Difference between revisions of "Part:BBa K1218011:Design"
| Line 20: | Line 20: | ||
==Egypt-AFCM Team Modeling== | ==Egypt-AFCM Team Modeling== | ||
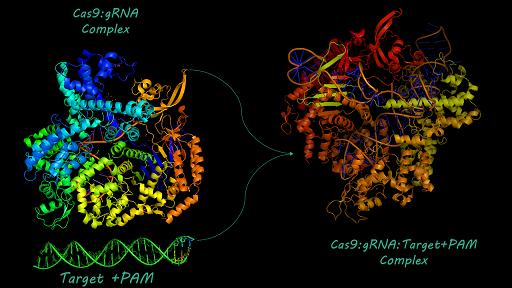
| − | [http://2017.igem.org/Team:AFCM-Egypt# Egypt-AFCM Team] simulated a structural model for [https://parts.igem.org/Part:BBa_K1218011# BBa_K1218011] using SWISS-Model to be ready for protein-nucleic acid docking using HADDOCK algorithm. Docking helped us to have insights about cleavage binding of cas9 to target DNA upstream to PAM site. Docking also revealed mechanisms of interaction between cas9 and gRNA. For more information about [http://2017.igem.org/Team:AFCM-Egypt | + | [http://2017.igem.org/Team:AFCM-Egypt# Egypt-AFCM Team] simulated a structural model for [https://parts.igem.org/Part:BBa_K1218011# BBa_K1218011] using SWISS-Model to be ready for protein-nucleic acid docking using HADDOCK algorithm. Docking helped us to have insights about cleavage binding of cas9 to target DNA upstream to PAM site. Docking also revealed mechanisms of interaction between cas9 and gRNA. For more information about [http://2017.igem.org/Team:AFCM-Egypt# Egypt-AFCM Team] structural Modeling you may visit [http://2017.igem.org/Team:AFCM-Egypt/Model# Egypt-AFCM Modeling] |
Part characterization and usage can be found at this composite part [https://parts.igem.org/wiki/index.php?title=Part:BBa_K2217026# BBa_K2217026_Experience]. | Part characterization and usage can be found at this composite part [https://parts.igem.org/wiki/index.php?title=Part:BBa_K2217026# BBa_K2217026_Experience]. | ||
Latest revision as of 17:26, 26 October 2017
Cas9
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 1642
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 3921
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 4863
Illegal BsaI.rc site found at 4840
Design Notes
Mutagenesis to remove internal EcoRI site
Cas9 for chloroplast
Sequence was codon optimized with software developed at UCL in Saul Purton's lab working on algal biotechnology. Can be found at the following link: https://www.ucl.ac.uk/algae/Genetic_engineering_tools. The sequence was additionally modified to remove all Bsa1 and BsmBI sites (illegal for phytobrick syntax) in the coding sequence and where possible the codon maintained as well as the genetic bias.
No direct source for part, need to be synthesized, sequence based on S. pyrogenes (http://www.uniprot.org/uniprot/Q99ZW2). Due to the large size of the protein, iDT synthesized part in 4 pieces which we were able to assemble through golden gate with the appropriate primers to maintain the coding sequence intact. Alternatively, Gibson could have been used for assembly.
Source
CRISPR-Cas system from streptococcus pyogenes; cloned from plasmids obtained from Bikard et al.
Egypt-AFCM Team Modeling
[http://2017.igem.org/Team:AFCM-Egypt# Egypt-AFCM Team] simulated a structural model for BBa_K1218011 using SWISS-Model to be ready for protein-nucleic acid docking using HADDOCK algorithm. Docking helped us to have insights about cleavage binding of cas9 to target DNA upstream to PAM site. Docking also revealed mechanisms of interaction between cas9 and gRNA. For more information about [http://2017.igem.org/Team:AFCM-Egypt# Egypt-AFCM Team] structural Modeling you may visit [http://2017.igem.org/Team:AFCM-Egypt/Model# Egypt-AFCM Modeling] Part characterization and usage can be found at this composite part BBa_K2217026_Experience.
References
http://nar.oxfordjournals.org/content/41/15/7429 David Bikard, Wenyan Jiang, Poulami Samai, Ann Hochschild, Feng Zhang3, and Luciano A. Marraffini. (2013) Programmable repression and activation of bacterial gene expression using an engineered CRISPR-Cas system. Nucleic Acids Research, 41 (15), 7429-7437.