Difference between revisions of "Part:BBa J04450:Experience"
(→Team OUC-China 2014) |
|||
| Line 2: | Line 2: | ||
This experience page is provided so that any user may enter their experience using this part.<BR>Please enter | This experience page is provided so that any user may enter their experience using this part.<BR>Please enter | ||
how you used this part and how it worked out. | how you used this part and how it worked out. | ||
| + | |||
| + | == '''Team Warwick 2015''' == | ||
| + | |||
| + | <p> | ||
| + | Our team considered using this part as part of a system of binding different coloured cells together in order to demonstrate specific cell placement. We characterised this part is order to determine the optimal amount of IPTG required for inducing the gene, and the copy number necessary to express the fluorescence brightly. The results for this can be seen below. | ||
| + | </p> | ||
| + | |||
| + | <p> | ||
| + | We cloned J04450 into three plasmid with varying copy numbers, in order from highest to lowest copy number they are: pSB1K3, pSB3K3, and pSB4K5. These plasmids were then transformed into electrocompetent MG1655 Z1 cells and grown overnight. THe next morning the cells were refreshed, and different concentrations of IPTG (0uM, 250uM, and 500uM) were added to induce them. For each of the three plasmids in each IPTG concentrations, three biological replicates were made, and when OD600 and RFP absorbance were measured, three technical replicates were made, for a total of 81 copies of the gene grown. The RFP absorbance and OD600 of these cells were measured over 20 hours. The OD600 over time was used to determine at what OD the cells were in steady state. This was then compared to the RFP measured at that time and graphed to show RFP expression per cell. | ||
| + | </p> | ||
| + | |||
| + | <img src="https://static.igem.org/mediawiki/2015/8/80/Warwick_J04450_Characterisation_graph.png"> | ||
| + | |||
| + | <p> | ||
| + | The graph shows that RFP expression was highest in the pSB1K3 and pSB4K5 plasmids, and that there was little difference in expression between the 250uM and 500uM concentration of IPTG. 0uM IPTG universally showed almost no expression. pSB1K3 should have the highest copy number and pSB4K5 should have the lowest copy number, so it's curious that they both expressed RFP very well. This could be due to a mutation in the pSB4K5 causing it to have a much higher copy number than usual. It is documented <a href="http://www.ncbi.nlm.nih.gov/pubmed/1283002">here</a> that a single point mutation can increase the copy number of a plasmid. The pSB4K5 plasmid we tested has been sent for sequencing in order to determine whether this is the case. | ||
| + | </p> | ||
| + | |||
| + | <p> | ||
| + | The raw data for this characterisation can be found <a href="https://static.igem.org/mediawiki/2015/7/70/Warwick_J04450_raw_data.txt">here</a>, and analysed data for this set can be found <a href"Warwick_J04450_in_pSB1K3_analysed_results.txt">here</a>, <a href="https://static.igem.org/mediawiki/2015/8/8f/Warwick_J04450_in_pSB3K3_analysed_results.txt">here</a>, and <a href="https://static.igem.org/mediawiki/2015/5/54/Warwick_J04450_in_pSB4K5_analysed_results.txt">here</a>. The analysed results used to generate the graph of RFP/OD are <a href="https://static.igem.org/mediawiki/2015/6/64/Warwick_J04450_RFP_OD_analysed_results.txt">here</a>. | ||
| + | </p> | ||
Revision as of 15:24, 22 September 2015
This experience page is provided so that any user may enter their experience using this part.
Please enter
how you used this part and how it worked out.
Team Warwick 2015
Our team considered using this part as part of a system of binding different coloured cells together in order to demonstrate specific cell placement. We characterised this part is order to determine the optimal amount of IPTG required for inducing the gene, and the copy number necessary to express the fluorescence brightly. The results for this can be seen below.
We cloned J04450 into three plasmid with varying copy numbers, in order from highest to lowest copy number they are: pSB1K3, pSB3K3, and pSB4K5. These plasmids were then transformed into electrocompetent MG1655 Z1 cells and grown overnight. THe next morning the cells were refreshed, and different concentrations of IPTG (0uM, 250uM, and 500uM) were added to induce them. For each of the three plasmids in each IPTG concentrations, three biological replicates were made, and when OD600 and RFP absorbance were measured, three technical replicates were made, for a total of 81 copies of the gene grown. The RFP absorbance and OD600 of these cells were measured over 20 hours. The OD600 over time was used to determine at what OD the cells were in steady state. This was then compared to the RFP measured at that time and graphed to show RFP expression per cell.
<img src=" ">
">
The graph shows that RFP expression was highest in the pSB1K3 and pSB4K5 plasmids, and that there was little difference in expression between the 250uM and 500uM concentration of IPTG. 0uM IPTG universally showed almost no expression. pSB1K3 should have the highest copy number and pSB4K5 should have the lowest copy number, so it's curious that they both expressed RFP very well. This could be due to a mutation in the pSB4K5 causing it to have a much higher copy number than usual. It is documented <a href="http://www.ncbi.nlm.nih.gov/pubmed/1283002">here</a> that a single point mutation can increase the copy number of a plasmid. The pSB4K5 plasmid we tested has been sent for sequencing in order to determine whether this is the case.
The raw data for this characterisation can be found <a href="https://static.igem.org/mediawiki/2015/7/70/Warwick_J04450_raw_data.txt">here</a>, and analysed data for this set can be found <a href"Warwick_J04450_in_pSB1K3_analysed_results.txt">here</a>, <a href="https://static.igem.org/mediawiki/2015/8/8f/Warwick_J04450_in_pSB3K3_analysed_results.txt">here</a>, and <a href="https://static.igem.org/mediawiki/2015/5/54/Warwick_J04450_in_pSB4K5_analysed_results.txt">here</a>. The analysed results used to generate the graph of RFP/OD are <a href="https://static.igem.org/mediawiki/2015/6/64/Warwick_J04450_RFP_OD_analysed_results.txt">here</a>.
Team NRP-UEA_Norwich 2014
Our team has converted this part into a series of four "GoldenGate Flipper modules" that can be used to convert four types of standard part defined in the GoldenGate MoClo Assembly Standard into standard BioBricks. This was done by inserting an inverted pair of BsaI recognition sequences either side of the original RFP reporter part. The flipper reaction is a one-pot, one-step, digestion-ligation GoldenGate cloning reaction. When cloning is successful the colonies become white as the RFP sequence is replaced by the new part. After the reaction is complete, no part of the BsaI recognition sequence will remain between the BioBrick suffix and the part.
Four types of Golden Gate Parts can be converted. The four new parts differ by the identity of the 4 base-pair 5' overhangs produced by digestion with BsaI.
Bba_K1467100 Pro + 5U MoClo Flipper (GGAG-AATG)
Bba_K1467200 CDS1 MoClo Flipper (AATG-GCTT)
Bba_K1467300 3U + Ter MoClo Flipper (GCTT-CGCT)
Bba_K1467400 Transcriptional Unit MoClo Flipper (GGAG-CGCT)
The image below shows the sequences inserted (in orange) between the RFP and the BioBrick prefix and suffix that enable the flipper to accept MoClo Pro+5U parts (which begin GGAG and end AATG) in a one-step digestion-ligation Golden Gate cloning reaction.
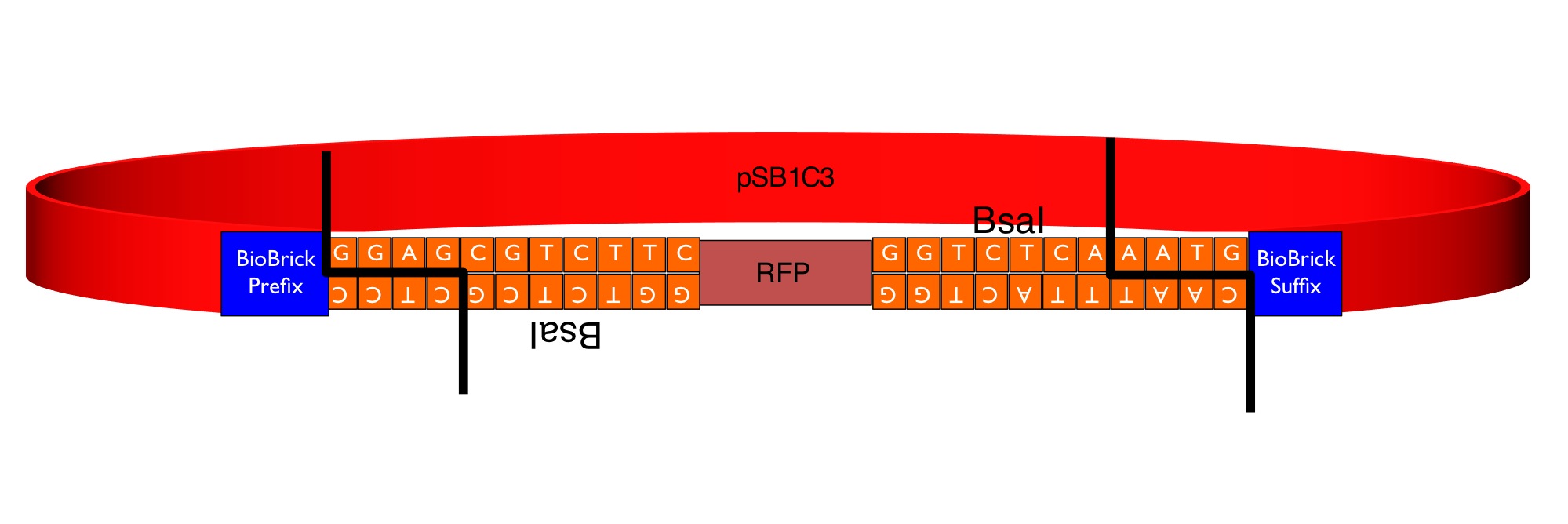
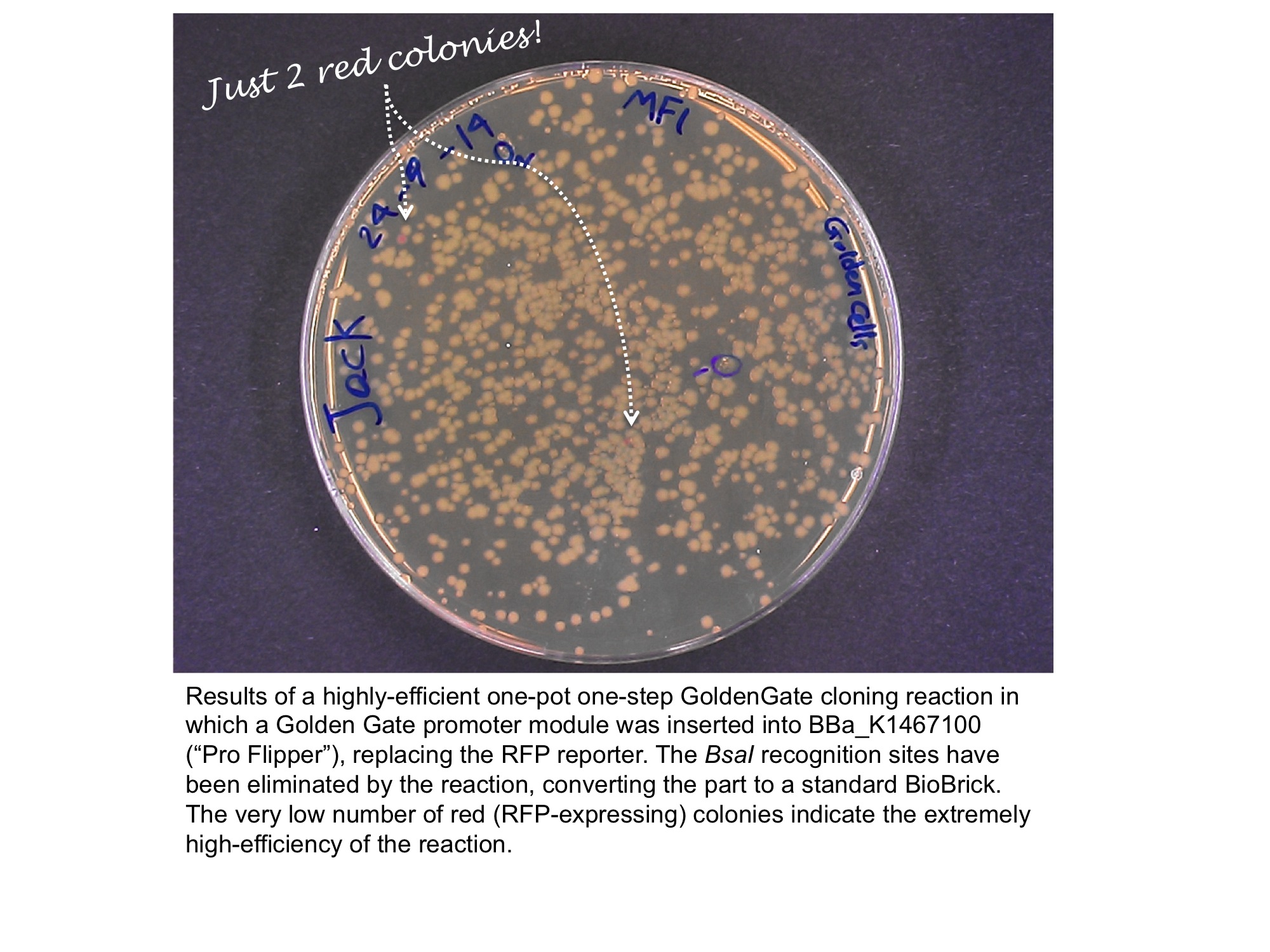
Below is an example of a Golden Gate one-step, one-pot cloning reaction with the improved BioBrick BBa_K1467100, where the restriction enzyme, ligase and (intact, uncut) donating and recipient plasmids were all incubated together in a single reaction before transformation into competent E.coli cells. Cloning was very successful as the vast majority of the colonies are white as the RFP sequence was replaced by the new part.
Team OUC-China 2014
This year our team use this part as a reporter gene link with BBa_k1439000. In order to mark our function plasmid. we find some problem about this part. The result shows Part BBa_J04450 didn’t work well as the reporter gene of the plasmid. The expression quantity of RPF is low or even next to zero under the condition of no resistance screening.

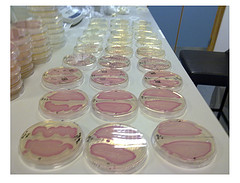
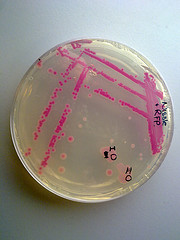
Figure1.
Figure1. Top10 strain containing BBa_J04450(pSB1C3).Culture medium contains no chloramphenicol to the left, to the right contains chloramphenicol. Under the same condition,37℃ 12h,RFP didn’t express in the culture medium without chloramphenicol.
Team NRP-UEA_Norwich 2013
Our team improved this part by adding an NdeI restriction site in front of the promoter region, upstream of the RFP gene. This means a restriction digest can be performed and the promoter or RFP gene exchanged to a allow further cloning. Please see part BBa_K1041000
Our aim was to evaluate RFP expression quantity of E. coli. In order to obtain this information, we agreed that the measurement of fluorescence intensity in certain OD values is the best approach. However, fluorescence quenching was a setback which we must overcome for accurate and reliable results. Because, as a result of quenching, fluorescence intensity counts do not give the actual amount of fluorescence expected from a certain OD value. Therefore we employed a curve fitting method suggested by Zhang et. al (2010) which is claimed to take the quenching problem into consideration

Where If is the fluorescence intensity; OD is the cell density; y0 is the offset; A1 is the amplitude; and t1 is the decay constant.
We chose BBa_J04450 for this experiment. And our negative control was again BBa_B0034.
There were nine bacterial concentrations prepared for one to five hours of IPTG induction. (All concentrations were prepared as triplicates.) Then a complete graph of all the induction durations was plotted with the help of the function shown above:

As seen in the graph, fluorescence behaves exactly as expected and does not increase linearly with the increased bacterial concentration. However, IPTG induction effect is clear on fluorescence intensity. (Error bars were too small to be visible.)

Finally, the variables A1 and t1 of the fittings of each induction duration were used to obtain a mean fluorescence intensity (MFI) value:



By conducting this experiment, we were able to characterize RFP expression pattern of the part BBa_J04450 in the host Top Ten. Theoretical mean fluorescence intensity reflects the fluorescence quantum efficiency of the fluorophore RFP, and was found to be unrelated to the cell concentration. Since it is obvious from the results we have obtained that change in the OD does not correlate linearly with the change in the fluorescence intensity. Therefore this model proposes a solution to the unpredictability of the fluorescence amount coming from a certain bacterial population.
The characterization was performed by METU-TURKEY 2010 IGEM TEAM.
Applications of BBa_J04450
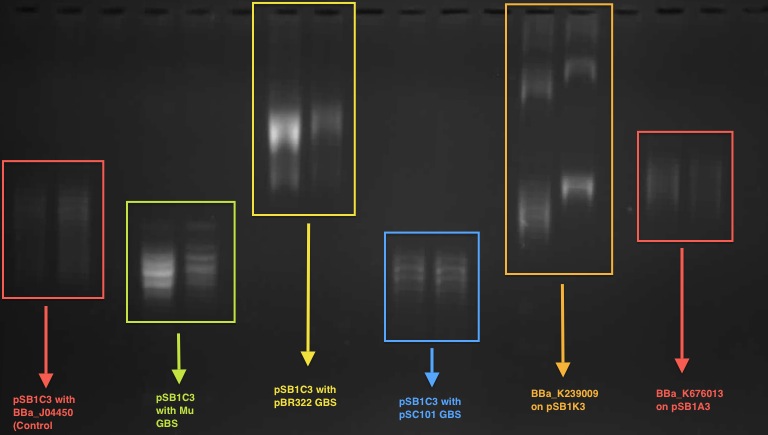
According to our result in the 1D chloroquine gel electrophoresis (at 2.5 μg/mL), the plasmid with the BBa_J04450 (on pSB1C3 backbone) migrated further compared to a plasmid with a known GBS (from pBR322 plasmid - BBa_K676012) on pSB1C£. At the chloroquine concentration which we used, the more negatively supercoiled plasmid will migrate further while the more relaxed molecules will travel slower through the gel.
So this indicates that the device BBa J04450 might have a potential gyrase binding site which facilitates the introduction of negative supercoils into plasmid compared to pBR322 GBS.
This characterisation was carried out by the UCL iGEM team 2011.
Tianjin iGEM 2012
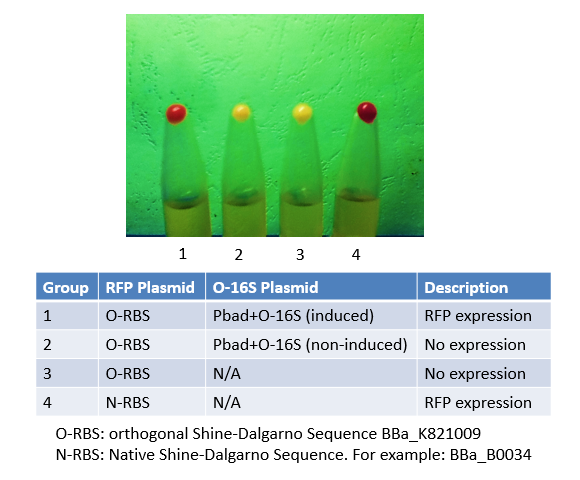
Originally, we used this BioBrick to express RFP in Group 4. We mutated the Shine-Dalgarno sequence of this BioBrick to make an orthogonal counterpart, BBa_K821001. As shown in the image, No. 4 is the product of this BioBrick, and No.1 is the product of its orthogonal counterpart. The results justified both this BioBrick and our BBa_K821001 worked very well, and expressed considerable amount of protein.
Glasgow iGEM 2011
It was found that the Red fluorescence of this protein was not consistently apparent when expressed in biofilm - although some results were observed. Numerous attempts to transform this construct into E. coli Nissle 1917 cells in a biofilm forming assay were unsuccessful, however, the RPF expressed readily into E. coli Nissle 1917 cells when grown on an agar plate.
The antibiotic resistance was still carried across when the culture was inoculated into the biofilm assay indicating that the plasmid was still present. The cultures were shaken at room temperature overnight.
It was thought that the lack of fluorescence was caused by the anoxic conditions generated by the biofilm assay, which may have resulted in poor folding of the chromophore. These results show that RFP is not the most efficient reporter molecule for biofilm, we would recommend something like iLOV, a new biobrick submitted by the Glasgow iGEM team this year [1].
Glasgow iGEM team 2011
Haynes Lab, 2012
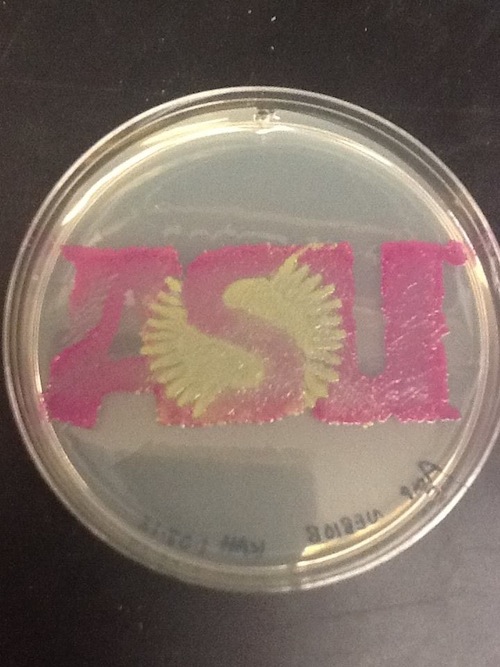
I used BBa_J04450 to demonstrate the expression of RFP in E. coli for a biomedical engineering course at Arizona State University.
- Cells: NEB10B
- Medium: LB Lennox agar, 100 ug/mL ampicillin
- The yellow/green area contains bacteria expressing GFP from BBa_I13522
User Reviews
UNIQcf8477de0f4760aa-partinfo-00000007-QINU
|
•••••
Antiquity |
This review comes from the old result system and indicates that this part worked in some test. |
|
•••••
|
We've used this part as an alternative to the suicide part p1010 (ccdB) for selecting against cells that have been transformed with a [http://ginkgobioworks.com/support/BioBrick_Assembly_Manual.pdf destination plasmid]. It worked perfectly, and if i might say, it looks smashingly in LB medium ;). |
|
•••••
|
We obtained this part from the Spring 2011 distribution plate. It expresses wonderfully in NEB10B cells from New England Biolabs. It does not express well in the fast-growing strain DH5-alpha Turbo, which we've found doesn't express transgenes very well in general. |
UNIQcf8477de0f4760aa-partinfo-0000000B-QINU