Difference between revisions of "Part:BBa R0071:Experience"
| Line 17: | Line 17: | ||
[[Image:prhl4.png|thumb|center|400px|Fig. 3 RFU of GFP / turbidity of imoroved Prhl mutants]]<br> | [[Image:prhl4.png|thumb|center|400px|Fig. 3 RFU of GFP / turbidity of imoroved Prhl mutants]]<br> | ||
| − | The sequences of these | + | The sequences of these two chosen mutants are shown in below. |
| − | + | Native Prhl (<partinfo>BBa_R0071</partinfo>) | |
TCCTGTGAAATCTGGCAGTTACCGTTAGCTTTCGAATTGGCTAAAAAGTGTTCtactagagAAAGAGGAGAAA | TCCTGTGAAATCTGGCAGTTACCGTTAGCTTTCGAATTGGCTAAAAAGTGTTCtactagagAAAGAGGAGAAA | ||
| − | Prhl( | + | Prhl(D6) (<partinfo>BBa_K1949060</partinfo>) |
| − | + | TCCTGTGAAATCTGGCAGTTACCGTTAGCTTTCGAATTGGCTA<b>t</b>AAAGTGTTCtactagagAAAGAGGAGAAA | |
| − | Prhl( | + | Prhl(H7) |
| − | + | TCCTGTGAAATCTGGCAGTTACCGTTAGCTTTCGAATTGGCTA<b>t</b>AAAGTGTTCtactag<b>ta</b>AAAGAGGAGAAA | |
| − | = | + | |
| − | + | =-==Materials and Methods==== | |
| + | |||
| + | ====Construction==== | ||
-Strain | -Strain | ||
| Line 40: | Line 42: | ||
-Plasmids | -Plasmids | ||
| − | I. Improved Prhl by | + | ======I. Improved Prhl by Tokyo_Tech 2014 and rhlR assay===== |
| − | A. Pcon | + | A. Pcon-rhlR-ssrA(<partinfo>BBa_C0071</partinfo>) (pSB6A1), Prhl(LR)(<partinfo>BBa_K1529310</partinfo>)-gfp (pSB3K3) |
| − | B. Pcon-rhlR(BBa_C0171) (pSB6A1), Prhl(LR)-gfp (pSB3K3) | + | B. Pcon-rhlR(<partinfo>BBa_C0171</partinfo>) (pSB6A1), Prhl(LR)-gfp (pSB3K3) |
| − | C. Pcon-rhlR- | + | C. Pcon-rhlR-ssrA (pSB6A1), Prhl(RL)(<partinfo>BBa_K1529300</partinfo>)-gfp (pSB3K3) |
D. Pcon-rhlR (pSB6A1), Prhl(RL)-gfp (pSB3K3) | D. Pcon-rhlR (pSB6A1), Prhl(RL)-gfp (pSB3K3) | ||
| − | E. Pcon-rhlR- | + | E. Pcon-rhlR-ssrA (pSB6A1), Pcon-gfp (pSB3K3) …Positive control |
| − | F. pSB6A1, pSB3K3 …Nagative control | + | F. pSB6A1, pSB3K3 …Nagative control |
| − | II. Improvment the | + | =====II. Improvment the native Prhl===== |
| − | A. Prhl(BBa_R0071)-gfp (pSB3K3) | + | A. Prhl(<partinfo>BBa_R0071</partinfo>)-gfp (pSB3K3) |
| − | B. Pcon-rhlR- | + | B. Pcon-rhlR-ssrA (pSB6A1), Prhl(D6)(<partinfo>BBa_K1949060</partinfo> )-gfp (pSB3K3) |
| − | C. Pcon-rhlR- | + | C. Pcon-rhlR-ssrA (pSB6A1), Prhl(H7)-gfp (pSB3K3) |
| − | D. Pcon-rhlR- | + | D. Pcon-rhlR-ssrA (pSB6A1), Pcon-gfp (pSB3K3) …Positive control |
| − | E. pSB6A1, pSB3K3 …Nagative control | + | E. pSB6A1, pSB3K3 …Nagative control |
| − | III. Comparison the improved Prhl by | + | =====III. Comparison the improved Prhl by Tokyo_Tech 2014 to our original improved Prhl===== |
| − | A. Pcon-rhlR- | + | A. Pcon-rhlR-ssrA (pSB6A1), Prhl(LR)-gfp (pSB3K3) |
| − | B. Pcon-rhlR- | + | B. Pcon-rhlR-ssrA (pSB6A1), Prhl(D6)-gfp (pSB3K3) |
| − | C. Pcon-rhlR- | + | C. Pcon-rhlR-ssrA (pSB6A1), Pcon-gfp (pSB3K3)…Positive control |
D. pSB6A1, pSB3K3…Negative control | D. pSB6A1, pSB3K3…Negative control | ||
-Medium | -Medium | ||
| + | |||
LB medium AK | LB medium AK | ||
| Line 86: | Line 89: | ||
| − | + | ====Assay protocol==== | |
| + | |||
The following experiments is performed at 37℃ unless otherwise stated. | The following experiments is performed at 37℃ unless otherwise stated. | ||
| − | I. Improved Prhl by | + | =====I. Improved Prhl by Tokyo_Tech 2014 and rhlR assay===== |
1) Prepare overnight culture for each sample in 3mL LB medium AK with vigorous shaking. | 1) Prepare overnight culture for each sample in 3mL LB medium AK with vigorous shaking. | ||
| Line 103: | Line 107: | ||
6) Add 100 microL of the samples into each well of a plate reader. | 6) Add 100 microL of the samples into each well of a plate reader. | ||
| − | 7) Measure the | + | 7) Measure the fluorescence intensity of GFP at 490 nm as an exciting wavelength, 525 nm as a measurement wavelength. |
| − | 8) Measure the | + | 8) Measure the turbidity at 600 nm. |
| − | II. Improvement the | + | =====II. Improvement the native Prhl (<partinfo>BBa_R0071</partinfo>)===== |
| − | 1) Introduce a point mutation to -10 region of Prhl (BBa_R0071) | + | 1) Introduce a point mutation to -10 region of Prhl (BBa_R0071)-gfp (pSB3K3) by inverse PCR using native Prhl(BBa_R0071)-gfp (pSB3K3) as a template and divergent primers. The sequences of the six forward premiers and of one reverse primer are shown below. |
| − | + | ||
| − | The sequences of the six forward premiers and of one reverse primer are shown below. | + | |
WT 5'-tcctgtgaaatctggcagttaccgttagctttcgaatt ggctaaaaagtgttctactagtagcg-3' | WT 5'-tcctgtgaaatctggcagttaccgttagctttcgaatt ggctaaaaagtgttctactagtagcg-3' | ||
| Line 130: | Line 132: | ||
R 5'-aattcgaaagctaacggtaactgcca-3' (5’ phosphorylated) | R 5'-aattcgaaagctaacggtaactgcca-3' (5’ phosphorylated) | ||
| − | 2) Prepare strains containing the pMu1~6 plasmids containing mutated Prhl- | + | 2) Prepare strains containing the pMu1~6 plasmids containing mutated Prhl-GFP (pSB3K3). |
3) Streak the transformants on an agar plate containing kanamycin (50 microg/ mL) and incubate overnight. | 3) Streak the transformants on an agar plate containing kanamycin (50 microg/ mL) and incubate overnight. | ||
| Line 136: | Line 138: | ||
4) Pore the LB medium K at the extent of covering the surface of the agar, and collect the colonies with a spreader. Then, incubate them in 5 mL LB medium K for 16 h with vigorous shaking. | 4) Pore the LB medium K at the extent of covering the surface of the agar, and collect the colonies with a spreader. Then, incubate them in 5 mL LB medium K for 16 h with vigorous shaking. | ||
| − | 5) Extract the plasmid, and introduce Prhl(mut)-gfp (pSB3K3) and Pcon-rhlR- | + | 5) Extract the plasmid, and introduce Prhl(mut)-gfp (pSB3K3) and Pcon-rhlR-ssrA to XL-1 Blue. |
6) Pore 200 microL LB medium AK containing C4 (6 microM) in one well of a 96-well plate and inoculate one colony to the well. Incubate the plate for 16 h. | 6) Pore 200 microL LB medium AK containing C4 (6 microM) in one well of a 96-well plate and inoculate one colony to the well. Incubate the plate for 16 h. | ||
| − | 7) Transfer 100 microL of the culture from the above samples to another plate that is suitable for | + | 7) Transfer 100 microL of the culture from the above samples to another plate that is suitable for fluorescence measurement. Measure the GFP fluorescence intensity at 490 nm as an exciting wavelength and 525 nm as an emission wavelength as well as the turbidity at 600 nm. |
| − | 8) Select mutants that have higher SN ratio than | + | 8) Select mutants that have higher SN ratio than native Prhl (BBa_R0071) and grow them in 3mL LB medium AK overnight with vigorous shaking. |
9) Dilute the overnight cultures to 1 / 60 in fresh LB medium AK (1.2 mL). | 9) Dilute the overnight cultures to 1 / 60 in fresh LB medium AK (1.2 mL). | ||
| Line 148: | Line 150: | ||
10) Incubate the fresh cultures for 1 h with vigorous shaking. | 10) Incubate the fresh cultures for 1 h with vigorous shaking. | ||
| − | 11) Add C4, C12 or DMSO | + | 11) Add C4, C12 or DMSO to each 500 microL sample at the final concentration 10 microM. |
12) Incubate the samples for 4 h with vigorous shaking. | 12) Incubate the samples for 4 h with vigorous shaking. | ||
| Line 154: | Line 156: | ||
13) Add 100 microL of the samples into each well of a plate reader. | 13) Add 100 microL of the samples into each well of a plate reader. | ||
| − | 14) Measure the | + | 14) Measure the fluorescence intensity of GFP at 490 nm as an exciting wavelength, 525 nm as a measurement wavelength. |
| − | 15) Measure the | + | 15) Measure the turbidity at 600 nm. |
| − | III. Comparison the improved Prhl promoter by | + | =====III. Comparison the improved Prhl promoter by Tokyo_Tech 2014 to our original improved Prhl promoter===== |
1) Prepare overnight cultures for each sample in 3mL LB medium AK with vigorous shaking. | 1) Prepare overnight cultures for each sample in 3mL LB medium AK with vigorous shaking. | ||
| Line 168: | Line 170: | ||
3) Incubate the fresh cultures for 1 h with vigorous shaking. | 3) Incubate the fresh cultures for 1 h with vigorous shaking. | ||
| − | 4) Add C4, C12 or DMSO | + | 4) Add C4, C12 or DMSO to each 500 microL sample at the final concentration 10 microM. |
5) Incubate the samples for 4 h with vigorous shaking. | 5) Incubate the samples for 4 h with vigorous shaking. | ||
| Line 174: | Line 176: | ||
6) Add 100 microL of the samples into each well of a plate reader. | 6) Add 100 microL of the samples into each well of a plate reader. | ||
| − | 7) Measure the | + | 7) Measure the fluorescence intensity of GFP at 490 nm as an exciting wavelength, 525 nm as a measurement wavelength. |
| + | |||
| + | 8) Measure the turbidity at 600 nm. | ||
| − | + | 6. Reference | |
| − | |||
(1) Gerardo Medina et al. (2003) Mechanism of Pseudomonas aeruginosa RhlR Transcriptional Regulation of the rhlAB Promoter. | (1) Gerardo Medina et al. (2003) Mechanism of Pseudomonas aeruginosa RhlR Transcriptional Regulation of the rhlAB Promoter. | ||
BACTERIOLOGY 185: 5976-5983 | BACTERIOLOGY 185: 5976-5983 | ||
Revision as of 03:34, 16 October 2016
Improvement and Characterize
Group: Tokyo Tech 2016
Author: Yoshio Takata
Summary of Improvement and Characterization:
I. Improved Prhl by iGEM 2014 Tokyo_Tech and characterize RhlR assay
II. Improvement of the wild type Prhl
III. Comparison of the improved Prhl by iGEM 2014 Tokyo_Tech to our original improved Prhl.
The sequences of these two chosen mutants are shown in below.
Native Prhl (BBa_R0071)
TCCTGTGAAATCTGGCAGTTACCGTTAGCTTTCGAATTGGCTAAAAAGTGTTCtactagagAAAGAGGAGAAA
Prhl(D6) (BBa_K1949060)
TCCTGTGAAATCTGGCAGTTACCGTTAGCTTTCGAATTGGCTAtAAAGTGTTCtactagagAAAGAGGAGAAA
Prhl(H7)
TCCTGTGAAATCTGGCAGTTACCGTTAGCTTTCGAATTGGCTAtAAAGTGTTCtactagtaAAAGAGGAGAAA
-==Materials and Methods===
Construction
-Strain
All the plasmids were prepared in XL1-Blue strain.
-Plasmids
=I. Improved Prhl by Tokyo_Tech 2014 and rhlR assay
A. Pcon-rhlR-ssrA(BBa_C0071) (pSB6A1), Prhl(LR)(BBa_K1529310)-gfp (pSB3K3)
B. Pcon-rhlR(BBa_C0171) (pSB6A1), Prhl(LR)-gfp (pSB3K3)
C. Pcon-rhlR-ssrA (pSB6A1), Prhl(RL)(BBa_K1529300)-gfp (pSB3K3)
D. Pcon-rhlR (pSB6A1), Prhl(RL)-gfp (pSB3K3)
E. Pcon-rhlR-ssrA (pSB6A1), Pcon-gfp (pSB3K3) …Positive control
F. pSB6A1, pSB3K3 …Nagative control
II. Improvment the native Prhl
A. Prhl(BBa_R0071)-gfp (pSB3K3)
B. Pcon-rhlR-ssrA (pSB6A1), Prhl(D6)(BBa_K1949060 )-gfp (pSB3K3)
C. Pcon-rhlR-ssrA (pSB6A1), Prhl(H7)-gfp (pSB3K3)
D. Pcon-rhlR-ssrA (pSB6A1), Pcon-gfp (pSB3K3) …Positive control
E. pSB6A1, pSB3K3 …Nagative control
III. Comparison the improved Prhl by Tokyo_Tech 2014 to our original improved Prhl
A. Pcon-rhlR-ssrA (pSB6A1), Prhl(LR)-gfp (pSB3K3)
B. Pcon-rhlR-ssrA (pSB6A1), Prhl(D6)-gfp (pSB3K3)
C. Pcon-rhlR-ssrA (pSB6A1), Pcon-gfp (pSB3K3)…Positive control
D. pSB6A1, pSB3K3…Negative control
-Medium
LB medium AK
LB medium containing ampicillin (50 microg/ mL) and kanamycin (50 microg/ mL)
LB medium K
LB medium containing kanamycin (50 microg/ mL)
Assay protocol
The following experiments is performed at 37℃ unless otherwise stated.
I. Improved Prhl by Tokyo_Tech 2014 and rhlR assay
1) Prepare overnight culture for each sample in 3mL LB medium AK with vigorous shaking.
2) Dilute the overnight cultures to 1 / 60 in fresh LB medium AK (1.2 mL).
3) Incubate the fresh cultures for 1 h with vigorous shaking.
4) Add C4 or DMSO to each 500 microL sample at the final concentration 10 microM or 100 microM.
5) Incubate the samples for 4 h with vigorous shaking.
6) Add 100 microL of the samples into each well of a plate reader.
7) Measure the fluorescence intensity of GFP at 490 nm as an exciting wavelength, 525 nm as a measurement wavelength.
8) Measure the turbidity at 600 nm.
II. Improvement the native Prhl (BBa_R0071)
1) Introduce a point mutation to -10 region of Prhl (BBa_R0071)-gfp (pSB3K3) by inverse PCR using native Prhl(BBa_R0071)-gfp (pSB3K3) as a template and divergent primers. The sequences of the six forward premiers and of one reverse primer are shown below.
WT 5'-tcctgtgaaatctggcagttaccgttagctttcgaatt ggctaaaaagtgttctactagtagcg-3'
F-Mu1 5'-tcctgtgaaatctggcagttaccgttagctttcgaatt ggntaaaaagtgttctactagtagcg-3'
F-Mu2 5'-tcctgtgaaatctggcagttaccgttagctttcgaatt ggcnaaaaagtgttctactagtagcg-3'
F-Mu3 5'-tcctgtgaaatctggcagttaccgttagctttcgaatt ggctnaaaagtgttctactagtagcg-3'
F-Mu4 5'-tcctgtgaaatctggcagttaccgttagctttcgaatt ggctanaaagtgttctactagtagcg-3'
F-Mu5 5'-tcctgtgaaatctggcagttaccgttagctttcgaatt ggctaanaagtgttctactagtagcg-3'
F-Mu6 5'-tcctgtgaaatctggcagttaccgttagctttcgaatt ggctaaanagtgttctactagtagcg-3'
R 5'-aattcgaaagctaacggtaactgcca-3' (5’ phosphorylated)
2) Prepare strains containing the pMu1~6 plasmids containing mutated Prhl-GFP (pSB3K3).
3) Streak the transformants on an agar plate containing kanamycin (50 microg/ mL) and incubate overnight.
4) Pore the LB medium K at the extent of covering the surface of the agar, and collect the colonies with a spreader. Then, incubate them in 5 mL LB medium K for 16 h with vigorous shaking.
5) Extract the plasmid, and introduce Prhl(mut)-gfp (pSB3K3) and Pcon-rhlR-ssrA to XL-1 Blue.
6) Pore 200 microL LB medium AK containing C4 (6 microM) in one well of a 96-well plate and inoculate one colony to the well. Incubate the plate for 16 h.
7) Transfer 100 microL of the culture from the above samples to another plate that is suitable for fluorescence measurement. Measure the GFP fluorescence intensity at 490 nm as an exciting wavelength and 525 nm as an emission wavelength as well as the turbidity at 600 nm.
8) Select mutants that have higher SN ratio than native Prhl (BBa_R0071) and grow them in 3mL LB medium AK overnight with vigorous shaking.
9) Dilute the overnight cultures to 1 / 60 in fresh LB medium AK (1.2 mL).
10) Incubate the fresh cultures for 1 h with vigorous shaking.
11) Add C4, C12 or DMSO to each 500 microL sample at the final concentration 10 microM.
12) Incubate the samples for 4 h with vigorous shaking.
13) Add 100 microL of the samples into each well of a plate reader.
14) Measure the fluorescence intensity of GFP at 490 nm as an exciting wavelength, 525 nm as a measurement wavelength.
15) Measure the turbidity at 600 nm.
III. Comparison the improved Prhl promoter by Tokyo_Tech 2014 to our original improved Prhl promoter
1) Prepare overnight cultures for each sample in 3mL LB medium AK with vigorous shaking.
2) Dilute the overnight cultures to 1 / 60 in fresh LB medium AK (1.2 mL).
3) Incubate the fresh cultures for 1 h with vigorous shaking.
4) Add C4, C12 or DMSO to each 500 microL sample at the final concentration 10 microM.
5) Incubate the samples for 4 h with vigorous shaking.
6) Add 100 microL of the samples into each well of a plate reader.
7) Measure the fluorescence intensity of GFP at 490 nm as an exciting wavelength, 525 nm as a measurement wavelength.
8) Measure the turbidity at 600 nm.
6. Reference
(1) Gerardo Medina et al. (2003) Mechanism of Pseudomonas aeruginosa RhlR Transcriptional Regulation of the rhlAB Promoter. BACTERIOLOGY 185: 5976-5983
(2) John S. Chuang et al. (2009) Simpson’s Paradox in a Synthetic Microbial System. SCIENCE 323: 272-275
Improvement
iGEM 2014 Tokyo_Tech
We, Tokyo Tech 2014, improved Prhl Promoter (BBa_R0071) by changing LuxR binding site of Plux Promoter to RhlR binding site.
We dsigned new Prhl promoter
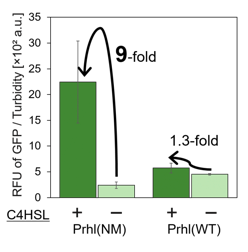
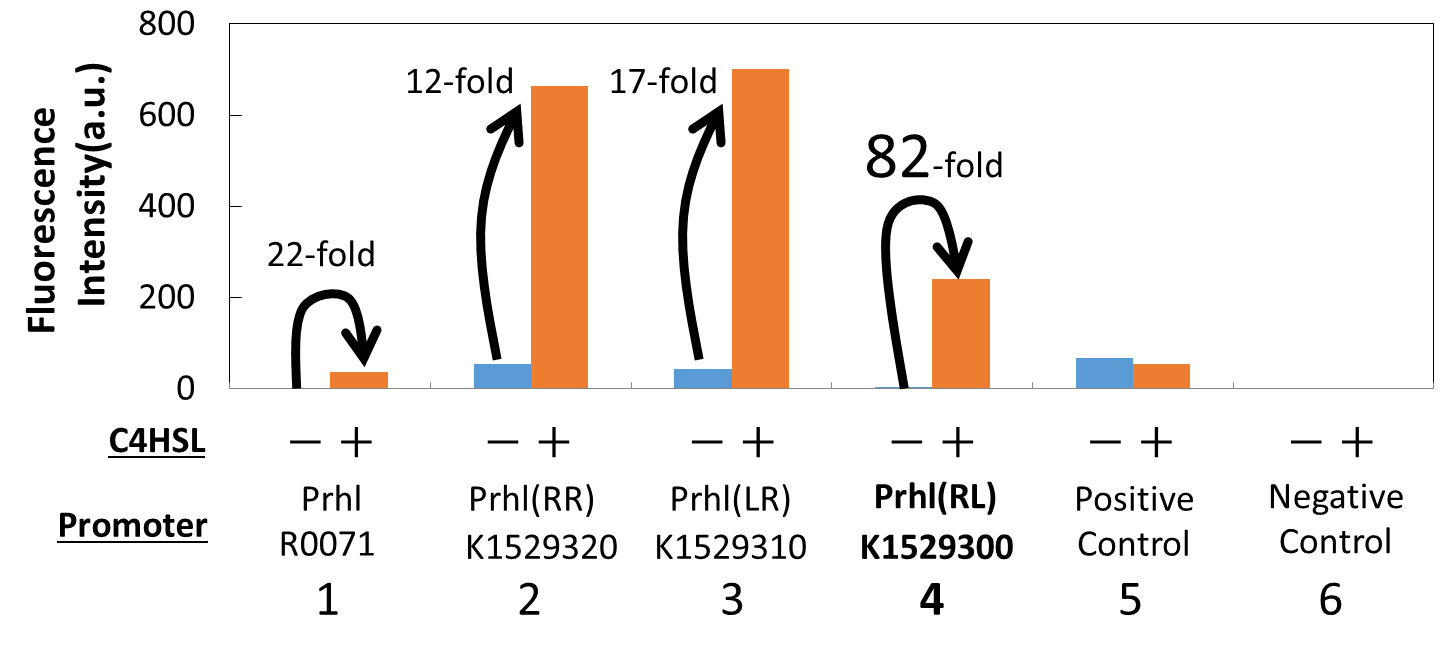
We measured the GFP expression with the four different promoters (Prhl : BBa_R0071, Prhl(RR) : BBa_K1529320, Prhl(LR) : BBa_K1529310 and Prhl(RL) : BBa_K1529300) by flow cytometer. Each promoter was tested in the presence and also in the absence of C4HSL.
Fig. 1 shows the fluorescence intensity detected by flow cytometer.
From the result, when induced by C4HSL, Prhl(RR) promoter showed higher maximum expression level and higher leak than the original Prhl promoter (BBa_R0071).
Although Prhl(RL) promoter had lower maximum expression level compared to Prhl(RR) promoter, it had the highest induced/not-induced ratio.
This means Prhl(RL) promoter has little leak.
Therefore, we can say that Prhl(RL):BBa_K1529300 promoter is the best improved Prhl promoter due to the advantages of less leak and higher expression level.
For more information, see [http://2014.igem.org/Team:Tokyo_Tech/Experiment/Prhl_reporter_assay our work in Tokyo_Tech 2014 wiki].
Applications of BBa_R0071
User Reviews
UNIQb0c8f070388e572d-partinfo-0000000E-QINU UNIQb0c8f070388e572d-partinfo-0000000F-QINU
|
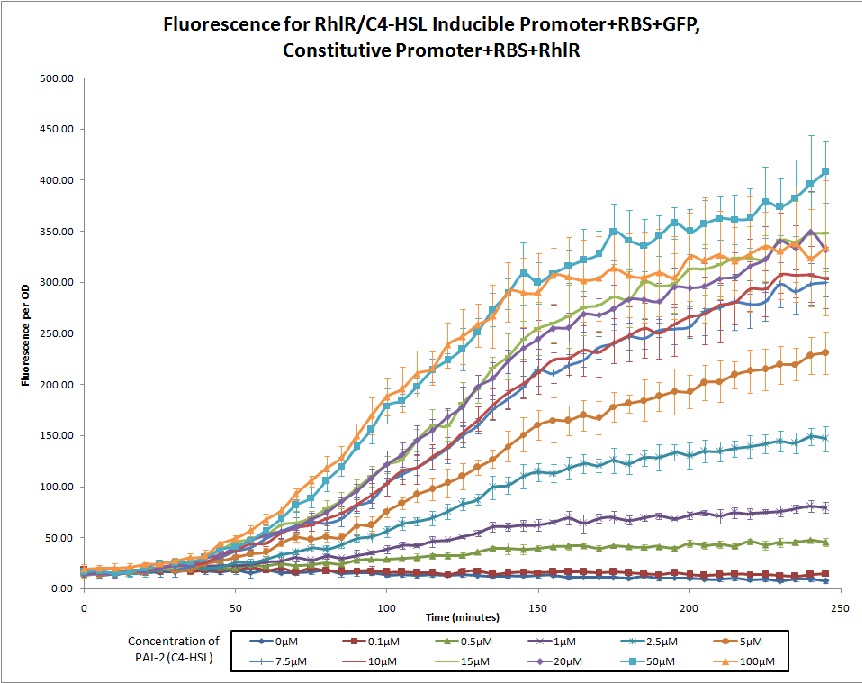
No review score entered. Northwestern 2011 |
The 2011 Northwestern iGEM team used this promoter as a unit within our Pseudomonas Aeruginosa biosensor. When this RhlR/C4-HSL regulated promoter is induced at varying concentrations of C4-HSL, we observed GFP fluorescence in accordance to the graph below. |