Difference between revisions of "Part:BBa K907000"
(edit by Lukas) |
|||
| Line 7: | Line 7: | ||
<li>'''Group:''' ETH Zurich 2016 | <li>'''Group:''' ETH Zurich 2016 | ||
<li>'''Author:''' Asli Azizoglu | <li>'''Author:''' Asli Azizoglu | ||
| − | <li>'''Summary:''' We cloned and characterised a codon optimised bxb1 for <i>E.coli</i>, and sent it to the registry as | + | <li>'''Summary:''' We cloned and characterised a codon optimised bxb1 for <i>E. coli</i>, and sent it to the registry as |
| − | a biobrick. Our biobrick can be found | + | a biobrick. Our biobrick can be found with a fast degradation tag [[BBa_K2116009]] and here without any degradation |
| − | + | tag [[Part:BBa_K2116026]]. The non-codon optimised version can be found here [[Part:BBa_K907000]]. Besides some favorable features, the codon optimized bxb1 also supports full assembly compatibility. | |
</ul> | </ul> | ||
| Line 16: | Line 16: | ||
<h2>Genetic Design</h2> | <h2>Genetic Design</h2> | ||
<ul> | <ul> | ||
| − | <li> Bxb1 was expressed under a tet promoter, without any degradation | + | <li> Bxb1 was expressed under a tet promoter, without any degradation tag [[Part:BBa_K2116056]]. The RBS has a strength of 1209.69 au (translation |
| − | rate), as calculated by the [https://salislab.net/software/ Salis RBS Calculator]. This construct was cloned on a medium | + | rate), as calculated by the [https://salislab.net/software/Salis RBS Calculator]. This construct was cloned on a medium |
copy plasmid with the p15A replication of origin. | copy plasmid with the p15A replication of origin. | ||
<li> TetR was expressed under the control of medium-strength constitutive promoter [[Part:Bba_J23118]], and cloned onto a | <li> TetR was expressed under the control of medium-strength constitutive promoter [[Part:Bba_J23118]], and cloned onto a | ||
low copy plasmid with pSC101/Rep101 replication of origin. | low copy plasmid with pSC101/Rep101 replication of origin. | ||
| − | <li> We also constructed a dual fluorescence reporter system [[Part:BBa_K2116024]] that expresses | + | <li> We also constructed a dual fluorescence reporter system [[Part:BBa_K2116024]] that expresses sfGFP [[Part:BBa_K2116017]] when bxb1 flips a directional |
| − | promoter. | + | promoter. In the non-flipped state the reporter expresses the red fluorescent protein mNectarine [[Part:BBa_K2116016]]. |
</ul> | </ul> | ||
<h2 id="experimental_setup"> Experimental Setup </h2> | <h2 id="experimental_setup"> Experimental Setup </h2> | ||
| Line 28: | Line 28: | ||
<li> The test construct with bxb1 was transformed together with the reporter construct, and either with TetR or with the empty | <li> The test construct with bxb1 was transformed together with the reporter construct, and either with TetR or with the empty | ||
backbone as <b>control</b>. | backbone as <b>control</b>. | ||
| − | <li> Cells were grown in LB medium for 4 hours, and then transfered to minimal M9 medium with a 1:100 dilution. aTc was added | + | <li> Cells were grown in LB medium for 4 hours, and then transfered to minimal M9 medium with a 1:100 dilution and grown over-night. The culture was again diluted 1:100 in minimal M9 medium the next day. aTc was added |
at given concentrations at OD<sub>600</sub>. Samples were taken at the specified time points, spun down and resuspended | at given concentrations at OD<sub>600</sub>. Samples were taken at the specified time points, spun down and resuspended | ||
in PBS at OD<sub>600</sub> 0.01 and kept on ice until measurement. | in PBS at OD<sub>600</sub> 0.01 and kept on ice until measurement. | ||
| Line 36: | Line 36: | ||
<h2>Results</h2> | <h2>Results</h2> | ||
<p>We observed that maximum flipping is reached at around 8h under these conditions. The response correlates with aTc concentration. | <p>We observed that maximum flipping is reached at around 8h under these conditions. The response correlates with aTc concentration. | ||
| − | However compared to the control, where there is no TetR, we see 6-fold less flipping. This could be improved by increasing | + | However, compared to the control, where there is no TetR, we see 6-fold less flipping. This could be improved by increasing |
| − | the concentration of aTc, however we have found that at 8000 ng/ | + | the concentration of aTc, however we have found that at 8000 ng/mL cell growth was inhibited.</p> |
[[File:T--ETH Zurich--bxb1kineticsfull.png|500px|thumb|center|Time and dose response of bxb1 flipping a directional promoter. | [[File:T--ETH Zurich--bxb1kineticsfull.png|500px|thumb|center|Time and dose response of bxb1 flipping a directional promoter. | ||
| Line 45: | Line 45: | ||
<h1> Comparison between Codon Optimised and Registry Bxb1 Integrase </h1> | <h1> Comparison between Codon Optimised and Registry Bxb1 Integrase </h1> | ||
<p>We compared the kinetics of our codon optimised bxb1 [[Part:BBa_K2116026]] and the one found previously on the registry | <p>We compared the kinetics of our codon optimised bxb1 [[Part:BBa_K2116026]] and the one found previously on the registry | ||
| − | [[Part:BBa_K907000]] that was not codon | + | [[Part:BBa_K907000]] that was not codon optimized. The same genetic and experimental setup was used as above. The only |
difference between the two constructs is that the non-codon optimised bxb1 has a weaker RBS, with a strength of 251.82au | difference between the two constructs is that the non-codon optimised bxb1 has a weaker RBS, with a strength of 251.82au | ||
(translation rate), as calculated by the [[https://salislab.net/software Salis RBS Calculator]]. Due to this difference | (translation rate), as calculated by the [[https://salislab.net/software Salis RBS Calculator]]. Due to this difference | ||
in construction we compared each bxb1 to it's own control (see above, [[#experimental_setup|Experimental Setup]]).</p> | in construction we compared each bxb1 to it's own control (see above, [[#experimental_setup|Experimental Setup]]).</p> | ||
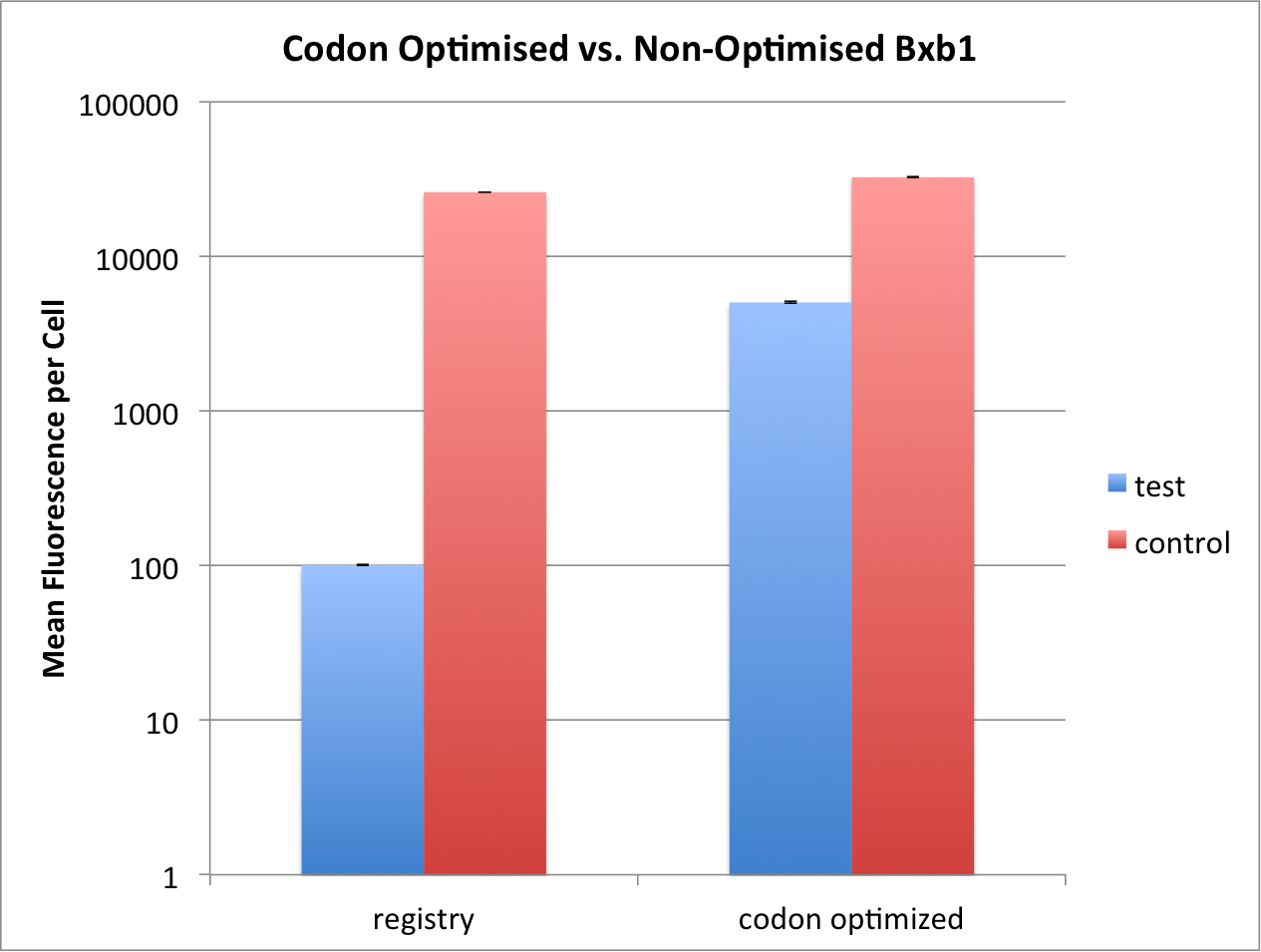
<p> We could show that the codon optimised bxb1 could reach ~1/6th flipping compared to control, whereas the non-codon optimised | <p> We could show that the codon optimised bxb1 could reach ~1/6th flipping compared to control, whereas the non-codon optimised | ||
| − | one reached only ~1/260th of its own control. We thus argue that the codon | + | one reached only ~1/260th of its own control. We thus argue that the codon optimized bxb1 is more efficient, especially |
| − | since the fold increase in efficiency is a lot more than the fold increase in RBS | + | since the fold increase in efficiency is a lot more than the fold increase in RBS strength. </p> |
[[File:T--ETH Zurich--codonoptimisation.png|500px|thumb|center|Comparison of codon optimised and non-optimised bxb1 flipping | [[File:T--ETH Zurich--codonoptimisation.png|500px|thumb|center|Comparison of codon optimised and non-optimised bxb1 flipping | ||
efficiency. Test constructs have bxb1 expressed under the Tet promoter, with TetR present in the system. Control doesn't | efficiency. Test constructs have bxb1 expressed under the Tet promoter, with TetR present in the system. Control doesn't | ||
have TetR in the system, and thus constitutively expresses bxb1. While non-codon optimised bxb1 can only reach ~1/260th | have TetR in the system, and thus constitutively expresses bxb1. While non-codon optimised bxb1 can only reach ~1/260th | ||
| − | of the full flipping reached by control, the codon optimised one reaches ~1/6th of its own control. Induction at OD<sub>600</sub> 0.5 with 2000ng/ | + | of the full flipping reached by control, the codon optimised one reaches ~1/6th of its own control. Induction at OD<sub>600</sub> 0.5 with 2000ng/mL. Data measured by flow cytometry at 16h. Values are shown as mean fluorescence per cell, error bars |
indicate SEM. ]] | indicate SEM. ]] | ||
Revision as of 11:13, 24 October 2016
Mycobacterium Phage Bxb1 gp35, DNA integrase
- Group: ETH Zurich 2016
- Author: Asli Azizoglu
- Summary: We cloned and characterised a codon optimised bxb1 for E. coli, and sent it to the registry as a biobrick. Our biobrick can be found with a fast degradation tag BBa_K2116009 and here without any degradation tag Part:BBa_K2116026. The non-codon optimised version can be found here Part:BBa_K907000. Besides some favorable features, the codon optimized bxb1 also supports full assembly compatibility.
Kinetic Characterisation
We investigated the kinetics of bxb1 flipping using flow cytometry.
Genetic Design
- Bxb1 was expressed under a tet promoter, without any degradation tag Part:BBa_K2116056. The RBS has a strength of 1209.69 au (translation rate), as calculated by the RBS Calculator. This construct was cloned on a medium copy plasmid with the p15A replication of origin.
- TetR was expressed under the control of medium-strength constitutive promoter Part:Bba_J23118, and cloned onto a low copy plasmid with pSC101/Rep101 replication of origin.
- We also constructed a dual fluorescence reporter system Part:BBa_K2116024 that expresses sfGFP Part:BBa_K2116017 when bxb1 flips a directional promoter. In the non-flipped state the reporter expresses the red fluorescent protein mNectarine Part:BBa_K2116016.
Experimental Setup
- The test construct with bxb1 was transformed together with the reporter construct, and either with TetR or with the empty backbone as control.
- Cells were grown in LB medium for 4 hours, and then transfered to minimal M9 medium with a 1:100 dilution and grown over-night. The culture was again diluted 1:100 in minimal M9 medium the next day. aTc was added at given concentrations at OD600. Samples were taken at the specified time points, spun down and resuspended in PBS at OD600 0.01 and kept on ice until measurement.
- Results are presented as mean fluorescence per cell and the standard error of the mean.
Results
We observed that maximum flipping is reached at around 8h under these conditions. The response correlates with aTc concentration. However, compared to the control, where there is no TetR, we see 6-fold less flipping. This could be improved by increasing the concentration of aTc, however we have found that at 8000 ng/mL cell growth was inhibited.
Comparison between Codon Optimised and Registry Bxb1 Integrase
We compared the kinetics of our codon optimised bxb1 Part:BBa_K2116026 and the one found previously on the registry Part:BBa_K907000 that was not codon optimized. The same genetic and experimental setup was used as above. The only difference between the two constructs is that the non-codon optimised bxb1 has a weaker RBS, with a strength of 251.82au (translation rate), as calculated by the [Salis RBS Calculator]. Due to this difference in construction we compared each bxb1 to it's own control (see above, Experimental Setup).
We could show that the codon optimised bxb1 could reach ~1/6th flipping compared to control, whereas the non-codon optimised one reached only ~1/260th of its own control. We thus argue that the codon optimized bxb1 is more efficient, especially since the fold increase in efficiency is a lot more than the fold increase in RBS strength.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 192
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 466
Illegal XhoI site found at 553 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 1105
Illegal NgoMIV site found at 1192
Illegal AgeI site found at 242 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI.rc site found at 1300

