Difference between revisions of "Part:BBa K1758340"
| Line 95: | Line 95: | ||
<p><i>In vitro</i> this sensor showed good results. The fluorescence level was high at low concentrations. Additionally, it showed that the expression level at 6 µg/L (Guideline of WHO for Mercury) reached the maximal signal. This result indicated the potential for measurement of concentrations under 6 µg/L.To confirm this hypothesis, it takes more experiments and tests with lower concentrations. Due to the high expression of sfGFP at low concentrations and the same expression level at different concentrations, it is not possible to quantify mercury with CFPS analyses . , Our model predicted this observation. During the measurement we noticed that the heavy metals have negative influences on the cell extract. Because of this fact, we used a correction factor, which resulted from the heavy metals influence on the CFPS system. This already optimized sensor showed the high potential of optimized sensors in CFPS.</p> | <p><i>In vitro</i> this sensor showed good results. The fluorescence level was high at low concentrations. Additionally, it showed that the expression level at 6 µg/L (Guideline of WHO for Mercury) reached the maximal signal. This result indicated the potential for measurement of concentrations under 6 µg/L.To confirm this hypothesis, it takes more experiments and tests with lower concentrations. Due to the high expression of sfGFP at low concentrations and the same expression level at different concentrations, it is not possible to quantify mercury with CFPS analyses . , Our model predicted this observation. During the measurement we noticed that the heavy metals have negative influences on the cell extract. Because of this fact, we used a correction factor, which resulted from the heavy metals influence on the CFPS system. This already optimized sensor showed the high potential of optimized sensors in CFPS.</p> | ||
| + | |||
| + | <b>References</b> | ||
| + | <p>iGEM Team Peking 2010</p> | ||
| + | |||
</html> | </html> | ||
Revision as of 19:35, 20 September 2015
Mercury repressor under control of constitutive promoter and strong RBS
Repressor of the mercurry responsive promoter PmerT under the control of konstitutive Promoter (K608002)
Usage and Biology
This device was used to create cell extract for our in vitro characterization of the mercury biosensor. It is based on BBa_K346001 desinged by team Peking 2010. Together with BBa_K1758342,BBa_K1758343 it represents our mercury sensor.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 462
Illegal NheI site found at 485 - 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Results
One of the already existing sensors we used for our system is the mercury sensor consisting of MerR the activator and the mercury specific promoter pmerT. The promoter is regulated by the MerR, which binds Hg2+-ions. Similar to the former sensors we added a sfGFP for detection via fluorescence.
For our mercury sensor we used parts of the mercury sensor constructed by iGEM team Peking 2010. These parts consist of the mercury dependent mer operon from Shigella flexneri R100 plasmid Tn21. The expression of the genes in the mer operon depends on the regulation by MerR its activator and promoter PmerT. For our sensor we used the codon optimized activator (BBa_K1758340), under control of a constitutive promoter,(BBa_K346001). Additionally to this activator we designed and constructed the specific promoter PmerT(BBa_K346002)(figure 2). For our sensor we added a 5’-UTR downstreamd of this promoter, which increased the fluorscence of the used reporter protein sfGFP.

We tested our mercury sensor with sfGFP as reporter gene, to test the functionality of the system. Moreover we tested different concentrations. The kinetic of our sensors response to different mercury concentrations is shown in figure 3. A strong increase in fluorescence levels is notecible after induction with mercury after 120 min. For better visualization the kinetics of figure 3 are represented as bars in figure 4. A fluorescence level difference for 120 min and 190 min is represented.
in vitro
For the characterization of the mercury sensor with CFPS we used parts differing from that we used in the in vivo characterization. For the in vitro characterization we used a cell extract out of cells, which contained the plasmid ( BBa_K1758340)(figure 5). In addition, we added plasmid DNA to the cell extract. This plasmid consisted of the mercury specific promoter pmerT with 5’-UTR-sfGFP. The entire sequence was placed under the control of of T7-promoter ( BBa_K1758344)(figure 6). The T7-promoter is needed to get a better fluorescence expression.




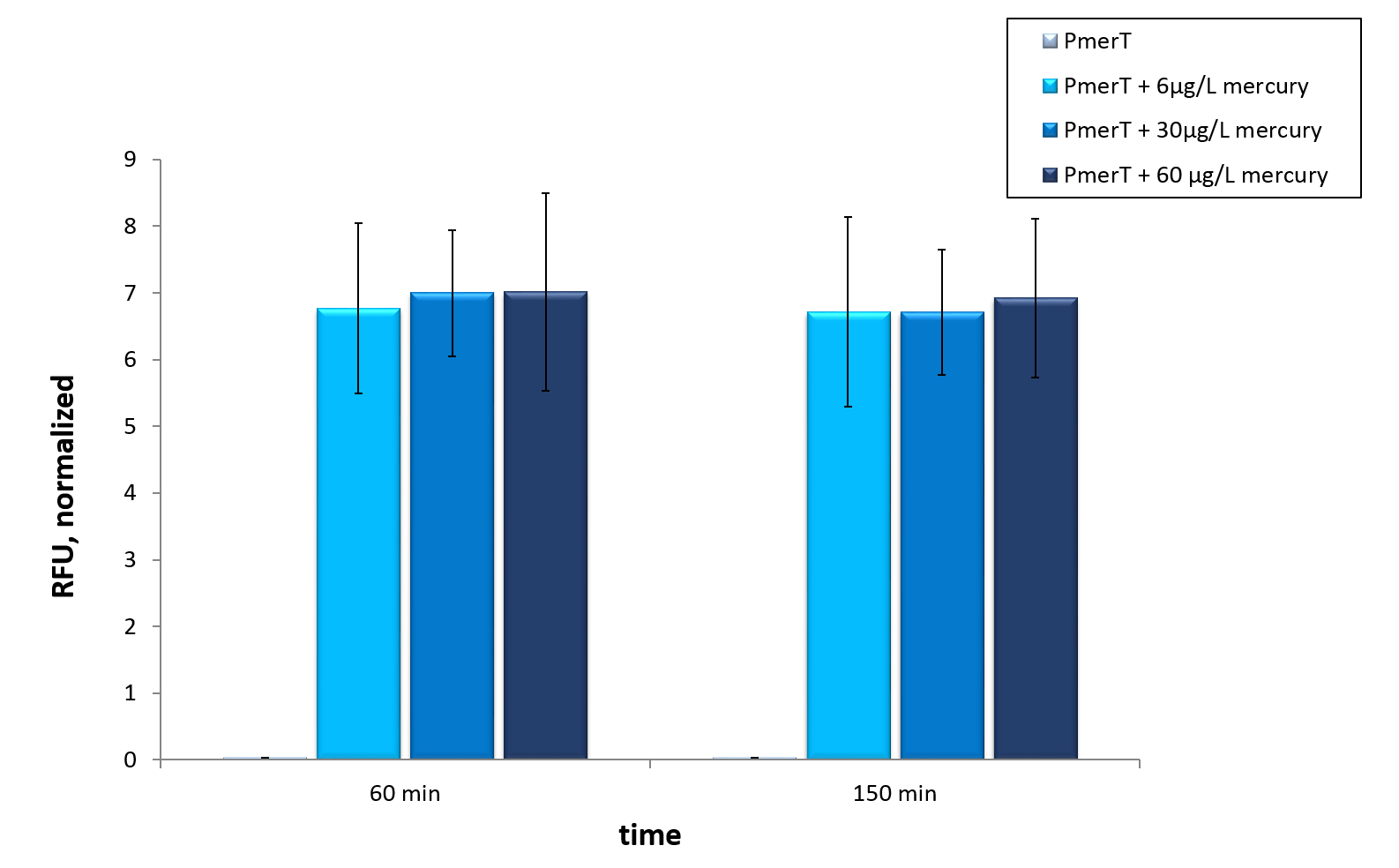
In vitro this sensor showed good results. The fluorescence level was high at low concentrations. Additionally, it showed that the expression level at 6 µg/L (Guideline of WHO for Mercury) reached the maximal signal. This result indicated the potential for measurement of concentrations under 6 µg/L.To confirm this hypothesis, it takes more experiments and tests with lower concentrations. Due to the high expression of sfGFP at low concentrations and the same expression level at different concentrations, it is not possible to quantify mercury with CFPS analyses . , Our model predicted this observation. During the measurement we noticed that the heavy metals have negative influences on the cell extract. Because of this fact, we used a correction factor, which resulted from the heavy metals influence on the CFPS system. This already optimized sensor showed the high potential of optimized sensors in CFPS.
ReferencesiGEM Team Peking 2010


