Difference between revisions of "Part:BBa J23118"
KalenClifton (Talk | contribs) |
|||
| Line 18: | Line 18: | ||
<partinfo>BBa_J23118 parameters</partinfo> | <partinfo>BBa_J23118 parameters</partinfo> | ||
<!-- --> | <!-- --> | ||
| + | |||
| + | ===Baltimore Biocrew 2019 Characterization=== | ||
| + | <strong>Goal</strong><br> | ||
| + | |||
| + | We, the Baltimore Biocrew, decided to characterize some of the Anderson promoters. These promoters are highly used by iGEM but the relative expression of these promoters have been routinely determined by measuring the fluorescence of a reporter protein. However, the function of a promoter is to start transcription of a gene so it may be more informative to measure the amount of RNA (instead of protein) produced by a reporter gene. Therefore, we decided to further characterize a selection of the Anderson promoters (J23100, J23101, J23103, J23105, J23118) by measuring RNA using Quantitative Polymerase Chain Reaction (qPCR). | ||
| + | |||
| + | <strong>Results</strong><br> | ||
| + | |||
| + | In our first trial of qPCR (8/03/19), we were able to measure the relative strengths for J23100, J23101, J23103, and J23105 which were 1.00, 0.00, 0.81, and 1.93, respectively. Since these strengths did not match the relative expression levels reported by iGEM2006_Berkeley, we repeated the qPCR (8/10/19) with the same cDNA. The strengths from this second trial were 1.00, 0.00, 0.37, and 0.20. We repeated it again and the relative strengths that we got on 10/12/19 for J23100, J23101, J23103, and J23103 were 1, 0, 2.91, and .32. Next, we made new cDNA by growing new liquid cultures, extracting RNA again, and repeating reverse transcription. From the new cDNA, we repeated the qPCR procedure two more times. The relative strengths for that we got on 9/28/19 for J23100, J23101, J23103, J23105, and J23118 were 1, 24.63, .36, 1.76, and .25. The relative strengths that we got on 10/12/19 for J23100, J23101, J23103, and J23105 were 1, 45.97, 3.20, and 1.26. In addition we measured promoter J23118 twice and got the strengths 1.13 and 1.32. | ||
| + | |||
| + | Here is the relative promoter strengths that we got from the qPCR. Baltimore BioCrew in blue compared to the 2006 Berkeley iGEM in orange. | ||
| + | [[File:Promoters_BaltimoreBiocrew2019.png|500px]] | ||
| + | |||
| + | To support our RNA measurements we also measured fluorescence of the liquid cultures we used to extract RNA. The cultures were grown overnight so we expected the bacteria to be at the stationary phase, but we measured OD to normalize any differences in growth. | ||
| + | |||
| + | |||
| + | <html> | ||
| + | |||
| + | <style> | ||
| + | table, th, td { | ||
| + | border: 1px solid black; | ||
| + | border-collapse: collapse; | ||
| + | } | ||
| + | th, td { | ||
| + | padding: 15px; | ||
| + | } | ||
| + | </style> | ||
| + | </head> | ||
| + | <body> | ||
| + | |||
| + | <table style="width:100%"> | ||
| + | <tr> | ||
| + | <th> Promoter </th> | ||
| + | <th> OD </th> | ||
| + | <th>fluorescence</th> | ||
| + | <th>fluorescence divided by OD</th> | ||
| + | <th>corrected relative expression</th> | ||
| + | <th>reported relative expression</th> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>BBa_J23100</td> | ||
| + | <td>0.876</td> | ||
| + | <td>250</td> | ||
| + | <td>285.38</td> | ||
| + | <td>1</td> | ||
| + | <td>1</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>BBa_J23101</td> | ||
| + | <td>0.674</td> | ||
| + | <td>255</td> | ||
| + | <td>378.33 </td> | ||
| + | <td>1.33</td> | ||
| + | <td>0.7</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>BBa_J23103</td> | ||
| + | <td>1.1</td> | ||
| + | <td>230</td> | ||
| + | <td>209.09 </td> | ||
| + | <td>0.73</td> | ||
| + | <td>0.01</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>BBa_J23105</td> | ||
| + | <td>1.08</td> | ||
| + | <td>215.74</td> | ||
| + | <td>209.09 </td> | ||
| + | <td>0.76</td> | ||
| + | <td>0.24</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>BBa_J23118</td> | ||
| + | <td>1.04</td> | ||
| + | <td>238</td> | ||
| + | <td>228.84 </td> | ||
| + | <td>0.80</td> | ||
| + | <td>0.56</td> | ||
| + | </tr> | ||
| + | </table> | ||
Revision as of 03:44, 20 October 2019
constitutive promoter family member
Variant RFP (au) J23112 1 J23103 17 J23113 21 J23109 106 J23117 162 J23114 256 J23115 387 J23116 396 J23105 623 J23110 844 J23107 908 J23106 1185 J23108 1303 J23118 1429 J23111 1487 J23101 1791 J23104 1831 J23102 2179 J23100 2547 |
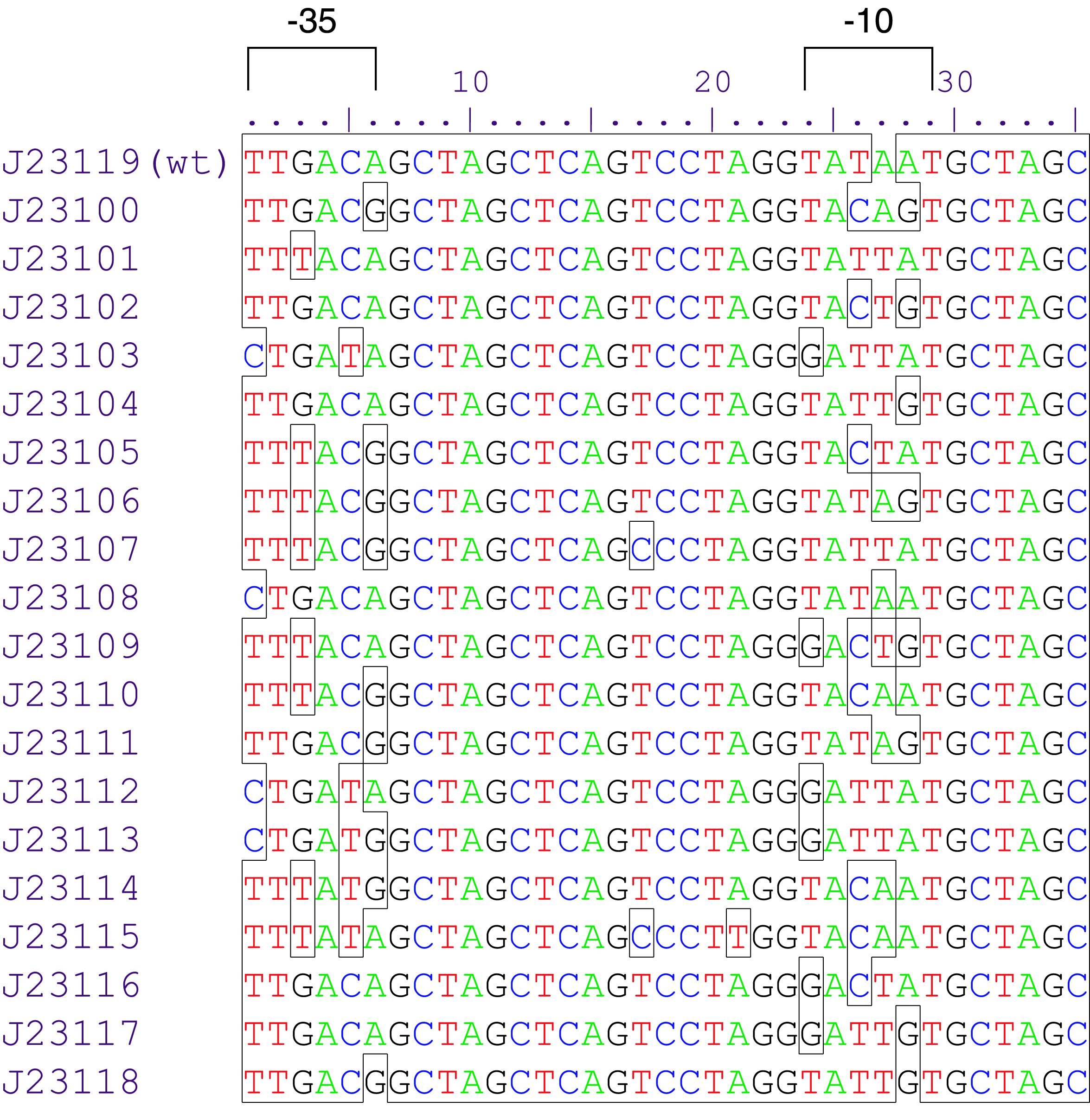
Constitutive promoter family
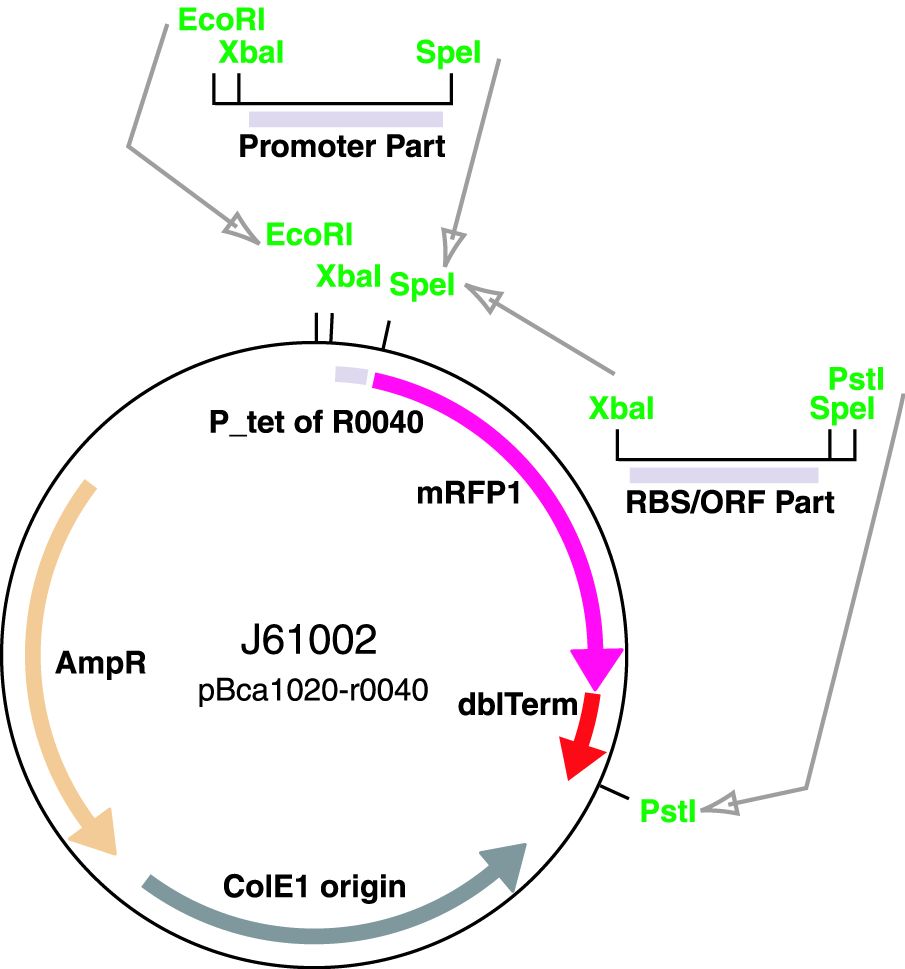
Parts J23100 through J23119 are a family of constitutive promoter parts isolated from a small combinatorial library. J23119 is the "consensus" promoter sequence and the strongest member of the family. All parts except J23119 are present in plasmid J61002. Part J23119 is present in pSB1A2. This places the RFP downstream of the promoter. Reported activities of the promoters are given as the relative fluorescence of these plasmids in strain TG1 grown in LB media to saturation. See part BBa_J61002 for details on their use.
These promoter parts can be used to tune the expression level of constitutively expressed parts. The NheI and AvrII restriction sites present within these promoter parts make them a scaffold for further modification. JCAraw
Saba: parts BBa_J23118 and BBa_J23110 are the same.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 7
Illegal NheI site found at 30 - 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000COMPATIBLE WITH RFC[1000]
Baltimore Biocrew 2019 Characterization
Goal
We, the Baltimore Biocrew, decided to characterize some of the Anderson promoters. These promoters are highly used by iGEM but the relative expression of these promoters have been routinely determined by measuring the fluorescence of a reporter protein. However, the function of a promoter is to start transcription of a gene so it may be more informative to measure the amount of RNA (instead of protein) produced by a reporter gene. Therefore, we decided to further characterize a selection of the Anderson promoters (J23100, J23101, J23103, J23105, J23118) by measuring RNA using Quantitative Polymerase Chain Reaction (qPCR).
Results
In our first trial of qPCR (8/03/19), we were able to measure the relative strengths for J23100, J23101, J23103, and J23105 which were 1.00, 0.00, 0.81, and 1.93, respectively. Since these strengths did not match the relative expression levels reported by iGEM2006_Berkeley, we repeated the qPCR (8/10/19) with the same cDNA. The strengths from this second trial were 1.00, 0.00, 0.37, and 0.20. We repeated it again and the relative strengths that we got on 10/12/19 for J23100, J23101, J23103, and J23103 were 1, 0, 2.91, and .32. Next, we made new cDNA by growing new liquid cultures, extracting RNA again, and repeating reverse transcription. From the new cDNA, we repeated the qPCR procedure two more times. The relative strengths for that we got on 9/28/19 for J23100, J23101, J23103, J23105, and J23118 were 1, 24.63, .36, 1.76, and .25. The relative strengths that we got on 10/12/19 for J23100, J23101, J23103, and J23105 were 1, 45.97, 3.20, and 1.26. In addition we measured promoter J23118 twice and got the strengths 1.13 and 1.32.
Here is the relative promoter strengths that we got from the qPCR. Baltimore BioCrew in blue compared to the 2006 Berkeley iGEM in orange.
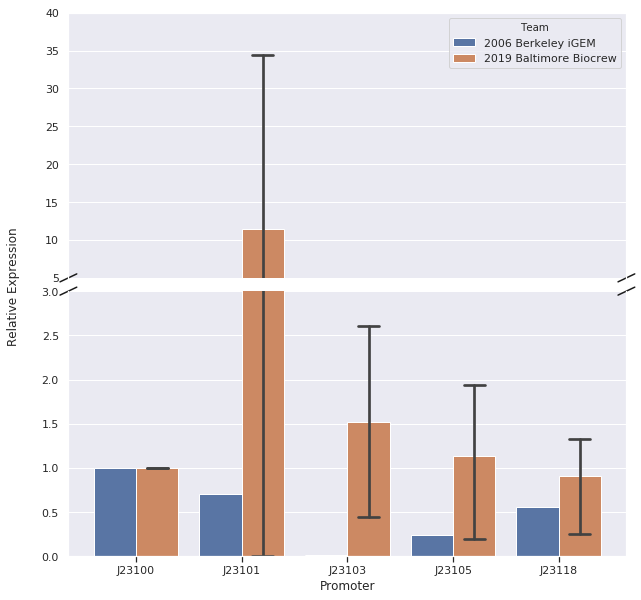
To support our RNA measurements we also measured fluorescence of the liquid cultures we used to extract RNA. The cultures were grown overnight so we expected the bacteria to be at the stationary phase, but we measured OD to normalize any differences in growth.
| Promoter | OD | fluorescence | fluorescence divided by OD | corrected relative expression | reported relative expression |
|---|---|---|---|---|---|
| BBa_J23100 | 0.876 | 250 | 285.38 | 1 | 1 |
| BBa_J23101 | 0.674 | 255 | 378.33 | 1.33 | 0.7 |
| BBa_J23103 | 1.1 | 230 | 209.09 | 0.73 | 0.01 |
| BBa_J23105 | 1.08 | 215.74 | 209.09 | 0.76 | 0.24 |
| BBa_J23118 | 1.04 | 238 | 228.84 | 0.80 | 0.56 |