Difference between revisions of "Part:BBa K3147000"
(→II. Proof of function) |
(→I : parts BBa_K3147000 (Pc-sfGFP-TEVcs-SSRA) function :) |
||
| Line 5: | Line 5: | ||
===I : parts BBa_K3147000 (Pc-sfGFP-TEVcs-SSRA) function :=== | ===I : parts BBa_K3147000 (Pc-sfGFP-TEVcs-SSRA) function :=== | ||
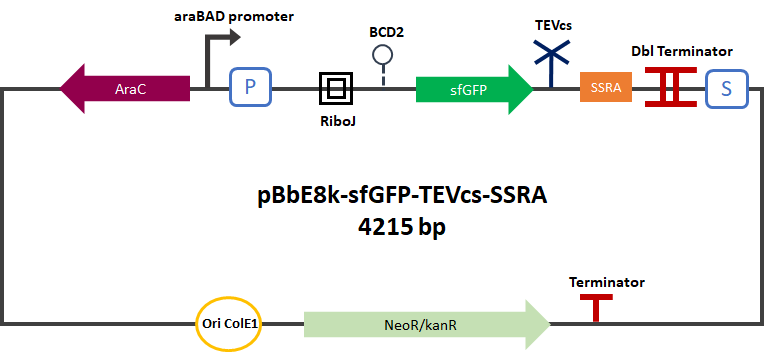
| − | The Montpellier 2019 team submitted a reporter gene construction in order to carry out their proof of concept. This construction produces an sfGFP(bs) [1] [2] [3] (BBA_K1365020) merged into C-ter with a fast degradation tag called ssrA [ | + | The Montpellier 2019 team submitted a reporter gene construction in order to carry out their proof of concept. This construction produces an sfGFP(bs) [1] [2] [3] (BBA_K1365020) merged into C-ter with a fast degradation tag called ssrA [5] (BBA_M0050). The TEV cutting site (BBa_J18918) has been added between the sfGFP and the ssrA tag. This construction can be used as a reporter gene. In the presence of TEV the ssrA is separated and allows the sfGFP to glow without being degraded. |
<div align="center">[[File:Pc-sfGFP-TEVcs-SSRA2.png|650px]]</div> | <div align="center">[[File:Pc-sfGFP-TEVcs-SSRA2.png|650px]]</div> | ||
Revision as of 16:00, 14 October 2019
sfGFP fuse to TEV-cleavable ssrA tag
I : parts BBa_K3147000 (Pc-sfGFP-TEVcs-SSRA) function :
The Montpellier 2019 team submitted a reporter gene construction in order to carry out their proof of concept. This construction produces an sfGFP(bs) [1] [2] [3] (BBA_K1365020) merged into C-ter with a fast degradation tag called ssrA [5] (BBA_M0050). The TEV cutting site (BBa_J18918) has been added between the sfGFP and the ssrA tag. This construction can be used as a reporter gene. In the presence of TEV the ssrA is separated and allows the sfGFP to glow without being degraded.
II. Proof of function
The experimental approach to test the activity of these reporters was to compare the basal fluorescence rate of the sfGFP with a TEV recognition site "cleaved" to this construction. To do so, we made a control construction: sfGFP-TEVcs(BBa_K3147001) similar to this one by removing the proteolysis tag and simulating a cut by the TEV.
We expressed this part in a plasmid with an arabinose promoter : pBbE8K-RFP backbone (https://www.addgene.org/35327/). We cloned it by Gibson Assembly.
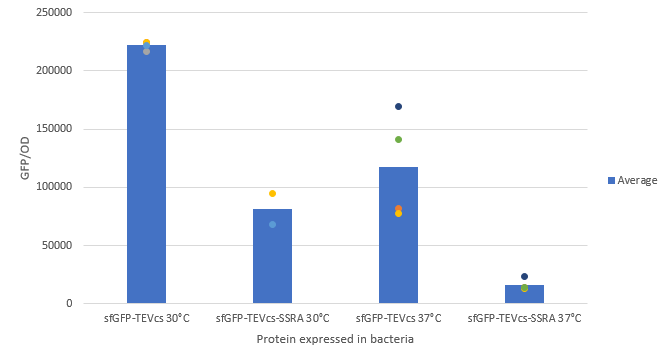
We compared the basal fluorescence of the E. coli strain NEB10β transformed with the sfGFP-TEVcs construction and the E. coli NEB10β transformed with the sfGFP-TEVcs-ssrA construction. Fluorescence was quantified after induction with arabinose concentrated at 1% with the plate reader all night. Here are the fluorescence of the sfGFP-TEVcs-ssrA and the of the sfGFP-TEVcs at 30 and 37°C. We can see that the ssrA tag is causing a lot of degradation of the protein.
Reference:
[1] McGinness, Baker, Sauer. 2006. Mol. Cell. 22:701.
[2] Overkamp, W. et al. (2013) Benchmarking various green fluorescent protein variants in Bacillus subtilis, Streptococcus pneumoniae, and Lactococcus lactis for live cell imaging. Appl. About. Microbiol. 79: 6481-6490
[3] Sarah Guiziou et al. 2016. “A part toolbox to tune genetic expression in Bacillus subtilis” Nucleic Acids Research, 2016, Vol. 44, No. 15 7495–7508.
[4] Fernandez-Rodriguez, Jesus, et Christopher A. Voigt. 2016. « Post-TranslationalControl of Genetic Circuits Using Potyvirus Proteases ». Nucleic Acids Research 44(13): 6493‑6502.
[5] Sunohara, T., Abo, T., Inada, T., & Aiba, H. (2002). The C-terminal amino acid sequence of nascent peptide is a major determinant of SsrA tagging at all three stop codons. RNA (New York, N.Y.), 8(11), 1416–1427. doi:10.1017/s1355838202020198
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 41
Illegal NheI site found at 53
Illegal NheI site found at 76 - 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 82
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]