Difference between revisions of "Part:BBa I746909"
| Line 7: | Line 7: | ||
<html> | <html> | ||
| − | <h2> | + | <h2> iGEM TUDelft 2019 </h2> |
| − | <p> <a href="https://2019.igem.org/Team:TUDelft">2019 iGEM | + | <p> <a href="https://2019.igem.org/Team:TUDelft">2019 iGEM TUDelft</a>characterized this part in PURE<sub>frex</sub>.<br><br></p> |
| + | ==Overview== | ||
| + | We have added characterization to https://parts.igem.org/Part:BBa_I746909 by performing an assay of the plasmid in the PUREfrex® cell-free expression system. This experiment allows for RNA levels to also be detected in tandem with protein levels, all in a predetermined T7 RNAP concentration, allowing for a more detailed understanding of the kinetic behavior (i.e. transcription and translation) of the construct. We believe the use of the PURE system in measurements like this contribute to the development of the iGEM cell-free chassis and to obtain better parameterization of the kinetics of genetic constructs like this one. | ||
| + | ==Method== | ||
| + | The following PUREfrex® system solutions were used: | ||
| + | Solution 1: Aminoacids, NTPs, tRNAs, and enzymatic substrates. | ||
| + | Solution 2: Proteins involved in T7 RNAP based transcription, in 30% glycerol. | ||
| + | Solution 3: 20 μM ribosomes, allowing for the translation of proteins. | ||
| + | In a 96 well plate we added the following components: | ||
| + | Blank: Solution A + SYBR Green (?). | ||
| + | DNA background: Solution A + Plasmid DNA + SYBR Green | ||
| + | RNA measurement: Solution A + Plasmid DNA + Solution B + SYBR Green. | ||
| + | Protein measurement: Solution A + Plasmid DNA + Solution B + Solution C. | ||
| + | We added 120 ng of DNA to all wells except for well 1.SYBR Green was used for detecting RNA and DNA concentrations. GFP fluorescence was also measured in the last well, which contained all 3 solutions of the PURE system. | ||
| + | Measurements were taken every 5 min for a total period of 7 hours. | ||
| + | For analysis, the DNA background (well 2) was subtracted from well 3 to obtain the kinetic growth curve of RNA transcripts | ||
| + | After the reaction was run, the final RNA concentration was isolated from well 3 and quantified using nanodrop, by taking three measurements and calculating the average. Assuming 100% efficiency, the measurement was used to determine the nanogram levels of RNA over the reaction`s time. | ||
| + | |||
| + | |||
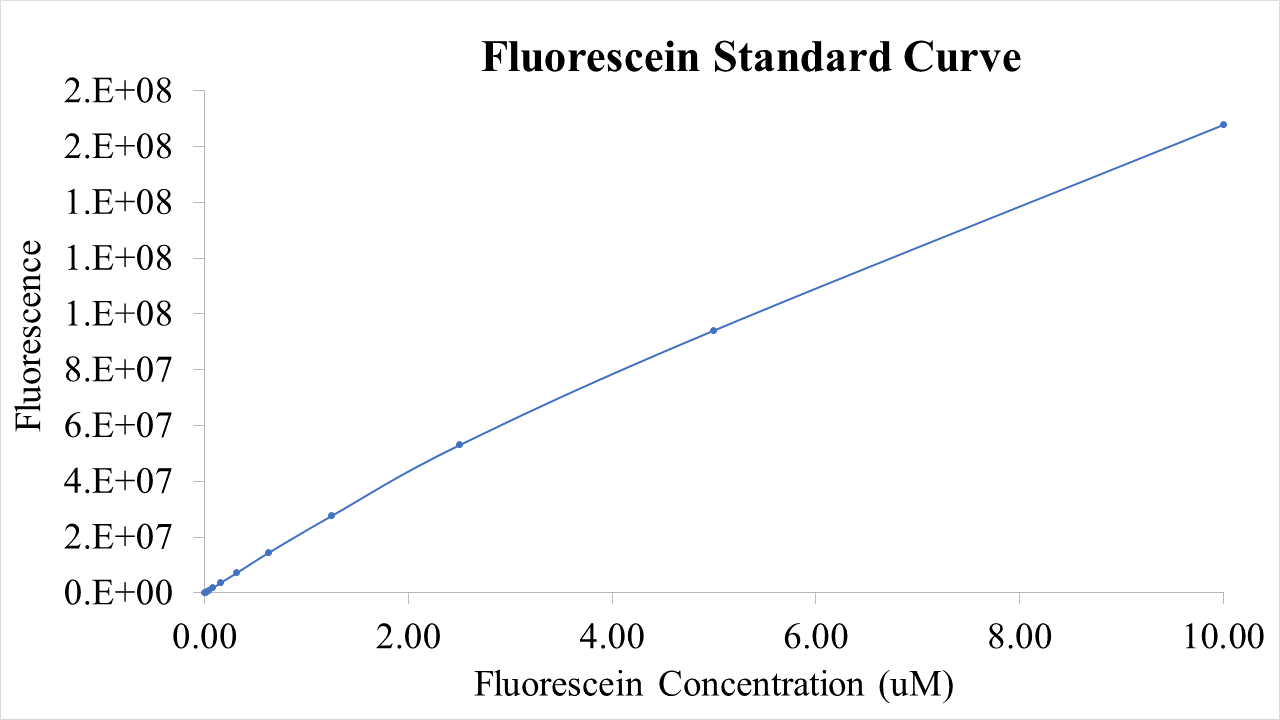
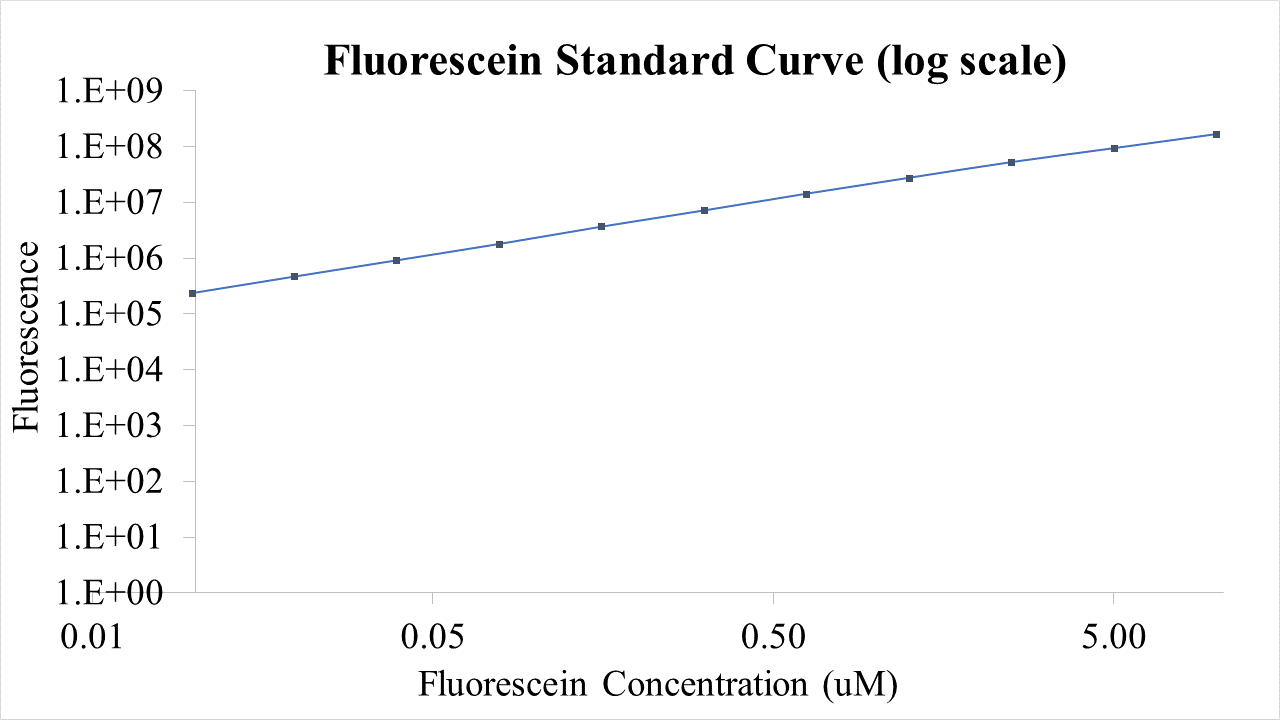
| + | ==Results== | ||
| + | Final RNA level: 267.7 +- 2.75 ng The following graphs were generated and a logistic growth curve was fitted to the data. A delay in protein translation can be observed when | ||
</html> | </html> | ||
Revision as of 12:16, 19 October 2019
superfolder GFP driven by T7 promoter
This is one of the constructs used to characterise superfolder GFP (see I746916 part description and http://openwetware.org/wiki/IGEM:Cambridge/2008/Improved_GFP for source and other information about this GFP variant)
iGEM TUDelft 2019
2019 iGEM TUDelftcharacterized this part in PUREfrex.
Characterization: PR4 vs. T7
Group: Queens_Canada, 2019
Author: Ruben Warkentin
Summary: We compared the amount of GFP expressed under a constitutive promoter (medium promoter, strong RBS) to T7 expression. Some proteins fold better under constitutive promoters; however, nobody had yet directly compared the amount of protein produced between constitutive vs. T7 expression.
Methods
BioBricks were transformed and expressed in E. coli (BL21). BL21 cells were cultured to an OD600=0.6 and 100 uL of culture was transferred into a 96 well plate. Colonies were transfered in quadruplicate. The fluorescence intensity of GFP was measured with a multi-mode microplate reader. The iGEM standardized fluorescence protocol was used for fluorescence measurement standardization (https://www.protocols.io/view/calibration-protocol-plate-reader-fluorescence-cal-6zrhf56).
Results
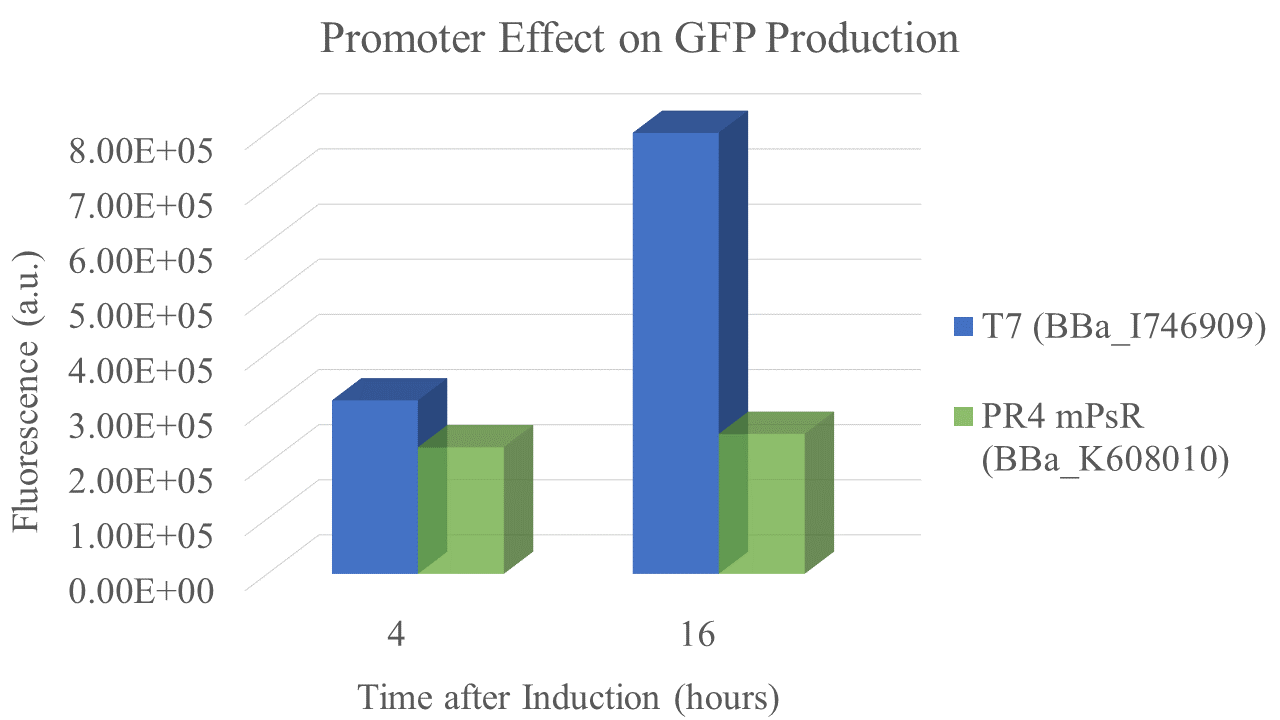
We found that the T7 promoter produced about 2.6 times as much fluorescent signal as the constitutive PR4 promoter, indicating that T7 is much more efficient at producing GFP (Fig. 5). Interestingly, the production of GFP under PR4 did not increase beyond the level observed at 4 hours after inoculation. It seems that PR4 leads to an initial production of protein; however, after the initial expression the promoter seems to be shut off.
Promoter and RBS:
PR4: medium Promoter (J23110) strong RBS (B0034)
T7: T7 Promoter (BBa_I746909)
| sample | PR4 | T7 |
| Fluorescence 4 hrs (a.u.) | 2.24E+05 | 3.09E+05 |
| Fluorescence 16 hrs (a.u.) | 2.48E+05 | 8.00E+05 |
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal SapI.rc site found at 62