Difference between revisions of "Part:BBa K1758320"
(→Results) |
|||
| Line 23: | Line 23: | ||
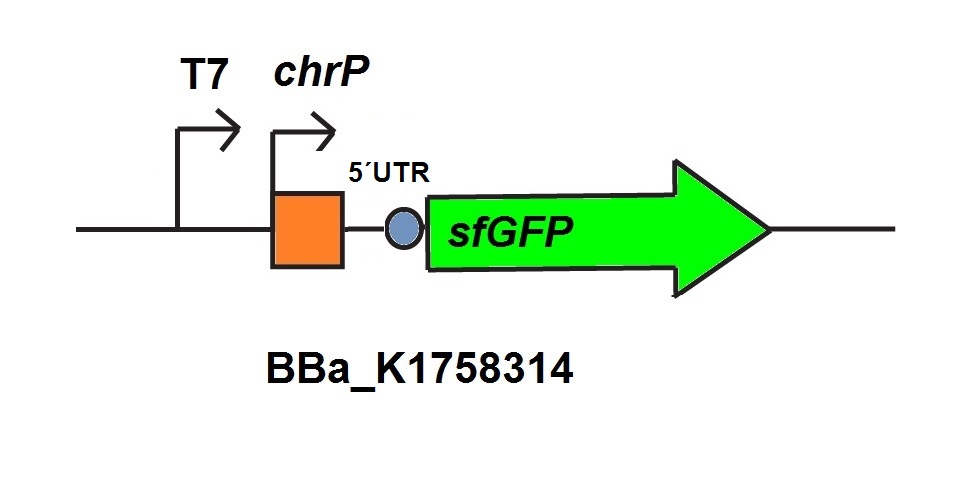
<p>For the characterization of the chromium sensor with CFPS we used parts differing from that we used in vivo characterization. For the in vitro characterization we used a cell extract produced from cells which contain the plasmid (<a href="https://parts.igem.org/Part:BBa_K1758310" target="_blank">BBa_K1758310</a>). The plasmid contains the gene <i>chrB</i> under the control of a constitutive promoter, so that the cell extract is enriched with repressor molecules. In addition to that we added plasmid-DNA of the chromium specific promoter <i>chrP</i> with 5’UTR-sfGFP under the control of T7-promoter (<a href="https://parts.igem.org/Part:BBa_K1758314" target="_blank">BBa_K1758314</a> (figure 6))to the cell extract. The T7-promoter is needed to get a better fluorescence expression. </p> | <p>For the characterization of the chromium sensor with CFPS we used parts differing from that we used in vivo characterization. For the in vitro characterization we used a cell extract produced from cells which contain the plasmid (<a href="https://parts.igem.org/Part:BBa_K1758310" target="_blank">BBa_K1758310</a>). The plasmid contains the gene <i>chrB</i> under the control of a constitutive promoter, so that the cell extract is enriched with repressor molecules. In addition to that we added plasmid-DNA of the chromium specific promoter <i>chrP</i> with 5’UTR-sfGFP under the control of T7-promoter (<a href="https://parts.igem.org/Part:BBa_K1758314" target="_blank">BBa_K1758314</a> (figure 6))to the cell extract. The T7-promoter is needed to get a better fluorescence expression. </p> | ||
| − | + | <figure> | |
| − | + | <img src=" https://static.igem.org/mediawiki/2015/e/e4/Bielefeld-CeBiTec_in_vitro_ChrB-part.jpeg" width="100 %"><figcaption> Figure 5: To produce the cell extract for <i>in vitro</i> characterization a construct (<a href="https://parts.igem.org/Part:BBa_K1758310" target="_blank">BBa_K1758310</a> ) with chromium repressor under the control of a constitutive promoter and strong RBS (BBa_K608002) is needed.</figcaption> | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
</figure> | </figure> | ||
| − | + | <figure> | |
| − | + | <img src="https://static.igem.org/mediawiki/2015/1/1f/Bielefeld-CebiTec_in_vitro_T7-chrP-UTR-sfGFP.jpeg" width="100%"> <figcaption> T7-chrP-UTR-sfGFP <a href="https://parts.igem.org/Part:BBa_K1758314" target="_blank">BBa_K1758314</a> used for<i>in vitro</i> characterization.</figcaption> | |
| − | + | ||
</figure> | </figure> | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | <figure | + | <figure> |
| − | + | <img src="https://static.igem.org/mediawiki/2015/9/99/Bielefeld-CeBiTec_Influence_of_chromium_on_the_cell_extract.jpeg" width="100%"> | |
<figcaption>Figure 7: Influence of different chromium concentrations on our crude cell extract. Error bars represent the standard deviation of three biological replicates. </figcaption> | <figcaption>Figure 7: Influence of different chromium concentrations on our crude cell extract. Error bars represent the standard deviation of three biological replicates. </figcaption> | ||
</figure> | </figure> | ||
| Line 46: | Line 37: | ||
<p>Chromium’s influence on the cell extract as shown in figure 7 is minimal for low concentrations. Higher chromium concentrations have a measurable impact on the viability of the cell extract, which is visible at concentrations of 120 µg/L and obvious at concentrations of 240 µg/L chromium.</p> | <p>Chromium’s influence on the cell extract as shown in figure 7 is minimal for low concentrations. Higher chromium concentrations have a measurable impact on the viability of the cell extract, which is visible at concentrations of 120 µg/L and obvious at concentrations of 240 µg/L chromium.</p> | ||
| − | + | <figure> | |
| − | + | <img src="https://static.igem.org/mediawiki/2015/f/fb/Bielefeld-CeBiTec_induction_chromium_in_chrB_cell_extract.jpg" width="100%"> | |
| − | + | ||
| − | + | ||
<figcaption>Figure 8: Chromium specific cell extract made from <i>E. coli</i> cells which already expressed the repressor before cell extract production. Induction with different chromium concentrations. Error bars represent the standard deviation of three biological replicates. </figcaption> | <figcaption>Figure 8: Chromium specific cell extract made from <i>E. coli</i> cells which already expressed the repressor before cell extract production. Induction with different chromium concentrations. Error bars represent the standard deviation of three biological replicates. </figcaption> | ||
</figure> | </figure> | ||
| Line 58: | Line 47: | ||
| − | + | <figure> | |
| − | + | <img src="https://static.igem.org/mediawiki/2015/1/1e/Bielefeld-CeBiTec_correction_induction_chromium_in_chrB-cell-extract.jpeg" width="100%"> | |
| − | + | ||
<figcaption>Figure 9: Chromium specific cell extract made from <i>E. coli</i> cells which already expressed the repressor before cell extract production. Induction with different chromium concentrations. The data are normalized on chromium’s influence to the cell extract. Error bars represent the standard deviation of three biological replicates. </figcaption> | <figcaption>Figure 9: Chromium specific cell extract made from <i>E. coli</i> cells which already expressed the repressor before cell extract production. Induction with different chromium concentrations. The data are normalized on chromium’s influence to the cell extract. Error bars represent the standard deviation of three biological replicates. </figcaption> | ||
</figure> | </figure> | ||
| − | |||
| − | |||
| − | |||
<p>Taking the influence of different chromium concentrations under consideration measured fluorescence can be normalized on chromium’s influence on the cell extract (figure 9). Normalized data suggest, that higher concentrations of chromium induce fluorescence in relevance to chromium’s influence on the cell extract. </p> | <p>Taking the influence of different chromium concentrations under consideration measured fluorescence can be normalized on chromium’s influence on the cell extract (figure 9). Normalized data suggest, that higher concentrations of chromium induce fluorescence in relevance to chromium’s influence on the cell extract. </p> | ||
| − | + | <figure> | |
| − | + | <img src="https://static.igem.org/mediawiki/2015/b/bd/Bielefeld-CeBiTec_induction_chromium_in_chrB_optimized_cell_extract2.jpg" width="100%"> | |
| − | + | ||
| − | + | ||
| − | <figure | + | |
| − | + | ||
<figcaption>Figure 10: Chromium sensor with alternative repressor build by team Dundee 2015, which has only the first 15 codons optimized in chromium specific cell extract under the induction with different chromium concentrations. Error bars represent the standard deviation of three biological replicates. </figcaption> | <figcaption>Figure 10: Chromium sensor with alternative repressor build by team Dundee 2015, which has only the first 15 codons optimized in chromium specific cell extract under the induction with different chromium concentrations. Error bars represent the standard deviation of three biological replicates. </figcaption> | ||
</figure> | </figure> | ||
| − | + | ||
| − | + | <figure> | |
| − | <figure | + | <img src="https://static.igem.org/mediawiki/2015/f/fe/Bielefeld-CeBiTec_Corr-induction-Cr-in-ChrBopt-CE.jpeg" width="100%"> |
| − | + | ||
<figcaption>Figure 11: Chromium sensor with alternative repressor build by team Dundee 2015, which has only the first 15 codons optimized in chromium specific cell extract under the induction with different chromium concentrations. Data are normalized on chromium’s influence to the specific cell extract Error bars represent the standard deviation of three biological replicates. </figcaption> | <figcaption>Figure 11: Chromium sensor with alternative repressor build by team Dundee 2015, which has only the first 15 codons optimized in chromium specific cell extract under the induction with different chromium concentrations. Data are normalized on chromium’s influence to the specific cell extract Error bars represent the standard deviation of three biological replicates. </figcaption> | ||
</figure> | </figure> | ||
| − | |||
| − | |||
<p>In addition to the measurements of our chromium sensor in CFPS we measured our chromium inducible promoter with the repressor of team Dundee (figure 10, 11), which works similar to ours. In contrast to our repressor only first 15 codons of their repressor are codon-optimized. Measurements with their repressor showed tendencies similar to our measured repressor. After normalization induction with higher chromium concentrations showed a detectable fluorescence response for both measured datasets. </p> | <p>In addition to the measurements of our chromium sensor in CFPS we measured our chromium inducible promoter with the repressor of team Dundee (figure 10, 11), which works similar to ours. In contrast to our repressor only first 15 codons of their repressor are codon-optimized. Measurements with their repressor showed tendencies similar to our measured repressor. After normalization induction with higher chromium concentrations showed a detectable fluorescence response for both measured datasets. </p> | ||
| − | |||
</html> | </html> | ||
Revision as of 05:52, 19 September 2015
copper activator under control constitutive promoter and strong RBS
Activator for copper induceble promoter copAP under the control of constitutive promoter (K608002)
Usage and Biology
cueR is a merR like regulator, which stimulates the transcription of copAP, a P-type ATPase pump (Outten et al. 2000). CopAP is the central component in obtaining copper homeostasis, it exports free copper from cytoplasm to the periplasm. This is enabled by copper induced activation of the operon transcription via CueR. The CueR-Cu+ is the DNA-binding transcriptional dual regulator which activates transcription (Yamamoto, Ishihama 2005).
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 7
Illegal NheI site found at 30 - 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Results
in vitro
For the characterization of the chromium sensor with CFPS we used parts differing from that we used in vivo characterization. For the in vitro characterization we used a cell extract produced from cells which contain the plasmid (BBa_K1758310). The plasmid contains the gene chrB under the control of a constitutive promoter, so that the cell extract is enriched with repressor molecules. In addition to that we added plasmid-DNA of the chromium specific promoter chrP with 5’UTR-sfGFP under the control of T7-promoter (BBa_K1758314 (figure 6))to the cell extract. The T7-promoter is needed to get a better fluorescence expression.


Chromium’s influence on the cell extract as shown in figure 7 is minimal for low concentrations. Higher chromium concentrations have a measurable impact on the viability of the cell extract, which is visible at concentrations of 120 µg/L and obvious at concentrations of 240 µg/L chromium.

The decrease of fluorescence for higher chromium concentrations in chromium specific cell extract is shown in figure 8. An increase of fluorescence at higher chromium concentrations would have been expected resulting out of the induction of the chromium sensor. A factor which should be considered is the influence of high chromium concentrations to the cell extract. The test for influence of chromium on the specific cell extract, illustrated in figure 7 showed that the influence of chromium at low concentrations is not significant. But the graphic shows that high concentrations of chromium induce fatal damages to the cell extract.

Taking the influence of different chromium concentrations under consideration measured fluorescence can be normalized on chromium’s influence on the cell extract (figure 9). Normalized data suggest, that higher concentrations of chromium induce fluorescence in relevance to chromium’s influence on the cell extract.


In addition to the measurements of our chromium sensor in CFPS we measured our chromium inducible promoter with the repressor of team Dundee (figure 10, 11), which works similar to ours. In contrast to our repressor only first 15 codons of their repressor are codon-optimized. Measurements with their repressor showed tendencies similar to our measured repressor. After normalization induction with higher chromium concentrations showed a detectable fluorescence response for both measured datasets.
