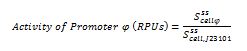Difference between revisions of "Part:BBa J23107:Experience"
| Line 58: | Line 58: | ||
| − | + | ||
| − | + | ||
| − | + | ||
<partinfo>BBa_J23100 AddReview 5</partinfo> | <partinfo>BBa_J23100 AddReview 5</partinfo> | ||
<I>University of Texas at Austin iGEM 2019</I> | <I>University of Texas at Austin iGEM 2019</I> | ||
Revision as of 19:17, 23 September 2019
This experience page is provided so that any user may enter their experience using this part.
Please enter
how you used this part and how it worked out.
Applications of BBa_J23107
Evaluation of Anderson promoter J23107 in B. subtilis by iGEM-Team LMU-Munich 2012
This Anderson promoter was evaluated without fused RFP with the lux operon as a reporter in B. subtilis. See the new BioBrick BBa_K823009 without RFP and have a look at the [http://2012.igem.org/Team:LMU-Munich/Data/Anderson Data] from the evaluation in B. subtilis.
User Reviews
UNIQ8788572844811bad-partinfo-00000000-QINU UNIQ8788572844811bad-partinfo-00000001-QINU
iGEM CINVESTAV_IPN_UNAM CHARACTERIZATION OF IGEM DISTRIBUTION BIOPARTS
For contribute to the parts registry our team decided to make the characterization of constitutive promoters, in E. coli, belonging to the family isolated from a small combinatorial library (J23101 , J23102, J23104, J23107, J23108, J2311, and J23115) which were attached to GFP, in psB1C3, to determine promoter activity, using the equipment Victor X3 Multilabel Plate Reader.
Fig. 1 Construction of the promoter J23107 expressing GFP.
Methods
With the selected colonies, an overnight culture was made in M9 media(minimal media supplemented with 0.2% CAA). After 12 hours the culture was transferred to a 96 well plate at a 1:10 dilution (20 μl of culture and 180 μL of fresh M9 medium). OD and fluorescence measurements of the selected colonies were performed at intervals of 30 minutes for 16 h. From the results the PopS were calculated (polymerases per second).
Modeling
The ecuations used for calulated de promoter activity were based on (R. K. Jason et. al 2009).
Results
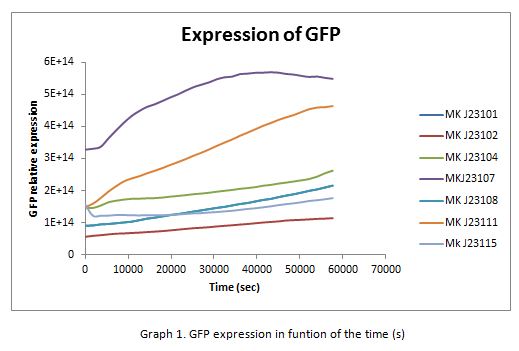
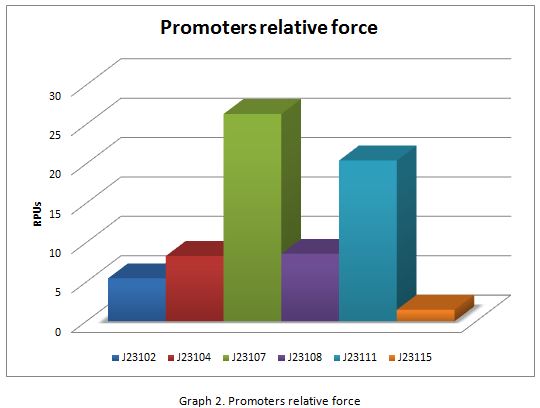
In the following graphs there is shown the GFP expression in function of th time and the realtive promotor intensity.
With the previous results of the characterization of the promoters there is concluded that the promoter J23107, is the strongest because it produces more RPUs”
University of Texas at Austin iGEM 2019 |width='60%' valign='top'|
Characterization of the Anderson series through burden monitoring by the University of Texas at Austin's 2019 iGEM team
Description
Our team transformed the Anderson series of RFP reporters (J23101, J23113, J23104, J23107, J23117 in pSBC13 backbone) into our constitutive GFP burden monitor E. coli strain and measured the relative burden of each part. They contained a constitutive GFP sequence in the genome which serves as a way to measure the constructs’ ribosome allocation. Our results show a certain reduction ingrowth rate for each part as a result of ribosome misallocation away from the genome and towards the plasmid containing the construct. The promoter strengths associated with each RFP reporter construct shows that parts with stronger promoters express less GFP and have a reduced growth rate when compared to the constructs containing weaker promoters. The promoters associated with these RFP reporters were used to create a series of BFP reporters, also transformed into our GFP burden monitor strain, and create the regression on the figure below. For more information on characterization of these parts through burden monitoring and evolutionary stability experiments, visit our team’s wiki page: [1]