Difference between revisions of "Part:BBa K2560261:Design"
| Line 7: | Line 7: | ||
===Design Notes=== | ===Design Notes=== | ||
| − | The part contains the 3' part of the <i>mcr</i> gene of <i>C. aurantiacus</i> (AY530019). This region is flanked by overhangs which are Phytobrick- and MoClo-compatible and by two BsaI recognition sites (Weber et al., 2011). | + | The part contains the 3' part of the <i>mcr</i> gene of <i>C. aurantiacus</i> (AY530019). This region is flanked by overhangs which are Phytobrick- and MoClo-compatible and by two BsaI recognition sites (Weber et al., 2011). It was built with the Marburg Toolbox, a golden gate based toolbox for modular cloning. According to the Marburg Toolbox, the part is designed as a 4-part (CDS). |
| + | |||
It was shown that a splitted version of Mcr with separated C- and N-terminal domains increases enzyme activity (Liu et al., 2013). If these splitted genes shall be used take [https://parts.igem.org/wiki/index.php?title=Part:BBa_K2560262 BBa_K2560262] and this part. If the gene for the whole enzyme shall be used take [https://parts.igem.org/wiki/index.php?title=Part:BBa_K2560260 BBa_K2560260] | It was shown that a splitted version of Mcr with separated C- and N-terminal domains increases enzyme activity (Liu et al., 2013). If these splitted genes shall be used take [https://parts.igem.org/wiki/index.php?title=Part:BBa_K2560262 BBa_K2560262] and this part. If the gene for the whole enzyme shall be used take [https://parts.igem.org/wiki/index.php?title=Part:BBa_K2560260 BBa_K2560260] | ||
Latest revision as of 22:08, 12 October 2018
mcrC - C-terminal domain of Malonyl-CoA Reductase from Chloroflexus aurantiacus
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 288
Illegal NheI site found at 1690 - 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 361
Illegal NgoMIV site found at 1975
Illegal AgeI site found at 1524 - 1000COMPATIBLE WITH RFC[1000]
Design Notes
The part contains the 3' part of the mcr gene of C. aurantiacus (AY530019). This region is flanked by overhangs which are Phytobrick- and MoClo-compatible and by two BsaI recognition sites (Weber et al., 2011). It was built with the Marburg Toolbox, a golden gate based toolbox for modular cloning. According to the Marburg Toolbox, the part is designed as a 4-part (CDS).
It was shown that a splitted version of Mcr with separated C- and N-terminal domains increases enzyme activity (Liu et al., 2013). If these splitted genes shall be used take BBa_K2560262 and this part. If the gene for the whole enzyme shall be used take BBa_K2560260
GGTCTCGAATG-coding_region-GCTTTGAGACC
The sequence was codonoptimized for V. natriegens ATCC 14048. According to Liu et al, we designed the part so that McrN contains amino acid 1-549 of the overall protein and McrC amino acid 550-1219 (Liu et al., 2013).
Source
Source of the part:
Genome: Chloroflexus aurantiacus strain OK-70-fl
Accession number of gene: AY530019, amino acid 550-1219
Accession number of encoded protein: AAS20429
mcrC was codonoptimized for V. natriegens and then synthetisized and integrated into the vector BBa_K2560002 via BsmBI
References
Liu, C., Wang, Q., Xian, M., Ding, Y., Zhao, G., 2013. Dissection of Malonyl-Coenzyme A Reductase of Chloroflexus aurantiacus Results in Enzyme Activity Improvement. PLoS One 8, 1–8. https://doi.org/10.1371/journal.pone.0075554
Strauss, G., Fuchs, G., 1993. Enzymes of a novel autotrophic CO2 fixation pathway in the phototrophic bacterium Chloroflexus aurantiacus, the 3-hydroxypropionate cycle. Eur. J. Biochem. 215, 633–43.
Weber, E., Engler, C., Gruetzner, R., Werner, S., Marillonnet, S., 2011. A Modular Cloning System for Standardized Assembly of Multigene Constructs. PLoS One 6, e16765. https://doi.org/10.1371/journal.pone.0016765
Marburg Toolbox
We proudly present the Marburg Collection, a novel golden-gate-based toolbox containing various parts that are compatible with the PhytoBrick system and MoClo. Compared to other bacterial toolboxes, the Marburg Collection shines with superior flexibility. We overcame the rigid paradigm of plasmid construction - thinking in fixed backbone and insert categories - by achieving complete de novo assembly of plasmids.
36 connectors facilitate flexible cloning of multigene constructs and even allow for the inversion of individual transcription units. Additionally, our connectors function as insulators to avoid undesired crosstalk.
The Marburg Collection contains 123 parts in total, including:
inducible promoters, reporters, fluorescence and epitope tags, oris, resistance cassettes and genome engineering tools. To increase the value of the Marburg Collection, we additionally provide detailed experimental characterization for V. natriegens and a supportive software. We aspire availability of our toolbox for future iGEM teams to empower accelerated progression in their ambitious projects.
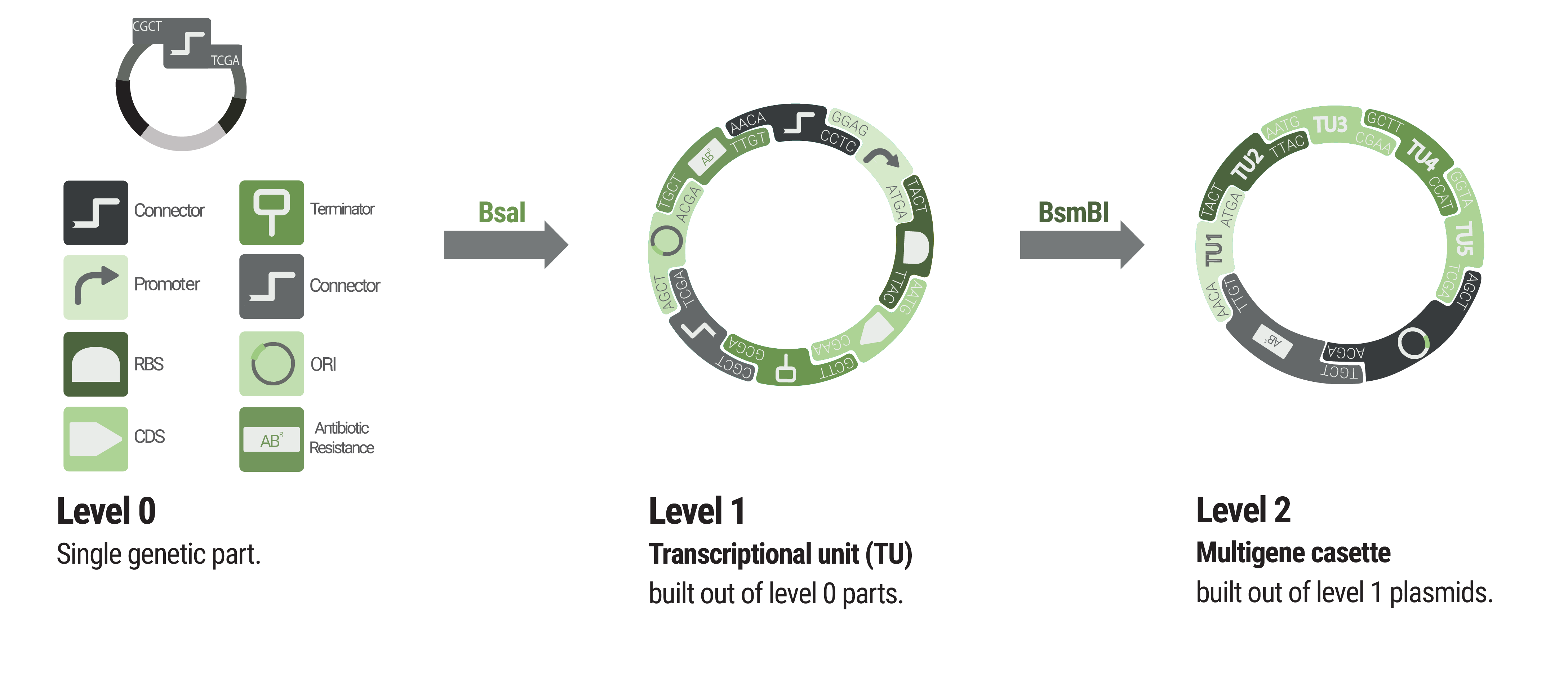
Basic building blocks like promoters or terminators are stored in level 0 plasmids. Parts from each category of our collection can be chosen to built level 1 plasmids harboring a single transcription unit. Up to five transcription units can be assembled into a level 2 plasmid.
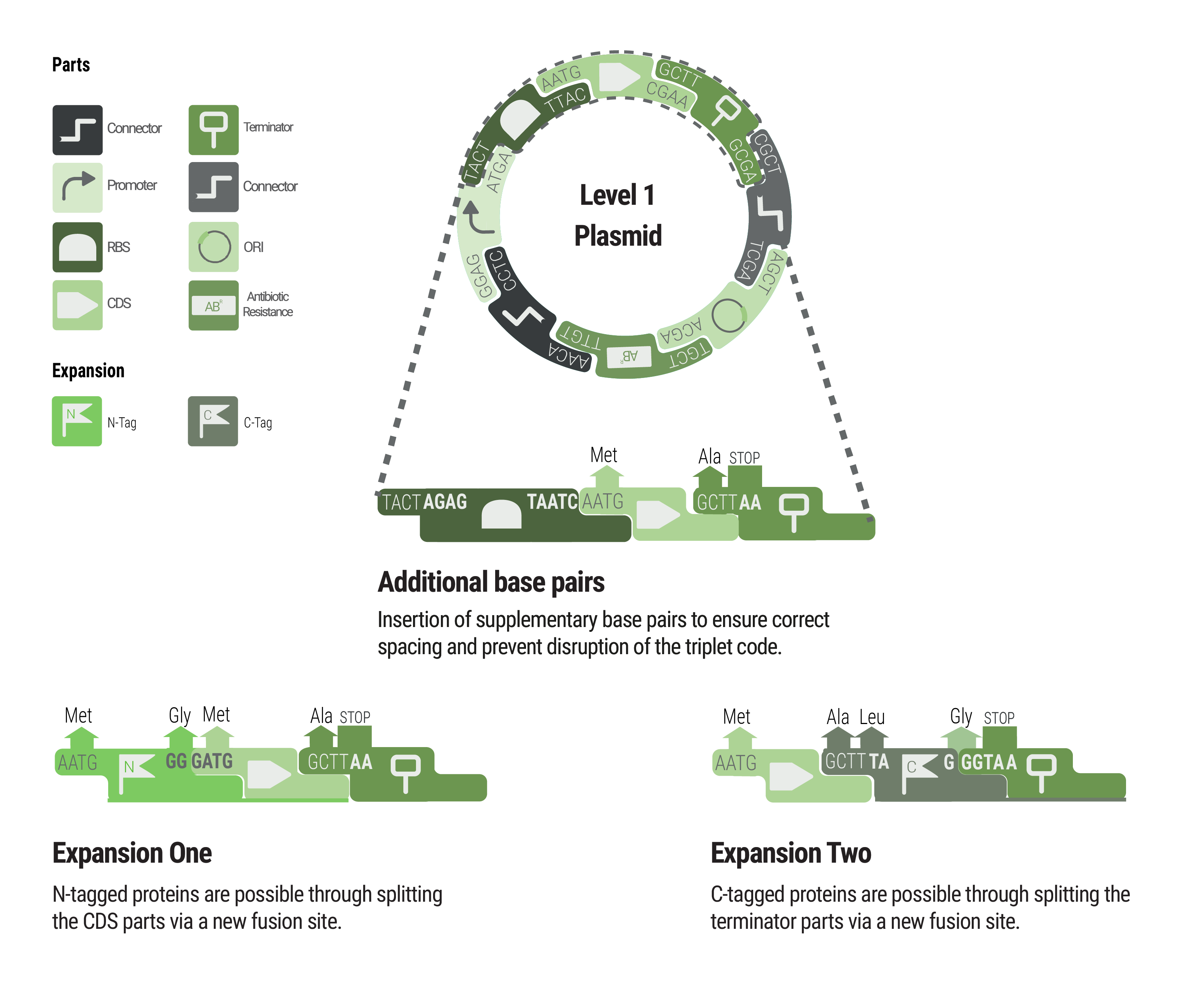
Between some parts, additional base pairs were integrated to ensure correct spacing and to maintain the triplet code. We expanded our toolbox by providing N- and C- terminal tags by creating novel fusions and splitting the CDS and terminator part, respectively.
Parts of the Marburg Toolbox

- K2560011 (5'Connector Dummy)
- K2560055
(1-6
Connector) - K2560065 (5'Con1)
- K2560066 (5'Con2)
- K2560067 (5'Con3)
- K2560068 (5'Con4)
- K2560069 (5'Con5)
- K2560075 (5'Con1
Short Res) - K2560076 (5'Con2
Short) - K2560077 (5'Con3
Short) - K2560078 (5'Con4
Short) - K2560079 (5'Con5
Short) - K2560095 (5'Con1 inv)
- K2560096 (5'Con2 inv)
- K2560097 (5'Con3 inv)
- K2560098 (5'Con4 inv)
- K2560099 (5'Con5 inv)
- K2560105 (5'Con5 inv
Ori) - K2560107 (5'Con1
Res)

- K2560007 (J23100)
- K2560009 (J23104)
- K2560014 (J23106)
- K2560015 (J23115)
- K2560017 (J23101)
- K2560018 (J23102)
- K2560019 (J23103)
- K2560020 (J23105)
- K2560021 (J23107)
- K2560022 (J23108)
- K2560023 (J23109)
- K2560024 (J23110)
- K2560025 (J23111)
- K2560026 (J23113)
- K2560027 (J23114)
- K2560028 (J23116)
- K2560029 (J23117)
- K2560030 (J23118)
- K2560031 (J23119)
- K2560123
(pTet) - K2560124 (pTrc)
- K2560131 (Promoter Dummy)

- K2560012 (3'Connector Dummy)
- K2560070 (3'Con1)
- K2560071 (3'Con2)
- K2560072 (3'Con3)
- K2560073 (3'Con4)
- K2560080 (3'Con5 Ori)
- K2560100 (3'Con1 inv
Short) - K2560101 (3'Con2 inv
Short) - K2560102 (3'Con3 inv
Short) - K2560103 (3'Con4 inv
Short) - K2560104 (3'Con5 inv
Short) - K2560106 (3'Con1 inv
Short Res) - K2560108 (3'Con1 inv)
- K2560109 (3'Con1 inv
Res) - K2560110 (3'Con2 inv)
- K2560111 (3'Con3 inv)
- K2560112 (3'Con4 inv)
- K2560113 (3'Con5 inv)

- K2560048 (Cam. Res. RFP)
- K2560056
(Kan. Res. (pSB3K3) RFP) - K2560057
(Kan. Res. (pSB3K3) GFP) - K2560058
(Tet. Res. (pSB3T5) RFP) - K2560059
(Tet. Res. (pSB3T5) GFP) - K2560125 (Carb. Res. RFP)
- K2560126 (Carb. Res. GFP)
- K2560127 (Carb. Res. into BBa_K2560002)
- K2560132 (Cam. Res. into BBa_K2560002)
- K2560133
(Kan. Res. into BBa_K2560002) - K2560134
(Tet. Res. into BBa_K2560002)
Tags and Entry Vectors







