184 as well as a key histidine residue H120 that can form a thiolate-imidazolium with the neighboring cysteine. An additional structural property that also other sortases show is the calcium binding site formed by the β3/β4 loop. The binding of a calcium ion slows the motion of the active site by coordinating to a residue in the β6/β7 loop. This helps binding the substrate and increasing the enzymatic activity nearly eightfold. [26] When a substrate gets into the active site, the cysteine attacks the amide bond between the threonine and the glycine in the LPXTG motif. After this the protonated imidazolium serves as an acid for the departing glycine with unbound NH2 of the former amide bond while the rest of the motif is bound to the cysteine residue. Another glycine nucleophile is then necessary in its deprotonated form to attack the thioester and re-establish an amide bond at the LPET-motif. This reaction is dead-ended if the used nucleophile is water.[26] Due to the fact that the mechanism is based on protonated forms of the catalytic residues the reaction is quite pH-dependent. Although the Sortase A in general is relatively stable between pH 3 and 11 the reaction works best around pH 8.[28]
Sortase variants
Due to the fact that the wildtype Sortase A shows rather slow kinetics, a pentamutant has been developed (Sortase A5M). [29] This version of the enzyme carries mutations in P94R/D160N/D165A/K190E/K196T which lead to a 140- fold increase in activity. Thereby, reaction rates are improved even at low temperature, however, Sortase A5M is still Ca2+-dependent. This dependence interferes with potential in vivo usage, as the concentrations of calcium in living cells can vary considerably. Hence a sortase mutant that acts across high differences in calcium concentrations or even works completely Ca2+-independently would be required for in vivo applications of sortase. To attain a high yield enzyme which is also calcium-independent Ca2+-independent mutations were combined with the Sortase A5M resulting in Sortase A7 variants such as the Sortase A7M. The newly achieved calcium-independence of these variants enable sortase applications not only in vitro but in vivo as well. [30]
Sortase A7M
For our project we chose to work with this optimized Sortase A7M. Its size is about 17.85 kDa and it has been shown to be stable for several weeks in the fridge at 4 °C. It also possesses the same properties of pH stability like other sortases [31] but comes with the advantage of being calcium independent. [32] "Sortagging" applications have included the cyclization of proteins and peptides [31] , modification and labeling of antibodies and the synthesis of protein conjugates with drugs, peptides, peptide nucleic acids and sugars. [27] Moreover it poses a lot of advantages for the binding of two proteins in vivo since it has relatively small tags which avoids putting too much metabolic burden on the cells when expressing the proteins of interest. This also avoids disturbing the folding of the proteins of interest and the later biological functions since the Sortase A7M is able to work under physiological conditions. [30] Other methods like the intein- based labeling of surfaces require large fusion-proteins with the intein domain which puts stress on the living cells and might cause folding and solubility issues. Another application for sortase-mediated systems is the anchoring of proteins on the cell wall of gram-positive bacteria which can be used for display of heterologous proteins. [26] It is also possible to attach non-biological molecules to the respective tag. The accessibility and flexibility determine the ability of a sortase enzyme to recognize the sorting motif and catalyzing the transacylation.
Methods
Cloning
h2>CloningThe methods used for cloning of the different mutants of the sortase were restriction and ligation via NdeI and SalI and Gibson assembly. The Sortase A5M was cloned into pET24(+) vector via restriction and ligation NdeI and SalI as restriction enzymes. The vector posesses a kanamycin resistance and is controlled through a T7 promoter, which can be induced with IPTG. Sortase A7M is controlled by the same T7 promoter. Sortas A, introduced by iGEM Stockholm 2016, was cloned via Gibson assembly into PSB1C3. This has a chloramphenicol resistance and is also controlled under a T7 promoter. Cloning of all products was checked via sequencing.
Expression and purification
After successfully transforming our sortase genes in BL21 cells, we inoculated 100 mL overnight cultures, with the respective antibiotic. The next day 1 L cultures were inoculated with the overnight culture to reach OD600 = 0.1. Subsequently the cultures were incubated under constant shaking at 37 °C until they reached OD600 = 0.6. At OD600 = 0.6 the cultures were induced with 0.5 mL of 1 M Isopropyl-β-D-thiogalactopyranosid (IPTG). The gene expression was performed at 30 °C under constant shaking overnight. After expression of Sortase A7M, Sortase A5M, and Sortase A from Stockholm (BBa_K2144008) in BL21 cultures the cells were crushed via EmulsiFlex (Avestin) and proteins were purified through affinity chromatography via Fast Protein Liquid Chromatography (FPLC) with the ÄKTA pure (GE Healthcare, Illinois, USA). His-Tag was used for purification of Sortase A7M and Sortase A (Stockholm) and Strep-Tag II was used for purification of Sortase A5M.
SDS-Page
To verify the successful production of of Sortase A7M, Sortase A5M, and Sortase A SDS-PAGEs were performed. The resulting bands were compared to the molecular weight of the different sortase variants. Also, SDS-PAGEs were completed to verify enzymatic activity in assays prior to measuring sortase properties via Fluorescence Resonance Energy Transfer (FRET).
Flourescence Resonance Energy Transfer (FRET)
To determine the kinetics of our transpeptidase variants, FRET assays were performed in 384 well-plates (dark) using a Tecan plate reader. A FRET relies on the phenomenon that an excited fluorophore (donor) transfers energy to another fluorophore (acceptor), thereby exciting it. This process only works if both fluorescent molecules are in close proximity and depends on the FRET-Pair. By transferring the energy from donor to acceptor, the donor's emission is reduced and the intensity of the acceptors emission is increased [1] . The efficiency depends on the distance between the fluorophore, the orientation and the spectral characteristics [2] . You can see the principle of FRET in Fig. 1.
Mass Spectrometry
To estimate the product yield of catalyzed reactions by Sortase A7M we performed mass spectrometry. The tested molecules can be distinguished between products and educts due to desorption and ionization. Therefore, we used the electrospray ionization (ESI) technique for the mass spectrometry. This technique has a low resolution but is a very soft ionization method, which makes it an optimal method for biological molecules. [1]
Results
Characterization of Sortase A7M (and comparison to Sortase A5M)
How do we measure if our purified sortases are active?
After purification of the sortases, we first performed SDS-PAGEs to verify that they are pure and monomeric. You can see in Fig. 3 that the purifications were successful. Next, we tested if the purified sortases connect two proteins that carry the important Sortase-recognition tags, N-terminal polyG and C-terminal LPETGG. Therefore, we added the sortases to a mix of GGGG-mCherry and mCherry-LPETGG. The reactions were performed in different buffers, at different enzyme-to-substrate ratios and for different time spans. We performed an SDS-PAGE, and prior to Coomassie staining, we recorded fluorescent images of the gel. Thereby, we could identify mCherry bands in the gel.

Figure x :SDS-PAGE of Sortase A7M and Sortase A5M where the bands show up at approximately 15 kDa. Our estimated size for Sortase A7M was 17.85 kDa, and for Sortase A5M 18.07 kDa. This confirms the result shown on the gel, since the band of Sortase A5M is a little higher than the one of Sortase A7M. caption

Figure x : caption

Figure x : Fluorescence gel of the sortase-reaction of GGGG-mCherry and mCherry-LPETGG
mediated by Sortase A7M incubated for 2 h and
4 h each. Reaction solutions were mixed with different ratios from enzyme to
substrate concentration(1:3;1:10) and each incubated in two different buffers(Tris-HCl
and Ammoniumdicarbonat).
Product bands at a height of about 57 kDa can be seen in lane 4, 5, 6, 8, 9 (from
left to right). The bands below the product at about 38 kDa could be semi-denatured
mCherry dimers.
b) Fluorescence gel on top of the coomassie-stained gel of the sortase-reaction
of GGGG-mCherry and mCherry-LPETGG mediated by Sortase A7M incubated for 2 h
and
4 h each. Reaction solutions were mixed with different ratios from enzyme to
substrate concentration(1:3;1:10) and each incubated in two different buffers (Tris-HCl
and Ammoniumdicarbonat).
Product bands at a height of about 57 kDa can be seen in lane 4, 5, 6, 8, 9 (from
left to right). The bands below the product at about 38 kDa could be semi-denatured
mCherry dimers. Additionally, Sortase A7M can be seen at 17 kDaA7M.The
unprocessed mCherry monomers can be seen at 28 kDa.
caption
As shown in Fig. 4, under certain conditions, a product band appeared at the expected size of 57.3 kDa (28.5+28.8 kDa). From this first activity test, we draw three conclusions:
- Our purified Sortase A7M is active
- The enzyme-substrate ratio affects the product yield
- The duration of the reaction affects the product yield
Additionally, TRIS buffer seems to alter the coomassie staining efficiency of Sortase A7M.
This endpoint measurement gave us a first impression that our Sortase A7M works nicely. Of
course, we wanted to further characterize the parameters of the reaction. When we understand
the Sortase better, modification of our VLPs will become more straightforward.
how do we measure sortase reaction kinetics
In the above described assays, we noticed the impact of enzyme-substrate ratio and reaction duration on the overall product yield. We thought about how to further measure the kinetics of the sortase reaction. In the literature, sortase reaction kinetics are often measured by FRET-assays. Therefore, we designed a suitable FRET-assay[1]. In the end, we came up with a new FRET pair not described in the literature to date: 5-TAMRA-LPETG and GGGG-sfGFP.
Development of a new FRET pair
For characterization of the reaction kinetics of Sortase A7M, Sortase A5M and Sortase A, we decided to develop a suitable FRET pair. In order to find an optimal FRET pair, we first recorded an emission and absorption spectrum of 5-Carboxytetramethylrhodamin-LPETG (TAMRA) and GGGG-mCherry to verify the suitability for the FRET effect, checking for a possible overlap between the donor's emission and the acceptor's extinction.

Figure x :Design of a FRET-pair of 5-TAMRA-LPETG (TAMRA) and GGGG-mCherry (mCherry). In this configuration TAMRA acts as donor and mCherry as acceptor. When the two fluorophores are not linked via the substrates of the sortase only TAMRA is being excited. After sortase mediated ligation of the two substrates mCherry is the fluorophore being excited via the FRET and the emission of mCherry intensifies. Meanwhile, the emission of TAMRA decreases. caption
TAMRA is a chemical fluorophore that has an absorbance maximum at 542 nm and an emission
maximum at
570 nm[1]. The
terminal carboxy
group of the dye was linked via a lysine linker to the LPETG sequence (see Fig. 5).
mCherry has
an N-terminal poly-glycine sequence and can therefore be linked to the LPETG motif of TAMRA via
the
Sortase A. For a sufficient FRET-effect, it is also necessary that the distance between
donor and
acceptor is lower than the Förster radius. The Förster radius describes the distance between two
fluorophores at which 50 % of the energy is transferred.
First, we wanted to identify which concentrations are needed for our experiment, then set up the
reaction
and measured fluorescence intensities. Over time, a decline in the emission of TAMRA can be
observed as
Sortase A7M/A5M is converting more educts to products.

Figure x : The graph shows the extinction and emission spectra of TAMRA and mCherry. Due to the large overlap of TAMRA emission and mCherry extinction it is possible to perform a FRET with this pair of fluorophores. The graph show the relative fluorescence unit (RFU[%]) in relation to the extincted/emitted wavelength [nm]. The peaks are normalized to 100 %. caption
The emission and extinction spectra of TAMRA and mCherry exhibit an overlap of emission of TAMRA and extinction of mCherry. Based on this output, a FRET-assay for the kinetics of Sortase A7M was performed to confirm whether the FRET-pair is working. As TAMRA is excited with light of a lower wavelength than mCherry, the former serves as FRET donor and the latter as acceptor. We chose the excitation wavelength at 485 nm to prevent unnecessary “leak” excitation of mCherry. Nevertheless, an extinction of mCherry could not be excluded and may have negative effects on the visibility of the FRET.

Figure x : caption

Figure x : Spectrum of TAMRA and mCherry, with Sortase A7M, over the course of 20 min in 5 min intervals. Depicted are the emission wavelengths against the RFU. The sortase-mediated ligation results in a decline of both emission peaks. caption
The analysis of the data shown in Fig. 7 confirmed the aforementioned suspicion that mCherry is also excited at 485 nm, which makes differentiation of the fluorescence more difficult. Furthermore, Fig. 8 shows that the difference in the decline of TAMRA is not significant. Accordingly, a decline in the emission maximum of TAMRA over time is also visible in the negative control. One reason might be bleaching of TAMRA through the excitation by the laser. Nevertheless, conversion by the Sortase A7M can be observed by comparing the results with the negative control.

Figure x :Sortase reaction in TAMRA mCherry FRET after subtracting the negative control. Depicted is the difference in RFU over time [min]. WIthin the first 20 min of the substrate conversion is the quickest. At 30 min a plateau is reached. After 60 min starts catalyzing the reverse reaction. The mean ΔRFU value was normalized to zero for better visualization. caption
To confirm the functionality of the Sortase A7M, another more sufficient FRET-pair was developed. The measured absorbance and emission spectra indicated that TAMRA and superfolder green fluorescence protein (sfGFP) are a possible FRET-pair. The sfGFP has an N-terminal polyglycine sequence and can therefore be linked to TAMRA with the sorting motif, in the same way as mCherry was connected. However, the small overlap between the extinction spectra of sfGFP and TAMRA could solve the previous “simultaneous excitation” problem we observed for the mCherry-TAMRA FRET-pair. Because of the lower excitation maximum of sfGFP compared to TAMRA, sfGFP was chosen as donor and TAMRA as acceptor. sfGFP was excited at 465 nm to minimize the unnecessary leak excitation of sfGFP.

Figure x : caption

Figure x :Design of a FRET-pair of 5-TAMRA-LPETG (TAMRA) and GGGG-sfGFP (sfGFP). In this configuration sfGFP acts as donor and TAMRA as acceptor. When the two fluorophores are not linked only sfGFP is being excited. After sortase-mediated ligation of the two substrates, TAMRA is the fluorophore being excited via FRET and the emission of TAMRA intensifies. Meanwhile, the emission of sfGFP decreases. caption
The transfer of energy from sfGFP to TAMRA can be seen by the decrease in emission of sfGFP and increase in emission from TAMRA. Compared to TAMRA as an acceptor, the sfGFP bleaches significantly less and is consequently more suitable as a donor for FRET. Furthermore, the afore mentioned problem of simultaneous donor and acceptor excitation seems to be solved. It seems that we have found a FRET-pair with superior properties.

Figure x : caption

Figure x :Spectrum of TAMRA and sfGFP, with Sortase A7M, over the course of 25 min in 5 min intervals. Depicted are the emission wavelengths against the RFU. The sortase-mediated ligation results in a decline of both emission peaks. caption
Due to the collected data of both FRET-pairs we decided to use the TAMRA-LPETG and GGGG-sfGFP FRET-pair for further characterization of our Sortase A variants. Two reasons justify this decision:
- TAMRA bleaches stronger than sfGFP when excited with a laser.
- The spectral overlap between TAMRA and mCherry disturbs “clean” energy transfer, thus the FRET-effect would be less visible and could not be used for analysis of the sortase-mediated reaction.
For recording of sortase reaction parameters we recommend using the FRET-pair sfGFP-TAMRA. As this pair of fluorophores proved to have near perfectly aligned spectra and since the bleaching effect is visibly lower on sfGFP than on TAMRA, we chose to use this FRET-pair in most of our following assay. Nevertheless, we do not rule out the use of TAMRA-mCherry as a FRET-pair since we used it in several FRET-assays as well.
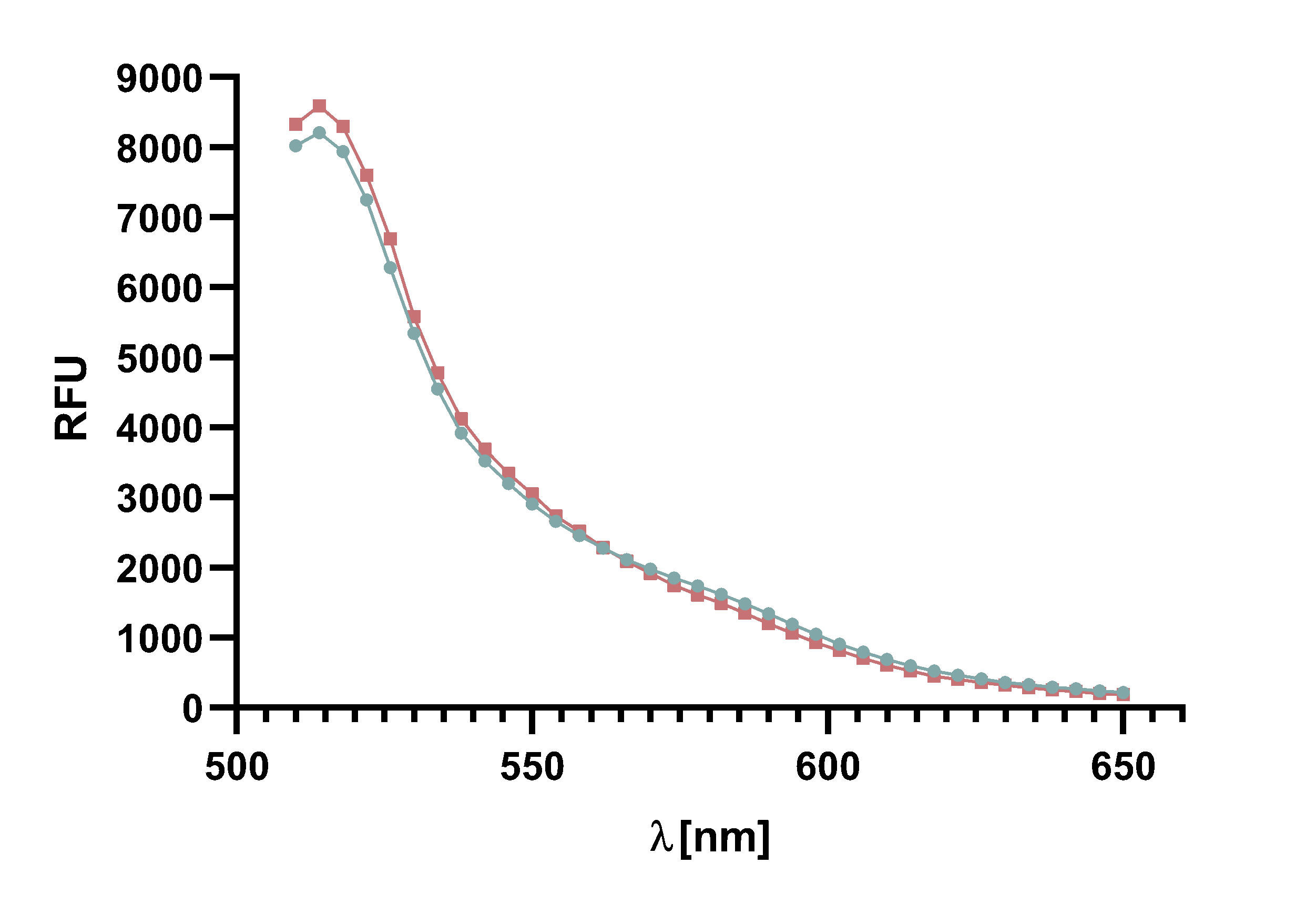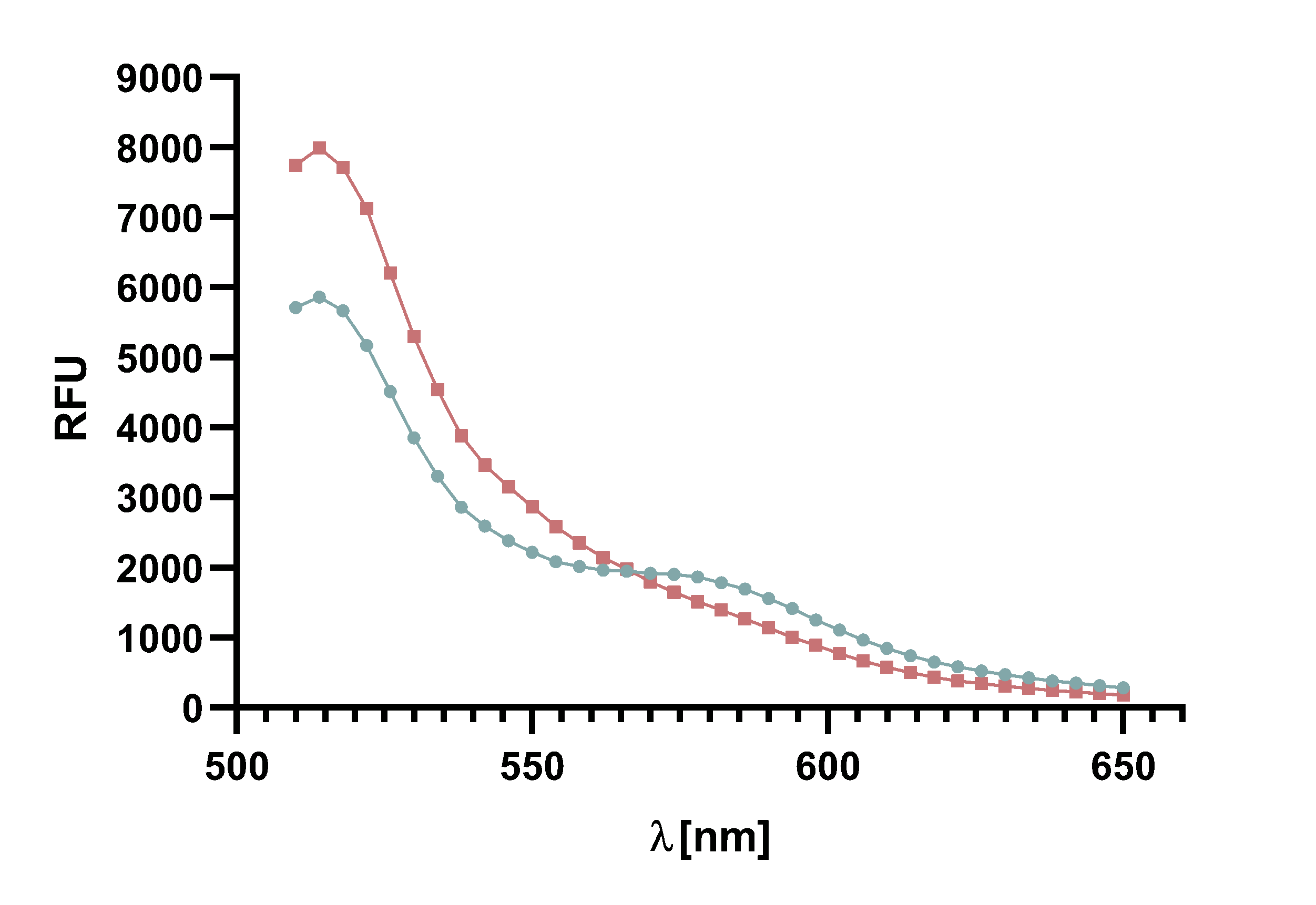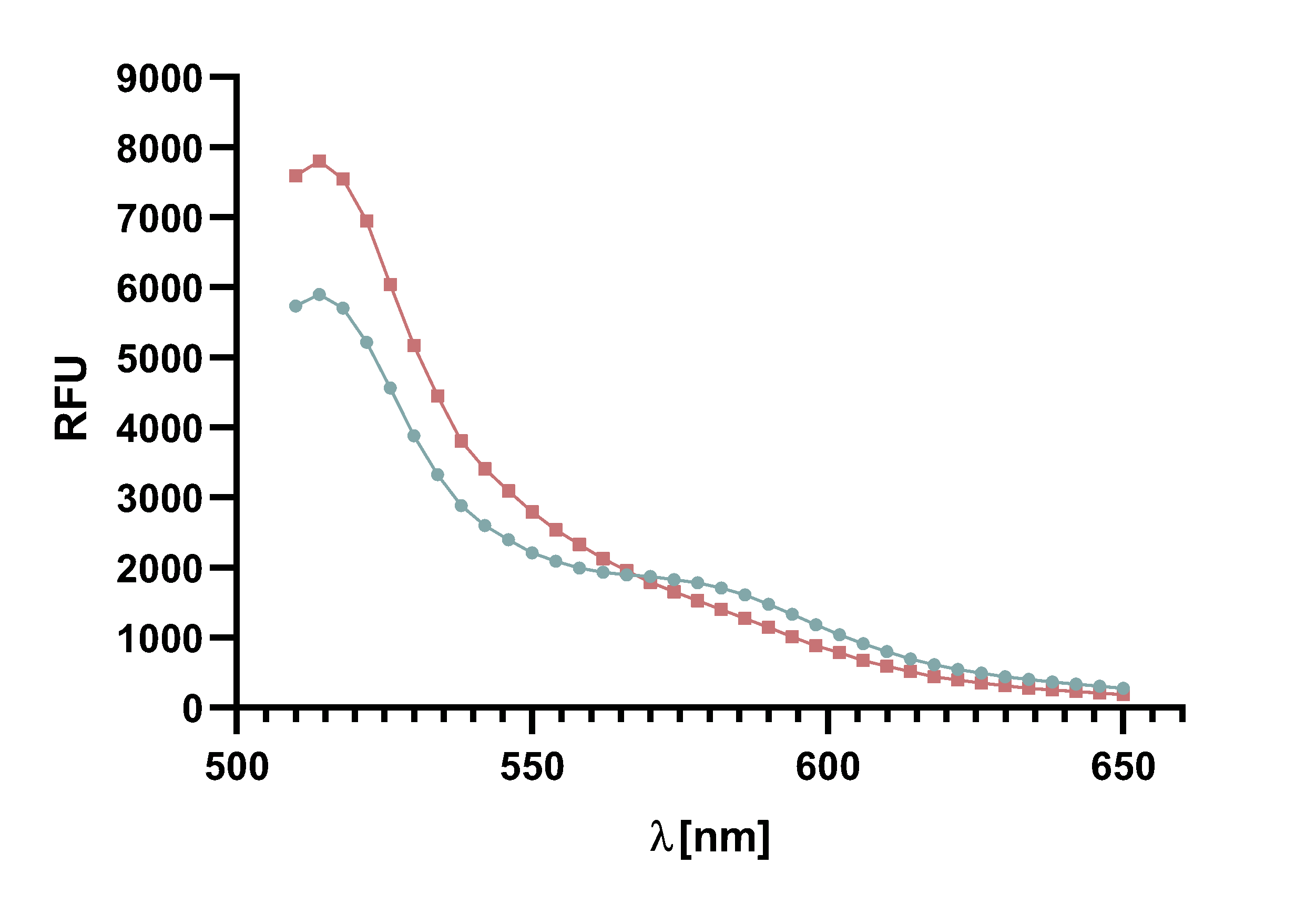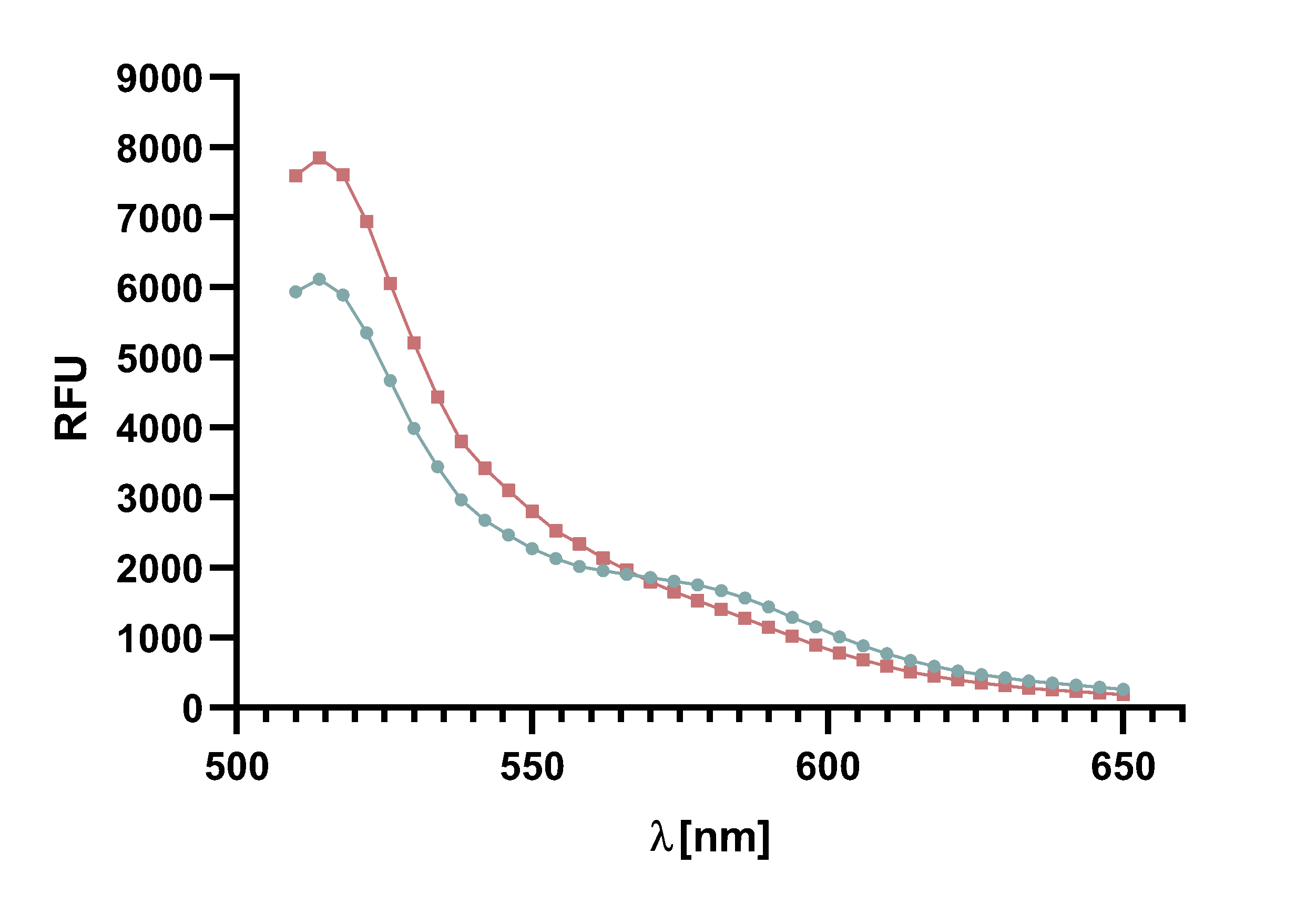
Figure x :Animation of Sortase A7M enzyme kinetics over the course of 3 h. The reaction speed increases radically in the beginning moving from RFU 8000 to RFU 6000 at λ = 550 nm where a plateau is reached (blue). The negative control (orange) is also reduced in its RFU due to bleaching. Nevertheless, a peak at λ = 580 nm arises already after short reaction time. This peak indicates the successful Fluorescence Resonance Energy Transfer. caption
Why are enzyme-substrate ratio and duration important parameters of the sortase reaction?
In one of our first FRET experiments, we addressed the simple theory: More sortase in the reaction mix improves the initial product formation. For this, we used the TAMRA-LPETG : GGGG-mCherry FRET pair. We measured the FRET change over time in a multiwell platereader (Fig. 14).

Figure x :Reaction kinetics of Sortase A7M in different concentrations at same level of substrate concentration. caption
However, in this assay we observed a striking feature of the sortase reaction. In the reaction with more Sortase A7M present, the FRET change started to decrease after a certain maximum was reached! We suspected some kind of dead-end product formation, as the sortase does also catalyze the reverse reaction of product to educts. Therefore, the overall reaction duration is a very important parameter. We gathered more details about the role of the reverse reaction during our comparison of Sortase A7M and Sortase A5M. Just keep reading if you want to know more!
Who wins - Sortase A7M or Sortase A5M
In our introduction we described that Sortase A7M and Sortase A5M are both fascinating enzymes, although each of them has a unique „selling point“. Sortase A5M is faster, whereas Sortase A7M is Ca2+-independent. We confirmed both of these points in extensive FRET-assays. According to the literature, Sortase A5M works best with a Ca2+-concentration of 2 mM. In contrast, Sortase A7M is a calcium-independent mutant of the enzyme. Moreover, Ca2+ even seems to inhibit this enzyme variant slightly [1].
Firstly, we confirmed that in contrast to Sortase A5M, Sortase A7M is Ca2+-independent. The results are shown in Fig. 15 Sortase A7M also works in presence of Ca2+, but these FRET experiments made us suspect that Ca2+ may even inhibit Sortase A7M.

Figure x :Sortase A7M FRET-assay of connecting TAMRA-LPETG with GGGG-sfGFP with and without Ca2+. The Sortase A7M reaction was measured with 6 mM Ca2+ every minute. Sortase A7M reaction without Ca2+ was measured every three minutes. It is shown that this enzyme variant works with calcium and without calcium as well, although it seems like Sortase A7M is slightly inhibited due to the presence of calcium which explains why the left graph is lower than the right one. caption
Secondly, we confirmed that Sortase A5M is inactive if Ca2+ is absent, which can be seen in Fig. 16 As expected, Sortase A5M shows increasing enzymatic activity with increasing Ca2+ levels. The reaction runs fastest with 2 mM Ca2+, and the maximal FRET change (in terms of ΔRFU) is reached after 37.5 min. Strikingly, the FRET change decreases afterwards. We observed this phenomenon before and assume this to be due to dead-end product formation caused by the reverse reaction.

Figure x : caption

Figure x : Both figures show Sortase A5M FRET assay connecting TAMRA-LPETG and GGGG-sfGFP with different Ca2+-concentrations. The right graph is showing that Sortase A5M does not work without calcium. The negative ΔRFU measurements are probably caused by measuring errors by the Tecan platereader. caption
According to the results of this assay, Sortase A7M is definitely Ca2+-independent, since it shows linking activity without calcium in the vicinity. The enzyme mutant also works in presence of Ca2+ (Fig. 15), but these FRET experiments made us suspect that Ca2+ may even inhibit Sortase A7M, since it shows less activity with calcium around than without calcium.
To better address this question, an ELISA was performed. Therefore, a piece of paper functionalized with GGGβA was connected to a protein domain, which binds antibodies to the LPTEG-tag. The results are shown in Fig. 17.

Figure x :
Absorbance at 450 nm at a temperature of 23.8˚ C
In well 1 additional 10 mM Ca2+ were
added which was not the case in well 2. Well 3 serves as a negative control since
the enzyme is missing in this reaction
caption
As shown in Fig. 17, the highest absorption was measured in well 2. Thus, Sortase A7M works more efficiently when no Ca2+ is around. The absorption is also relatively high for the negative control, which can be explained by poor washing before the substrate for Horeseradish peroxidase (HPR) was added. This assay shows the functionality of Sortase A7M even in context of surfaces since we confirmed that Sortase A7M is able to connect tags attached to paper. This shows that the surface structure is not a relevant factor for the enzyme.

Figure x :Absorbance at 450 nm at a temperature of 23.8˚ C
In well 1 additional 10 mM Ca2+ were
added which was not the case in well 2. Well 3 serves as a negative control since
the enzyme is missing in this reaction.
caption
When we compare the reaction speed of Sortase A5M and Sortase A7M, Sortase A5M is the clear winner. However, this means of course that the reverse reaction is also faster in the case of Sortase A5M. Consequently, Sortase A7M is the best variant for in vivo modification of our VLPs as it is Ca2+-independent. On the other hand, Sortase A5M is a suitable enzyme variant for in vitro modification due to its high efficiency.
What about other substrates?
Primary Amines
The literature describes Sortase A7M as somewhat „promiscuous“ towards other substrates than GGGG(polyG) as long as the substrate possesses a primary amine. To confirm this, we performed additional assays with other substrates in the lab of Prof. Kolmar. The Sortase A7M used for this assay was stored in the fridge at 4 °C for two weeks. The substrates were TAMRA with a KLPETG bound to TAMRA via the lysine side chain and 3-azidopropanamine as the example for a primary amine. The reaction was performed for two hours at 37 °C. It was then analyzed by electron spray ionization mass spectrometry (ESI-MS) (Fig. 18).

Figure x : Mass spectrum before the reaction of TAMRA-LPETG with 3-azidopropanamine showing the educt at 1054 g/mol. caption
Fig. 18 shows the educt-peak in the mass spectrum. TAMRA with the LPETG-tag weighed 1054 g/mol. Shown above in green is the single charged molecule at 1054.27 g/mol and the double charged molecule at 528.75 g/mol.

Figure x : Mass spectrum after the reaction of TAMRA-LPETG with 3-azidopropanamine showing the product at 1079.37g/mol. caption
Yield
For the characterization of Sortase A7M an assay was designed to show the coupling efficiency between the TAMRA-LEPTG and the tetrapeptide GGG-Beta-Alanin (GGGβA) catalyzed by the Sortase. The β-Alanin functions as a spacer. The Sortase reaction was performed for 1h at 30˚C and was stopped by enzyme separation through centrifugal filtration. For analysis mass spectrometry (ESI-MS) was used. The mass spectrometry enables differentiation between products and educts. It allowed us to make an estimate of the product yield. The calculated theoretical molecular masses are 1054 g/mol for TAMRA and 1240 g/mol for TAMRA-LPETGGGβA. Therefore, peaks are expected at mass/n, with n ∈ N. By comparison of the number of corresponding peaks, estimation of the product yield is possible as both molecules possess the same amount of ionizable groups and thus the difference in the ionizability of both molecules is negligible.

Figure x :Mass spectrum of the sortase-mediated ligation of TAMRA-LPETG and GGGβA showing the difference in height of the educt-peak and the product-peak which can be used to estimate the yield of our Sortase A7M. caption
In Fig. 20 the 621.56 peak can be assigned to the TAMRA-LEPTGGGβA and the 528.85 to the TAMRA-LPETG. The count ratios of the two molecules mentioned show an excess of the product.
Is Sortase A7M able to attach cargo to P22 coat protein?
One of the most important questions in our project was: Can Sortase A7M attach proteins to tagged P22 coat protein (CP-LPETGG)? This is one of the absolute requirements of our MVP platform.
We performed the linking reaction with CP-LPETGG and GGGG-mCherry as substrates and applied them to an SDS-PAGE. We saw products at the expected size (28 kDa + 49 kDa = 77 kDa) thus the requirement is fulfilled. However, a lot of additional bands appeared that we did not expect. These bands also appeared when only Sortase A7M and CP were mixed.

Figure x : caption

Figure x : caption
Figure 21:
a) Sortase A7M band is at expected height (17.85 kDa). The two negative controls containing only GGGG-mCherry (28 kDa) and CP-LPETGG (49 kDa) at the expected respective heights. b) Shown are sfGFP-SP and CP-LPETGG each incubated with both Sortase A7M and Sortase A5M. Both gels display multimers when coat and a sortase variant are in a sample together.
To investigate this issue, we had a look at the literature and found a matching description in the publication of Patterson et al.. They performed a similar experiment with P22 capsid proteins and observed the same multimers in their SDS-PAGEs [2] . Comparing both SDS-PAGEs, we came to the following assumption:
Because of the promiscuity of Sortase A7M to accept primary amines as substrates, as we discussed previously, the formation of CP multimers occurs, unspecifically catalyzed by Sortase A7M.
Parallel to these experiments, we successfully modified the exterior of pre-assembled VLPs in vitro (VLP assembly). These modified VLPs were homogenous and overall correctly assembled. Therefore, we conclude that the described multimer problem only occurs when Sortase A7M encounters free CP.
Does methionine affect Sortase linking?
Sortase A7M preferably attaches N-terminal poly-G to C-terminal LPETGG. However, the first amino acid of a protein is methionine (to be specific, formylmethionine in bacteria). For our constructs that possess N-terminal polyG-tags, we have to ask ourselves the question: If the initial methionines are not cleaved off after the proteins have been produced, will this interfere with the Sortase reaction?
To investigate this, we cloned and purified another protein: TVMVsite-GGGG-mCherry. This protein can be treated with TVMV-protease, leading to *GGGG-mCherry. This *GGGG-mCherry was then compared to (M)GGGG-mCherry we used in all previous assays.
Also, we tested if it is possible to also link another substrate with another protease cleavage site infront of the N-terminal GGGG-sequence also aplicable. Fig. XY shows that this is possible. Due to these findings we modified our VLPs with *GGGG-sfGFP.

Figure x : Sortase-mediated ligation of TAMRA-LPETG and GGGG-sfGFP (with TEV cleavage site) one cut with TEV protease and one not. The sample with the unprocessed substrate shows no increase in RFU. In contrast the processed substrate shows a clear increase in delta RFU. After 90 min the reverse reaction begins.
We performed FRET-assays with TAMRA-LPETG and either of the following reaction partners:
- (M)GGGG-mCherry, a protein sample that might still carry an N-terminal methionine
- *GGGG-mCherry that does not carry any additional N-terminal residue
Before the FRET-assay was started, we adjusted the mCherry-concentrations of both fluorescent protein solutions to the same level. To do so, we diluted them until both showed the same fluorescence at 610 nm.

Figure x : caption

Figure x : caption
Figure x: FRET of the sortase reaction connecting TAMRA-LPETG and GGGG-sfGFP mediated by Sortase A7M. The concentration of the Sortase A7M was kept at the same level why the concentration of sfGFP was either 7.8 mM or 1 mM. The graphs show that the reverse reaction happens earlier if if the GGGG-substrate concentration is lower.
Strikingly, only the (M)GGGG-mCherry construct showed a clear decrease in delta RFU after the maximum delta RFU was reached (at about XXX min).

Figure x : Sortase-mediated ligation of TAMRA-LPETG and GGGG-mCherry one cut with TVMV protease and one with a methionin infront of the GGGG-tag. As visible the reverse reaction happens earlier if the methionine is not cleaved of the GGGG-tag. The delta RFU is referring to the negative controls without Sortase A7M.
We assume the following: Although we adjusted the overall mCherry concentration by fluorescence, we cannot determine the absolute amount of MGGGG-mCherry in the (M)GGGG-mCherry sample. However, if this amount was relatively high, the effective substrate concentration that could enter the sortase reaction would be low. That is because MGGGG is a worse sortase substrate than GGGG – if any at all. If we furthermore consider that a low substrate concentration correlates with a faster reverse reaction, we can explain the observed decrease in delta RFU for the (M)GGGG-mCherry sample that contrasts the delta RFU trend of the *GGGG-mCherry sample.
On this basis we can assume that a certain, yet unknown portion of the (M)GGGG-mCherry sample still carries an N-terminal methionine.
These FRET-assays let us assume that methionine disturbs or at least interferes with the sortase reaction mechanism. Indeed, our modeling suggests that methionine affects the interaction of polyG and the flexible loop near the active site of Sortase A7M. Click here if you want to know more about our modeling results!
We propose that potential users of our platform introduce a protease cleavage site in front of the GGGG-protein in order to ensure successful modification of the VLP surface.
This strengthens our hypothesis: If there is any amino acid in front of the poly-glycine sequence, substrate binding to Sortase A7M is negatively influenced.
Modeling
Introduction
In synthetic biology, theoretical models are often used to gain insights, predict and
improve
experiments. In our project we are modifying Virus-like particles (VLPs) by attaching
proteins to the
surface of the P22 capsid
through a linker. The linking is
catalyzed using
the enzyme Sortase A7M, which is a calcium independent mutant of the wild type Sortase A
from Staphylococcus aureus. We performed
modeling to predict the unknown structure of the
Sortase A7M, to improve the linker between proteins and therefore optimizing the
modification
efficiency of our platform.
Two different modeling approaches were used to determine the structure of Sortase A7M.
We compared
machine learning approaches to traditional comparative, Monte-Carlo based modeling
methods. The
results were evaluated using an energy-scoring function and molecular dynamics (MD)
simulations. The
most promising Sortase A7M structures were used to perform a docking simulation to
screen for
optimal linkers.
Structure determination
In silico modeling and simulation of proteins requires a 3D structure, which can be obtained from the RCSB Protein Data Bank. However, if no 3D structures are annotated, as it is the case with sortase A7M, the structure has to be determined by other means. The structure prediction of sortase A7M was done using two different approaches.
RosettaCM
Background
In our second approach we used the RosettaCommons comparative modeling (RosettaCM), which is based on homology modeling. Homology modeling is a protein modeling method, which requires one or more template structures as base the protein to be modeled on. The protein sequences are aligned with the sequence of the target protein. Unaligned sections are modeled using fragment or protein libraries, which leads to creating protein structures based on different sequence homologues of the protein of interest. Ab-initio or de novo modeling on the other hand attempts to find protein structures solely based on physicochemical principles applied to the primary sequence, which can be compared to the refolding of a denaturated protein.
RosettaCM combines ab-initio modeling with homology modeling. The homologus structures for which a resolved 3D structure with sufficiently similar sequence exists are generated using homology modeling. Afterwards the unaligned sequences are modeled de novo. By combining the two methods RosettaCM represents a precise and resource efficient tool for protein structure prediction. Rosetta applications rely on the Monte-Carlo Optimization, which is a probabilistic approach to finding a local minimum in the energy landscape of protein conformations. The underlying equation serving as the fundament of the statistical Monte-Carlo method is the Metropolis acceptance criterion:

where kB is the Boltzmann constant, ΔE the difference in
energy of the two states and T the temperature. The term kBT can also
be written as a single factor β.
During the statistical protein folding based on the Monte-Carlo method, the initial structure is changed by small random perturbations of the atom locations. Whether the structure is accepted or not is decided by the Metropolis acceptance criterion. If ΔE < 0, the structure is accepted, otherwise the newly proposed structure is accepted with probability p as described in the Metropolis acceptance criterion.
Procedure
The RosettaCM protocol requires evolutionary related structures and sequences, as well as fragment files of the target structure. The fragment files serve as a structure template for the proteins and they consist of peptide fragments of sizes 3 and 9. We gathered five evolutionary related structures from the RCBS PDB with the accession numbers:
- 1ija
- 1itw
- 1itp
- 1ito
- 2mlm
The five RCBS entries represent different structures of sortases from Staphylococcus aureus. Fragment files can be created with the Robetta online server or with the Rosetta FragmentPicker application.
The RosettaCM procedure is best described in the following steps:
- sequence and structural alignment of templates
- fragment insertion in unaligned sections
- replacement of random segment with segment from a different template structure
- energy minimization
- all-atom optimization
The alignment can be performed with various tools. We used MAFFT to generate the multiple sequence alignments. Prior to using the alignments as an input, they were converted to the grishin alignment format as RosettaCM requires the alignments to be in said format. The minimization is performed using the Rosetta controid energy function. For the centroid function to be applied, the protein is converted to the centroid representation. A protein in centroid representation consists of the backbone atoms N, Cα;, OCarbonyl and an atom of varying size representing the side chain. The advantage of using the centroid representation is that the energy landscape can be traversed easier due to the smoother nature of the centroid energy landscape. Finally the generated structure undergoes a second minimization in an all-atom model by means of Monte-Carlo optimization. This is similar to the energy minimization but without the amino acids being represented as centroids of their functional groups. Structures computed through all-atom optimizations can reach atomic resolutions {{Quelle rosetta paper}} which is crucial for a model meant to be used to estimate atomic interactions.
Results
The run yielded 15,000 structures which have been compared using the Rosetta scoring functions (talaris2013). From the 15,000 structures generated, we inspected the ten best scoring structures.
As can be seen in figure 5, the most prominent differences can be found in the regions close to the N- and C-terminus. As fluctuations in those regions are not untypical, we decided to use the best scoring structure, candidate S_14771 (figure 6), as the input for the simulations to follow.

Figure x : The structural alignment of the ten best scoring sortase structures displaying minor differences with the exception of the C- and N-terminal regions. N- and C-terminal regions tend to show strong fluctuations, thus it is unsurprising to find the terminal regions to be unaligned

Figure x : Sortase A7M candidate S_14771 created through RosettaCM.
In order to evaluate the secondary structure of the Sortase A7M candidate S_14771 a Ramachandran plot has been created and compared to the five sortases used as input for the comparitive modeling. Comparisons were also drawn with the Sortase predicted by Deep Learning as well as a database of randomly sampled proteins. Ramachandran plots of dihedral angles (fig x) can be a first indicator whether the structures computed are valid.

Figure x : Caption?

Figure x : Caption?

Figure x : Caption?
Figure 5: The comparison of the ramachandran plot of structure S_14771 and the ramachandran plot found on Protopedia suggests that secondary structures are present. Hence the structure appears to contain α-helices, β-sheets and a small amount of lefthanded α-helices.
Conclusion
We used machine learning methods, as well as monte-carlo simulations
to
determine the structure of the mutated transpeptidase Sortase A7M.
The machine
learning approach using AlQuarishi's Deep Neural Network yielded a
structure which seemed to
not have any secondary structures. To exclude the possibility of an
error in the
PyMOL visualization software by Schroedinger, a Ramachandran plot
(figure xyz)
was created. The plot shows that no typical secondary structures are
present
which is a strong indicator of a failed approach to determine a
structure.
The approach, using Rosetta Comparative Modeling, yielded
15,000
structures scored with the talaris2013 scoring function. The ten
best structures
were aligned and exhibited almost identical secondary structures
(figure xzy).
The greatest structural differences are present in the N- and
C-terminal
regions. Since terminal regions tend to fluctuate more strongly than
non-terminal segments of the protein, we deemed those fluctuations
non-relevant
for the proteins functionality.
Being the best scoring candidate, structure S_14771 was analyzed
structurally
using a Ramachandran plot (figure xyz). The plot shows all the
relevant and
typical structures sortases exhibits and serves as an indicator for
a
successful structure prediction.
In the steps to follow, a molecular dynamics (MD)
simulation will be performed on both structures. Even though
structure CASP12
does not seem to be a valid structure, refolding processes during a
MD
simulation might lead to a relaxation of the protein and allow for a
promising
prediction of the sortase A7M structure.
Molecular dynamics
Introduction
The structure predictions made so far were based on statistical methods with physical constraints. The Deep Learning algorithm uses a neural network trained to find a function associating the amino acid sequence and the final 3D positions of the atoms within the protein. On the other hand, predictions were made with Rosetta using the Monte Carlo Method. Here random movement of individual atoms occurs, and the energy is estimated after each step.
Even though both methods use physical constraints to find plausible protein structures, neither of them actually simulates the behavior of these molecules within a physical force field. Moreover, both methods do not necessarily output fully relaxed protein structures and simulate water implicitly by preferring hydrophilic parts of the proteins to be on the outside. Thus, we conducted a molecular dynamics (MD) simulation to verify the plausibility of our protein structure and allow equilibration. The molecular dynamics simulation provides the opportunity to simulate water as discrete molecules, creating a solvated protein. This step is crucial to validate the structures, as the interaction with water is one of the primary mechamism for protein folding. Since neither candidate CASP12 nor S_14771 have been modeled with explicit water an according MD simulation is imperative, to verify the correctness of the candidates conformation. This of course is much more expensive in terms of computational ressources. As the protein has to be placed in a simulation box and said box is filled with water molecules. This is called solvation and is visualized for candidate S_14771 in figure eeeeee.

Figure x : Sortase A7M in a force field surrounded by discrete water molecules. Image was made with gmxSolvate.
We used GROMACS (GROningen MAchine for Chemical Simulations) as the tool for our molecular dynamic simulations. GROMACS solves Newtons equations of motion for individual atoms [1] . While this classical simulation is much more accurate than predictions made by the other methods, approximations are used nonetheless: Forces are cut after a certain radius and the system size is quite small. [1] Additionally, atoms are assumed to be classical particles, which is not the case, as quantum mechanics plays a role in particle-particle interactions. Still, this simulation is very computationally expensive. Therefore, only time periods less than one second could be simulated.
Methods
To perform the molecular dynamics simulations we mostly followed the GROMACS Lysosome tutorial as it serves our purpose perfectly. We created our simulation box to be of dodecahedral shape and a 0.7 nm distance of the solute to the box borders. We used periodic boundry conditions and a Na+ Cl- concentration of 0.012 mol/L. The main difference of our approach was that we used the CHARMM36 force field instead of the OPLS-AA/L force field and have adjusted our molecular dynamics parameters accordingly. The simulation was performed on a NVIDIA GTX 760 graphics card allowing us to simulate approximately 1 ns per hour.
To analyse the MD simulation we used the Python programming language and the Biotite package as well as GROMACS analysis tools as covar and anaeig. The first analyses are a root-mean-square deviation (RMSD), a root-mean-square fluctuation (RMSF) and a gyration radius analysis. RMSD calculations have been described in the structure prediction section. To compute the RMSF the movement distance of each residue is computed as a root-mean-square over time as:

Figure x : caption
where v(t)i is the position of atom i at time t. The radius of gyration is The final analysis performed on the MD simulation is called Principle Component Analysis (PCA). By applying PCA to a protein it is possible to gain insights into the relevant vibrational motions and thereby the physical mechanism of the protein .
Results
The first possible indicators of a stable protein structure are converging root-mean-square deviation (RMSD), small root-mean-square fluctuation (RMSF) values as well as converging radii of gyration. Using the Python software package and the module Biotite we calculated these quantities and plotted the results for both candidate S_14771 and candidate CASP12.

Figure x : The RMSD is one of three main indicators of a stable protein structure of the MD simulation of S_14771 over the period of 200,000 ps. As time progressed the RMSD increased with a smaller slope. The value stabilizes at a time of 110,000 ps and fluctuated around the value of 6 Å.

Figure x : At t = 40,000 ps already the RMSD has arived at a stable value, while at the same time the gyration (fig x) radius decreases over time continuously. This information suggests the protein might be folding and potentially develpoing secondary structures not present previously.

Figure x : The prominent fluctuations of the residues from ranges 105 to 115 might indicate a binding site or another form of functional structure. The radius of gyration, just as the RMSD fig xyz, stabilizes around a simulation time of of 110,000 ps and converges towards a value of 16.7 Å.

Figure x : As from t = 40,000 ps the radius of gyration decreases constantly. At the end of the simulation the gyration radius reaches a value of 17 Å. This behavior indicates folding of the protein structure.

Figure x : The fluctuations (RMSF) of most residues appear insignificant compared to the first, the last residues and the residues close to residue 110 . Typically the N- and C-terminus tend to fluctuate more intensively due to the lack of stabilizing structures. The prominent fluctuations in the range of residue 105 to 115 can indicate a binding site or another form of functional structure.

Figure x : The prominent fluctuations of the residues from ranges 105 to 115 might indicate a binding site or another form of functional structure. The radius of gyration, just as the RMSD fig xyz, stabilizes around a simulation time of of 110,000 ps and converges towards a value of 16.7 Å.
Typical RMSDs and radii of gyration converge towards a value dependent on the size of the protein. Convergence of those quantities can be interpreted as a stable state of the protein structure. As it can be seen in Figures x and y both the RMSD and the radius of gyration stabilize at the same time as the simulation reaches 110,000 ps (110 ns), suggesting a now stabilized structure of candidate S_14771 solvated in water. Another indicator of a functional protein is the RMSF. Instead of being averaged over all atoms, the RMSF is averaged over time with respect to each amino acid. It provides insights in both protein stability and functionality. Fig xzf reveals the RMSF of residues 105 to 115 to be significantly higher than that of other residues. This hints at the presence of a functional unit along these residues. As commented on in the section describing our structure prediction approaches, the N- and C-terminal regions tend to fluctuate more strongly as a result of the absence of stabilizing structures.
RMSD and gyration of radius calculations of candidate CASP12 (figures x and y) provide evidence of folding. However, the RMSF values show values significantly higher, an effect possibly caused by instability or refolding. Nevertheless, the strongest fluctuations, disregarding the terminal regions, can be seen in the region of residue 105 to 115. This insight consolidates the theory that residues 105 to 115 might be a part of a functional unit.
We were unsure whether candidate CASP12 can be considered a plausible structure and how to interpret the findings concerning the prominent fluctuations. Therefore, we decided to perform a Principle Component Analysis.
Principle component analysis
To analyze our system further Principle Component Analysis (PCA) was performed using GROMACS.

Animation 4: A Principle Component Analysis of a fast (blue) and a slow (red) mode showing the most prominent movements of the Cα-chain of candidate S_14771. Both modes show movement of the β6/β7 loop consisting of residues 105 to 115 towards the active site . Thus we can assume that the closing β6/β7 loop is involved in the reaction mechanism.
The results from the Principle Component Analysis of candidate S_14771 (animation xy) show a movement of the residues 105 to 115 towards the active site, supporting our theory that residues 105 to 115 are important for the reaction mechanism. Since the slow mode (red), which shows the most relevant movement of the sortase, moves further towards the active site, it is possible that the β6/β7 loop either closes the binding site of the ligand peptides or even transports one peptide towards the other.
Animation xyz shows the results of the Principle Component Analysis of candidate CASP12. As the RMSF calculations suggested (fig xyz), the whole protein seems to be moving randomly with no directed movement. In addition the active site amino acids are spread across the protein confirming our assumption that the protein is not in a stable or plausible conformation.
Conclusion
We gained evidence that at least on of our Sortase A7M models is a valid and stable candidate by performing various methods to analyse the structural stability and validity of our two Sortase A7M candidates. The candidate S_14771 that was generated using RosettaCM appears to be a fitting candidate not only due to successful analyses, but also since the residues of the active site are close enough to each other to catalyze a ligation reaction. Our model created through deep learning excelled only in terms of RMSD and gyration radius calculations. Not only the RMSF and Principle Component Analysis but also the conformation of the active site have proven candidate CASP12 to be of no use for further calculations as it does not portray a valid conformation of Sortase A7M.
Docking
Now that the binding site of the Sortase had been found, the peptide ligand needed to be inserted into the binding site to create a peptide-protein complex. The procedure of choice for the introduction of a ligand into the binding site of a protein is called docking. In the following sections, we will present the protocol and methods we used as well as the results they yielded.
Background
Enzymes are one of the most relevant macromolecules in biology. Their functionality is determined through the way they interact with their ligands. Although enzymes are highly specific concerning the ligands they interact with, similar compounds can often bind to the same enzyme albeit with different affinity. To determine the best possible binding conformation of the protein-ligand complex, we use FlexPepDock, an algorithm provided by the the RosettaCommons software package.
Procedure
The ab-initio FlexPepDock protocol consists of multiple steps and is documented on the RosettaCommons online documentation. We modified the protocol as the one provided did not work with our approach. The modified protocol has the following form:
- secondary structure determination
- complex creation
- FlexPepDock refinement
To determine the secondary structure of the peptide, fragment files (3- and
5-mers) had to be generated and a PSIPRED secondary structure prediction had to
be performed. As the peptides had a sequence length less than 20 amino acids, we
were not able to use the online services such as Robetta and the PSIPRED online service.
Instead we used the Rosetta FragmentPicker
application and the PSIPRED command line tool.
The generated structures serve as the input for the refinement protocol.
The generation of the peptide-protein complex can be divided into three steps:
- peptide creation
- peptide relaxation
- coarse complex creation
The peptide structure was created through ab-initio modeling.
Initial creation of the peptide was followed by insertion of the peptide into
the sortase binding site. This lead to a coarse model of the peptide sortase
complex. Here we used insight gained from the molecular dynamics simulation to
place the peptide close to the binding site.
In the final step the FlexPepDock refinement protocol is executed and 50,000
complex structures are generated. We used the inputs as described in
{{fuhrman paper}}, written by the authors of the FlexPepDock documentation.
To get a better overview over our data we performed a clustering in python,
using the scikit-learn package. We clustered the structures with respect to:
- total score: the total score of the docking provided by the Rosetta scoring function
- interface score: the sum of the energy of the residues in the interfacing region
- reweighted score: a score calculated by double weighting the contribution of the residues in the interfacing region
- root-mean-square deviation: the root-mean-square deviation of the peptides in relation to the structure with the highest score
- peptide direction: the direction the peptide is facing
Here clustering is used to group the docking results and thereby descrease the samlple size. From the 50,000 results we picked the results with the 500 best total scores, the 500 best interface scores and the 500 best reweighted scores. As we aimed to create an unbiased set for clustering, the abscence of duplicates in the set was ensured. We decreased the sample size to 100 groups representing the best scoring structures from the three categories.
Results
For sequences MGGGGPPPPPP(M-polyG), GGGGPPPPPP(polyG) and PPPPPPLPETGG(LPETGG) 50,000 structures have been created and clustered. After the clustering the sample consisted of 100 structures of docked complexes.

Figure x : The three best scoring structures (total score, interface score, reweighted score) of the LPETGG-tag are shown. Only two results are visible as the best reweighted score candidate is identical to the best interface score candidate. The reacting section of the LPETGG-tag namely glycine is colored yellow as is the active site. The glycin of both ligand peptides is facing the active site.
Analysis of the scores has shown a similar score for all the three dockings. The best scoring results of the LPETGG docking show a tendency of the glycines to face the active site while also being in close proximity to the active site.

Figure x : The three best scoring structures (total score, interface score, reweighted score) of the poly-g peptide are shown. Only two results are visible as the best reweighted score candidate is identical to the best interface score candidate. Instead of facing the active site (yellow) the reacting glycines (yellow) appear to interact with the β6/β7 loop of the sortase.

Figure x : The three best scoring structures (total score, interface score, reweighted score) of the poly-g peptide are shown. Only two results are visible as the best reweighted score candidate is identical to the best interface score candidate. Concerning the M-poly-G peptide no uniform directional orientation can be observed. The structure with the best interface score (light blue) is oriendted towards the loop while the structure with the best total/reweighted (dark blue) is oriented towards the β-sheets.
Figure lpetgg shows the docking result of the LPETGG peptide to the sortase. The results shown are the best scoring structures of the clustering with respect to the total score, interface score and reweighted score. As the best scoring structure is the same for the total score and the reweighted score only two peptides are shown. This also applies to figures x and y. For both results the reacting glycin residues (yellow) are facing the active site. Additionally, the same residues are in close proximity to the active site.
The figures x ad y show the docking of the both polyG and M-polyG. While polyG results align well and seem to be interacting with the β6/β7 loop rather than with the active site, this does not seem to be the case for M-polyG. Instead of both structures interacting with the β6/β7 loop or active site one (best interaction score; dark blue) interacts with the β6/β7 loop and the other (best reweighted/total score; light blue-gray) appears to interact with the active site.

Figure x : The close up of the M-polyG peptide (best total/reweighted score) indicates an interaction of methionine with arginine139 and cysteine126.

Figure x : Methionine of the result with the best interface score interacted with the β6/β7 loop rather than the active site. Still the reactive glycine residues appear to be bound to the β6/β7 loop.
As can be seen in figure 16 visualizing the result of the the docking simulation total/reweighted score) suggests an interaction of methionine and two of the active sites namely arginine139 and cysteine126. Visualizing the result of the according docking simulation, as can be seen in figure 16, suggests an interaction between methionine and two active site residues, namely arginine139 and cysteine126. Figure 17 shows the interaction of M-polyG with the β6/β7 loop. The glycines still interact with the β6/β7 loop. Instead of binding above the β6/β7 loop, which is the case for polyG as illustrated in fig z, the interaction seems to be influenced by methionine. By interacting with the residues in the β-helix methionine could potentially hinder binding of glycine to the β6/β7 loop by partial immobilization of the peptide. Overall peptide binding and orientation is less uniform compared polyG without the leading methionine, which could be an indicator of lesser binding affinity of M-PolyG towards the β6/β7 loop.
Conclusion
To computationally investigate binding affinities of the polyG and M-polyG as well as the LPETGG tags we performed docking simulations using the Rosetta FlexPepDock application. We used a modified version of the recommended protocol as the modified version was easier to automate and served our purpose better than the standard protocol. From the calculated scores only, we could not see a difference in binding affinities. Thus, we inspected the best scoring structures regarding the total score, the interface score and the reweighted score using PyMOL. Since the best structures with respect to total score and reweighted score were the same for all simulations, only two structures have been inspected per run. A polyproline tag was appended to all the peptides to simulate the modification of the VLPs with a small peptide.
As expected, the results showed that for LPETGG, the glycines of both peptides oriented towards the active site. This is unsurprising as peptides with the sequence LPXTGG are known to be substrate of the Sortase. It was more surprising to see the polyG tag oriented away from the active site since polyG also is a known substrate of the sortase. Both polyG peptides were facing the β6/β7 loop (residues 105 to 115) uniformly and appeared to be interacting with it. The M-polyG peptides did not show a uniform orientation or interaction scheme. On one hand the visualization of the best result concerning the total and reweighted score has shown interaction of methionine with the cysteine126 and arginine139, two residues of the active site. On the other hand, the visualization of the best result with respect to the interface score shows the M-polyG facing the mobile β6/β7 loop. In contrast to the polyG peptide the lacking the methionine, the M-polyG peptide is pulled down below the β6/β7 loop by the methionine interacting with one of the β-sheets leading to the active site. This is not the case with the polgG results, which lie aligned in one plane with the β6/β7 loop.
Modeling Conclusion
generic filler text
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21INCOMPATIBLE WITH RFC[21]Illegal XhoI site found at 445
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
