Difference between revisions of "Part:BBa K592009:Experience"
m (→User Reviews) |
|||
| Line 18: | Line 18: | ||
<!-- End of the user review template --> | <!-- End of the user review template --> | ||
<!-- DON'T DELETE --><partinfo>BBa_K592009 EndReviews</partinfo> | <!-- DON'T DELETE --><partinfo>BBa_K592009 EndReviews</partinfo> | ||
| + | |||
| + | {|width='80%' style='border:1px solid gray' | ||
| + | |- | ||
| + | |width='10%'| | ||
| + | <partinfo>BBa_E1010 AddReview 5</partinfo> | ||
| + | <I>Immudzen</I> | ||
| + | |width='60%' valign='top'| | ||
| + | |||
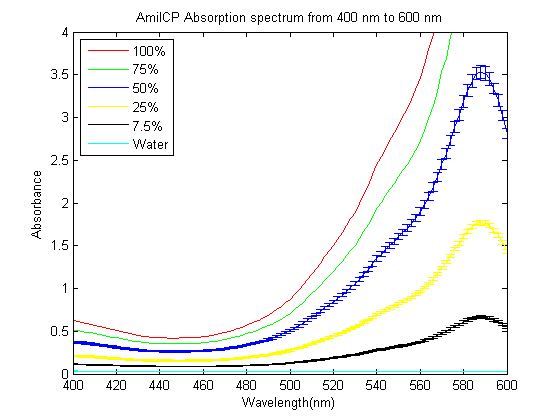
| + | As part of the 2013 CU Boulder project we worked on separating RFP from AmilCP and we measured the spectrum of AmilCP from 400 nm to 600 nm. | ||
| + | |||
| + | [[File:Cu_Amilcp.png]] | ||
| + | |||
| + | We also found that when running on an agarose gel RFP will run down on the gel while AmilCP runs up on the gel. | ||
| + | |}; | ||
| + | |||
{|width='80%' style='border:1px solid gray' | {|width='80%' style='border:1px solid gray' | ||
Revision as of 00:08, 23 September 2013
This experience page is provided so that any user may enter their experience using this part.
Please enter
how you used this part and how it worked out.
Applications of BBa_K592009
User Reviews
UNIQdf5215ba55d10779-partinfo-00000000-QINU UNIQdf5215ba55d10779-partinfo-00000001-QINU
|
•••••
Immudzen |
As part of the 2013 CU Boulder project we worked on separating RFP from AmilCP and we measured the spectrum of AmilCP from 400 nm to 600 nm. We also found that when running on an agarose gel RFP will run down on the gel while AmilCP runs up on the gel. |
|
•••••
iGEM 2013 Hong_Kong_HKUST Trevor Y.H. HO |
Error creating thumbnail: Invalid thumbnail parameters This part was employed in testing our Gibson Assembly protocol. In the experiment design, the mRFP coding DNA sequence in J04450 was replaced with that of amilCP. The resulting plate from transformation of the post-reaction mixture is shown. The color of the blue chromoprotein was clearly distinguished from that of mRFP, which aided in my evaluation of the performance of Gibson Assembly. |
|
BBa_K592009 iGEM Groningen 2012 |
Our team has managed to couple this biobrick part with our promoters: alsT, fnr, and sboA. The cloning was done in BBa_K818000 (plasmid backbone for B. subtilis, engineered by team Groningen 2012), to allow color expression in B. subtilis. We utilized a strong RBS BBa_B0034 for pigment expression in E. coli and B. subtilis. The purple/blue colour was strongly visible in E.coli without any induction, while the expression in B. subtilis was more subtle (B. subtilis colony looks slightly blue on the plate agar). After induction of the promoter before AmilCP, B. subtilis also turned clearly purple/blue. Please have a look at the page of BBa_K818400 for more information.
|
|
|
iGEM Team Uppsala University 2012 We have observed that expression of amilCP confers a noticeable fitness cost in E coli. This is surprising, given that such behaviour is not seen in other homologous chromoproteins and fluorescent proteins. We thus recommend using the new aeBlue (BBa_K864401) chromoprotein for blue color expression, which also has a more clear blue color. |