Difference between revisions of "Part:BBa K3610007"
(→Usage and Biology) |
(→Usage and Biology) |
||
| (8 intermediate revisions by the same user not shown) | |||
| Line 6: | Line 6: | ||
===Usage and Biology=== | ===Usage and Biology=== | ||
| + | |||
| + | [[File:T--UZurich--EFR ecto.png|300px|thumb|left|Figure 1: EFR (Uniprot COLGT6)]] | ||
Elongation factor-thermo unstable receptor (EFR) from A. thaliana is a plant pattern-recognition receptor (PRR). It is a cell surface receptor and part of the plants firts defence mechanism against potential pathogens. The EFR receptor is also a leucin-rich-repeats (LRR) receptor-like serine/threonine-protein kinase. The protein consists of an extracellular domain with leucin-rich repeats, a ligand binding domain found in many receptors, a single-pass transmembrane domain and finally an intracellular kinase domain. The ligand binding domain from EFR has high specificity to a bacterial pathogen-associated moleculat pattern (PAMP), namely the epitope elf18 of the abundant protein Elongation Factor Tu (EF-Tu), which is catalyzes the binding of aminoacyl-tRNA (aa-tRNA) to the ribosome in most prokaryotes and therefore is evolutionarily highly conserved. This makes the EFR a receptor that can be activated by the presence of a huge variety of bacteria. Upon binding of the ligand to the extracellular domain, the receptor dimerizes with its coreceptor BRI1-associated receptor kinase (BAK1). This interaction triggers the activation of the intracellular kinase domain of EFR and BAK1, initiating a signal cascade leading to an upregulation of immune response mechanisms. | Elongation factor-thermo unstable receptor (EFR) from A. thaliana is a plant pattern-recognition receptor (PRR). It is a cell surface receptor and part of the plants firts defence mechanism against potential pathogens. The EFR receptor is also a leucin-rich-repeats (LRR) receptor-like serine/threonine-protein kinase. The protein consists of an extracellular domain with leucin-rich repeats, a ligand binding domain found in many receptors, a single-pass transmembrane domain and finally an intracellular kinase domain. The ligand binding domain from EFR has high specificity to a bacterial pathogen-associated moleculat pattern (PAMP), namely the epitope elf18 of the abundant protein Elongation Factor Tu (EF-Tu), which is catalyzes the binding of aminoacyl-tRNA (aa-tRNA) to the ribosome in most prokaryotes and therefore is evolutionarily highly conserved. This makes the EFR a receptor that can be activated by the presence of a huge variety of bacteria. Upon binding of the ligand to the extracellular domain, the receptor dimerizes with its coreceptor BRI1-associated receptor kinase (BAK1). This interaction triggers the activation of the intracellular kinase domain of EFR and BAK1, initiating a signal cascade leading to an upregulation of immune response mechanisms. | ||
When the ectodomains of EFR and another PRR are swapped, the new chimeric receptors are sometimes still functional (depending on which second receptor is chosen). It has further been shown, that for binding of the ligand and dimerization with BAK1 the extracellular domain and the transmembrane domain are needed. The kinase domain is not necessary for initiating this interaction. | When the ectodomains of EFR and another PRR are swapped, the new chimeric receptors are sometimes still functional (depending on which second receptor is chosen). It has further been shown, that for binding of the ligand and dimerization with BAK1 the extracellular domain and the transmembrane domain are needed. The kinase domain is not necessary for initiating this interaction. | ||
| − | For visualizing the interaction of EFR with BAK1, the cytoplasmic domain can be replaced with a split fluorescent protein or another protein that generates a visual output. In our iGEM project, we fused this sequence to a split-mCherry protein and a split-luciferase in another experiment and then coexpressed the part with BAK1 carrying the counterpart to the split protein instead of the intracellular kinase domain in S. cerevisiae. | + | For visualizing the interaction of EFR with BAK1, the cytoplasmic domain can be replaced with a split fluorescent protein or another protein that generates a visual output. In our iGEM project, we fused this sequence to a split-mCherry protein and a split-luciferase in another experiment and then coexpressed the part with BAK1 carrying the counterpart to the split protein instead of the intracellular kinase domain in <i>S. cerevisiae</i>. |
==Characterization== | ==Characterization== | ||
| − | === | + | ===Expression with YFP=== |
We fused the ectodomain of EFR to the yellow fluorescent protein and expressed it in <i>S. cerevisiae</i>. For a more elaborate characterization of this part see [[Part:K3610045]]. | We fused the ectodomain of EFR to the yellow fluorescent protein and expressed it in <i>S. cerevisiae</i>. For a more elaborate characterization of this part see [[Part:K3610045]]. | ||
| Line 21: | Line 23: | ||
After successful transformation of yeast cells we checked for enhanced fluorescence with confocal fluorescence microscopy. | After successful transformation of yeast cells we checked for enhanced fluorescence with confocal fluorescence microscopy. | ||
| − | [[File:T--UZurich-- | + | [[File:T--UZurich--Control.png|800px|thumb|none|Figure 2: Control]] |
| − | [[File:T--UZurich-- | + | [[File:T--UZurich--eEFR.png|800px|thumb|none| Figure 3: <i>S. cerevisiae transformed with eEFR construct]] |
| − | + | ||
Imaging of the <i>S. cerevisiae</i> cells, which had been previously transfected with plasmids containing this construct, revealed increased fluorescence levels than the untransfected control. These results imply increased expression of YFP, indicating expression of the EFR ectodomain. | Imaging of the <i>S. cerevisiae</i> cells, which had been previously transfected with plasmids containing this construct, revealed increased fluorescence levels than the untransfected control. These results imply increased expression of YFP, indicating expression of the EFR ectodomain. | ||
Additionally, the imaging results suggest that the proteins are in part localized at the cellular membrane, which is in alignment with our expectations as there is a secretion signal peptide and a transmembrane domain in the construct. | Additionally, the imaging results suggest that the proteins are in part localized at the cellular membrane, which is in alignment with our expectations as there is a secretion signal peptide and a transmembrane domain in the construct. | ||
| + | |||
| + | To see whether there actually is localization at the cell membrane, additional imaging with the membrane stain FM4-64 was carried out. | ||
| + | |||
| + | [[File:T--UZurich--UT Membrane Stain.png|440px|thumb|left|Figure 4: Untransformed Control,(A) : YFP, (B) : FM4-64, (C): light field. (D): merge.Imaging of untransfected S. cerevisiae cells reveals hardly any fluorescence within the YFP spectrum]]<br> | ||
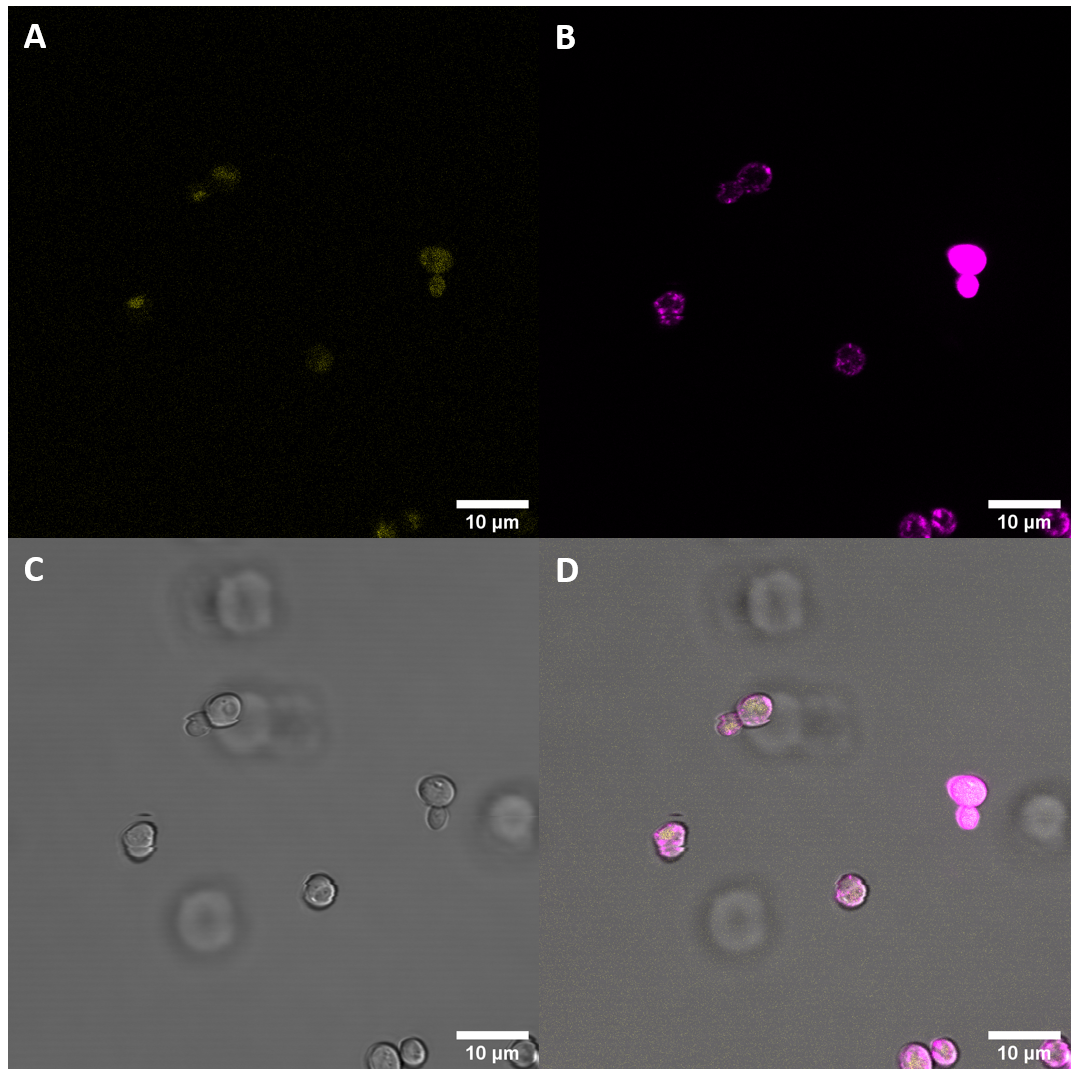
| + | [[File:T--UZurich--eEFR Membrane Stain.png|450px|thumb|none|Figure 5: eEFR S. cerevisiae: (A) : YFP, (B) : FM4-64, (C) : light field. (D): merge. The construct eEFR:YFP is expressed and in about 25% of the cells clearly co-localized with the FM4-64 stain.]] | ||
| + | |||
| + | Confocal fluorescence microscopy with membrane staining confirmed what had been suggested previously with initial microscopy imaging. The results suggest that the receptor protein fused to the YFP gets epxpressed in <i>S. cerevisiae</i> and shows clear ring structures co-localized with the FM4-64 stain, indicating that there is localization at the plasma membrane. | ||
====Spectrometry==== | ====Spectrometry==== | ||
| Line 62: | Line 70: | ||
<br> | <br> | ||
| − | [[File:T--UZurich-- | + | [[File:T--UZurich--Spectrometer1.png|500px|none|left|Figure 6: Fluorescence values standardized for OD600 of the different receptors (C=Control). Cells with BAK+ showed only weak fluorescence, while BAK-, eBAK and eEFR showed a strong increase in the fluorescence levels. CORE did not display any increase when compared with untreated <i>S. cerevisiae</i> cells (autofluorescence).]] |
Results of the plate reader suggested increased fluorescence through expression of YFP. The same was observed when cells were examined with the microscope. Therefore, <i>S. cerevisiae</i> cells express the plasmid containing the EFR ectodomain. Additionally, microscopy revealed localization at the cell membrane. This localization at the cell periphery is a big success as it shows, that the signal peptide from the alpha-Mating factor did, in fact induce translation into the membrane. Additionally, it was shown that the intracellular kinase domain is not necessary for membrane localization. | Results of the plate reader suggested increased fluorescence through expression of YFP. The same was observed when cells were examined with the microscope. Therefore, <i>S. cerevisiae</i> cells express the plasmid containing the EFR ectodomain. Additionally, microscopy revealed localization at the cell membrane. This localization at the cell periphery is a big success as it shows, that the signal peptide from the alpha-Mating factor did, in fact induce translation into the membrane. Additionally, it was shown that the intracellular kinase domain is not necessary for membrane localization. | ||
| − | + | ||
| + | ====Flow Cytometry==== | ||
| + | It has been important to us to examine a sample with different approaches simultaneously, which is why we were eager to also measure fluorescence intensity by flow cytometry. In a first phase, 100,000 cells were measured from each biological replicate (488/530 FITC channel in a BD FACSCanto II flow cytometer). | ||
| + | In the next phase, the biological replicates for each construct were pooled together and 200,000 cells from each sample were measured. | ||
| + | |||
| + | [[File:T--UZurich--FACs.png|800px|thumb|none|Figure 7: Left: single biological replicates. right: pooled samples. With the exception of the construct with eCORE, cells transfected with our constructs showed considerably higher overall fluorescence intensities than the negative control.]] | ||
| + | |||
| + | Flow catometry provided further evidence for expression of the construct in <i>S. cerevisiae</i>. Cells transfected with plasmids containing eEFR showed significantly increased fluorescence intensities when examined as single biological replicates, as well as when the replicates were pooled together in one sample. | ||
| + | |||
| + | This observations are a huge success. There is compelling evidence for the claim that the EFR receptor ectodomain gets expressed in <i>S. cerevisiae</i> and, if fused to the signal peptide of the alpha-Factor from yeast, also does get localized at the cell memrbane. | ||
| + | |||
| + | This makes plant PRRs, or at least the receptor EFR, an attractive tool for many applications in Synthetic biology, especially when we consider, that similar observations were made when the same experiment was conducted with the ectodomain of EFR's coreceptor BAK1 ([[Part:BBa K3610003]]) | ||
| + | |||
| + | |||
| + | ===Usage with split-NanoLuc=== | ||
| + | |||
| + | We fused this part to the SmallBit part of the NanoBit system, a split luciferase (eEFR:SBit). We managed to express the CORE:SBit together with the ectodomain of the plant PRR BAK1 which was fused to the LargeBit part of the split NanoLuc protein (eBAK1:LBit). We were able to express both constructs in <i>S. cerevisiae</i>. For a more detailed description see the parts: [[Part:BBa K3610038]] and [[Part:BBa K3610043]]. | ||
| + | |||
| + | After transfection of the cells with the two plasmids, we tested whether the split-NanoLuc proteins would be able to interact, which would reconstitute its functionality. We further were interested in the interaction between the EFR and BAK1 ectoomain. EFr and BAK1 naturally interact upon binding of elf18. Should both receptors be expressed properly in the cell membrane of <i>S. cerevisiae</i>, the ligand will be able to bind to the EFR ectodomain. If this is sufficient to drive dimerization of the two receptor in <i>S. cerevisiae</i>, then the NanoLuc parts will be more likely to interact as well, which would lead to a higher amount of functional luciferase proteins in the cell. | ||
| + | |||
| + | To test for dimerization of the luciferase parts and also the two receptors, an assay with a luminometer was performed. | ||
| + | |||
| + | ====Luminescence assay==== | ||
| + | Samples with cells which were transfected with the two mentioned plasmids (eBAK1 and eCORE) and samples containing <i>S. cerevisiae</i> cells that were not transfected with any plasmids (UT). | ||
| + | |||
| + | Optical densities (OD600) of all samples were adjusted to 0.34.<br> | ||
| + | For each type of sample, three types of measurements were made: | ||
| + | *1 µL of deionized water added (no elicitor) | ||
| + | *1 µL of epitope elf18 added | ||
| + | *1 µL of epitope csp22 added | ||
| + | |||
| + | Each measurement was done four times with sample size 50µL. To each well 50 µL NanoGlo solution was added (50:1 buffer to furimazine)<br> | ||
| + | |||
| + | The average values of the four measurements are displayed in the plot below. | ||
| + | [[File:T--UZurich--NLuc_Assay_1.png|600px|thumb|none|Figure 1: Average Luminescence Levels over time after 30 minutes incubation]] | ||
| + | The results suggest that the receptors do get expressed in the yeast cells. It further is shown that the receptor either drive interaction between the two NanoBit parts or at least do not inhibit this interaction. | ||
| + | Luminescence levels were, however, not increased when the bacterial epitope elf18 was present, the elicitor which normally initiates ligand-dependent interaction between BAK1 and EFR. | ||
| + | These results suggest that there is no or very little elf18-driven dimerization of the two receptor-ectodomains EFR and BAK1 when they are expressed in <i>S. cerevisiae</i>. | ||
| + | |||
| + | ====Luminescence Assay 2==== | ||
| + | |||
| + | The assay was repeated in the a similar manner with the following differences: | ||
| + | *We did not test again for samples which had been transformed with eCORE plasmids. So we only had samples for untransformed yeast cells and for cells transfected with the plasmids containting [[Part:BBa K3610038]] and [[Part:BBa K3610043]]. | ||
| + | *The optical density of all samples was adjusted to OD600 = 0.26. | ||
| + | *Measurement started immediately after addition of the bacterial elicitor and the NanoGlo solution. | ||
| + | |||
| + | [[File:T--UZurich--NLuc_Assay_2.png|600px|thumb|none|Figure 10: Average Luminescence Levels over time, measured directly after addition of bacterial elicitor and furimazine]] | ||
| + | |||
| + | Our observations of the second luminescence assay matched our expectations after the first one. Luminescence levels were indeed increased when the cells had previously been transformed with our constructs. It was again the case, however, that reconstitution of the NanoLuc protein was not driven by the ligand-dependent interaction of the plant receptors. It seemed, again, to be the case, that addition of bacterial elicitor did in fact decrease the measured luminescent output. | ||
| + | |||
| + | |||
| + | ====Luminescence Assay 3==== | ||
| + | The assay was repeated again with the following differences (compared to the first assay): | ||
| + | *We did not test again for samples which had been transformed with eCORE plasmids. So we only had samples for untransformed yeast cells and for cells transfected with the plasmids containting [[Part:BBa K3610038]] and [[Part:BBa K3610043]]. | ||
| + | *NanoGlo was added before the measurements were made, the bacterial elicitor was added right before measurement | ||
| + | *Optical density was adjusted to OD600 = 0.5. We waited, however, for almost two hours, during which the cells were kept in TE buffer. During this time, a lot of the cells died, which decresaed the amount of whole living cells as TE supports cell lysis. | ||
| + | *Only one bacterial epitope (elf18) was added. | ||
| + | |||
| + | [[File:T--UZurich--NLuc_Assay_3.png|600px|thumb|none|Figure 11: Average Luminescence Levels over time, furimazine added 30 minutes before measurement and bacterial elicitor added immediately before.]] | ||
| + | |||
| + | A third time, we were able to observe that expression of our constructs increased luminescence intensity when the substrate furimazine was added. We, additionally, observed that the increase in luminescence was not initiated or significantly enhanced by presence of the bacterial epitope elf18. | ||
| + | |||
| + | ====Summary of Luminescence Assays==== | ||
| + | We conducted for assays with a plate reader of the type <i>Synergy H1</i>. Each time, luminescence was strongly increased when the samples had been transfected with plant PRRs fused to the NanoBits. This has especially been the case when the plant recpetors EFR and its coreceptor BAK1 were used. | ||
| + | The increase in luminescence intensity did not seem to be driven by epitope.dependent interaciton of the two receptors. It is possible that, unlinke plants, <i>S. cerevisiae</i> cells lack any mechanisms to prevent unprovoked interaction between the target-receptor EFR and BAK1. Another possible explanation for the results could be that the receptors proteins are not processed the same way as they are in plant cells. For example, in plant cells, the receptor proteins get highly glycosylated. It is possible that this glycosylation influences the structure and the functionality of the receptor proteins. | ||
| + | |||
| + | ==Conclusion UZürich 2020== | ||
| + | The EFR ectodomain was a crucial part of our project. Nothing we set out to do could have worked without this part. We are happy with what we have achieven, even though there is so much more left to discover, much more to try and apply. | ||
| + | |||
| + | We are nevertheless happy, to have used this part in the following ways: | ||
| + | *Demonstration expression by fusing this part to YFP and transfecting <i>S. cerevisiae</i> cells with the construct. We saw that the cells express the receptor and, in addition, some of it gets trafficked to the plasma membrane. Compared to other receptors, the EFR ectodomain has the highest proportion of cells with clear localization. | ||
| + | *Dimerization Assay with eBAK and split-mCherry. Fused to the split-mCherry parts, coexpression of EFR and BAK1 did not reveal increased fluorescence when compared to the control. Interestingly, there seems to be an effect of the bacterial elicitor elf18 on fluorescence intensity. | ||
| + | *Dimerization Assay with BAK1 and split-NanoLuc. Fused to the NanoBit parts, coexpression of this part and BAK1 leads to a luminescent output in the prescence of the NanoLuc substrate furimazine. It does, however, seem that dimerization of the split-NanoLuc is not driven by ligand-dependent interaction of the plant PRRs. | ||
<!-- --> | <!-- --> | ||
Latest revision as of 21:20, 27 October 2020
EFR Ectodomain without signal sequence
This part is the ectodomain of the plant PRR EFR, a cell surface receptor determining the preception of the bacterial Elongation-Factor Tu.
Usage and Biology
Elongation factor-thermo unstable receptor (EFR) from A. thaliana is a plant pattern-recognition receptor (PRR). It is a cell surface receptor and part of the plants firts defence mechanism against potential pathogens. The EFR receptor is also a leucin-rich-repeats (LRR) receptor-like serine/threonine-protein kinase. The protein consists of an extracellular domain with leucin-rich repeats, a ligand binding domain found in many receptors, a single-pass transmembrane domain and finally an intracellular kinase domain. The ligand binding domain from EFR has high specificity to a bacterial pathogen-associated moleculat pattern (PAMP), namely the epitope elf18 of the abundant protein Elongation Factor Tu (EF-Tu), which is catalyzes the binding of aminoacyl-tRNA (aa-tRNA) to the ribosome in most prokaryotes and therefore is evolutionarily highly conserved. This makes the EFR a receptor that can be activated by the presence of a huge variety of bacteria. Upon binding of the ligand to the extracellular domain, the receptor dimerizes with its coreceptor BRI1-associated receptor kinase (BAK1). This interaction triggers the activation of the intracellular kinase domain of EFR and BAK1, initiating a signal cascade leading to an upregulation of immune response mechanisms. When the ectodomains of EFR and another PRR are swapped, the new chimeric receptors are sometimes still functional (depending on which second receptor is chosen). It has further been shown, that for binding of the ligand and dimerization with BAK1 the extracellular domain and the transmembrane domain are needed. The kinase domain is not necessary for initiating this interaction.
For visualizing the interaction of EFR with BAK1, the cytoplasmic domain can be replaced with a split fluorescent protein or another protein that generates a visual output. In our iGEM project, we fused this sequence to a split-mCherry protein and a split-luciferase in another experiment and then coexpressed the part with BAK1 carrying the counterpart to the split protein instead of the intracellular kinase domain in S. cerevisiae.
Characterization
Expression with YFP
We fused the ectodomain of EFR to the yellow fluorescent protein and expressed it in S. cerevisiae. For a more elaborate characterization of this part see Part:K3610045.
Fluorescent Microscopy
After successful transformation of yeast cells we checked for enhanced fluorescence with confocal fluorescence microscopy.
Imaging of the <i>S. cerevisiae cells, which had been previously transfected with plasmids containing this construct, revealed increased fluorescence levels than the untransfected control. These results imply increased expression of YFP, indicating expression of the EFR ectodomain. Additionally, the imaging results suggest that the proteins are in part localized at the cellular membrane, which is in alignment with our expectations as there is a secretion signal peptide and a transmembrane domain in the construct.
To see whether there actually is localization at the cell membrane, additional imaging with the membrane stain FM4-64 was carried out.
Confocal fluorescence microscopy with membrane staining confirmed what had been suggested previously with initial microscopy imaging. The results suggest that the receptor protein fused to the YFP gets epxpressed in S. cerevisiae and shows clear ring structures co-localized with the FM4-64 stain, indicating that there is localization at the plasma membrane.
Spectrometry
In addition to analyzing the cells with a microscope, we conducted a fluorescence assay with a plate reader. We conducted this experiment for multiple receptors at the same time. This way we were able to compare the expression levels of different receptors.
The results of these measurements are displayed in the table below.
| Fluorescence normalized for OD600 | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Control | BAK+ | BAK- | eBAK | eCORE | eEFR | ||||
| 4185,221063 | 9731,614266 | 26067,19254 | 28118,24739 | 3712,946478 | 23379,84399 | ||||
If we set the values for the Control to 1 (Control = 1), then we get the fluorescence levels relative to the control, which is again diplayed in the table below.
| Control | eCORE | eEFR | BAK- | BAK+ | eBAK |
|---|---|---|---|---|---|
| 1 | 0,8871565975 | 5,586286516 | 6,228390841 | 2,325233033 | 6,718461693 |
Results of the plate reader suggested increased fluorescence through expression of YFP. The same was observed when cells were examined with the microscope. Therefore, S. cerevisiae cells express the plasmid containing the EFR ectodomain. Additionally, microscopy revealed localization at the cell membrane. This localization at the cell periphery is a big success as it shows, that the signal peptide from the alpha-Mating factor did, in fact induce translation into the membrane. Additionally, it was shown that the intracellular kinase domain is not necessary for membrane localization.
Flow Cytometry
It has been important to us to examine a sample with different approaches simultaneously, which is why we were eager to also measure fluorescence intensity by flow cytometry. In a first phase, 100,000 cells were measured from each biological replicate (488/530 FITC channel in a BD FACSCanto II flow cytometer). In the next phase, the biological replicates for each construct were pooled together and 200,000 cells from each sample were measured.
Flow catometry provided further evidence for expression of the construct in S. cerevisiae. Cells transfected with plasmids containing eEFR showed significantly increased fluorescence intensities when examined as single biological replicates, as well as when the replicates were pooled together in one sample.
This observations are a huge success. There is compelling evidence for the claim that the EFR receptor ectodomain gets expressed in S. cerevisiae and, if fused to the signal peptide of the alpha-Factor from yeast, also does get localized at the cell memrbane.
This makes plant PRRs, or at least the receptor EFR, an attractive tool for many applications in Synthetic biology, especially when we consider, that similar observations were made when the same experiment was conducted with the ectodomain of EFR's coreceptor BAK1 (Part:BBa K3610003)
Usage with split-NanoLuc
We fused this part to the SmallBit part of the NanoBit system, a split luciferase (eEFR:SBit). We managed to express the CORE:SBit together with the ectodomain of the plant PRR BAK1 which was fused to the LargeBit part of the split NanoLuc protein (eBAK1:LBit). We were able to express both constructs in S. cerevisiae. For a more detailed description see the parts: Part:BBa K3610038 and Part:BBa K3610043.
After transfection of the cells with the two plasmids, we tested whether the split-NanoLuc proteins would be able to interact, which would reconstitute its functionality. We further were interested in the interaction between the EFR and BAK1 ectoomain. EFr and BAK1 naturally interact upon binding of elf18. Should both receptors be expressed properly in the cell membrane of S. cerevisiae, the ligand will be able to bind to the EFR ectodomain. If this is sufficient to drive dimerization of the two receptor in S. cerevisiae, then the NanoLuc parts will be more likely to interact as well, which would lead to a higher amount of functional luciferase proteins in the cell.
To test for dimerization of the luciferase parts and also the two receptors, an assay with a luminometer was performed.
Luminescence assay
Samples with cells which were transfected with the two mentioned plasmids (eBAK1 and eCORE) and samples containing S. cerevisiae cells that were not transfected with any plasmids (UT).
Optical densities (OD600) of all samples were adjusted to 0.34.
For each type of sample, three types of measurements were made:
- 1 µL of deionized water added (no elicitor)
- 1 µL of epitope elf18 added
- 1 µL of epitope csp22 added
Each measurement was done four times with sample size 50µL. To each well 50 µL NanoGlo solution was added (50:1 buffer to furimazine)
The average values of the four measurements are displayed in the plot below.
The results suggest that the receptors do get expressed in the yeast cells. It further is shown that the receptor either drive interaction between the two NanoBit parts or at least do not inhibit this interaction. Luminescence levels were, however, not increased when the bacterial epitope elf18 was present, the elicitor which normally initiates ligand-dependent interaction between BAK1 and EFR. These results suggest that there is no or very little elf18-driven dimerization of the two receptor-ectodomains EFR and BAK1 when they are expressed in S. cerevisiae.
Luminescence Assay 2
The assay was repeated in the a similar manner with the following differences:
- We did not test again for samples which had been transformed with eCORE plasmids. So we only had samples for untransformed yeast cells and for cells transfected with the plasmids containting Part:BBa K3610038 and Part:BBa K3610043.
- The optical density of all samples was adjusted to OD600 = 0.26.
- Measurement started immediately after addition of the bacterial elicitor and the NanoGlo solution.
Our observations of the second luminescence assay matched our expectations after the first one. Luminescence levels were indeed increased when the cells had previously been transformed with our constructs. It was again the case, however, that reconstitution of the NanoLuc protein was not driven by the ligand-dependent interaction of the plant receptors. It seemed, again, to be the case, that addition of bacterial elicitor did in fact decrease the measured luminescent output.
Luminescence Assay 3
The assay was repeated again with the following differences (compared to the first assay):
- We did not test again for samples which had been transformed with eCORE plasmids. So we only had samples for untransformed yeast cells and for cells transfected with the plasmids containting Part:BBa K3610038 and Part:BBa K3610043.
- NanoGlo was added before the measurements were made, the bacterial elicitor was added right before measurement
- Optical density was adjusted to OD600 = 0.5. We waited, however, for almost two hours, during which the cells were kept in TE buffer. During this time, a lot of the cells died, which decresaed the amount of whole living cells as TE supports cell lysis.
- Only one bacterial epitope (elf18) was added.
A third time, we were able to observe that expression of our constructs increased luminescence intensity when the substrate furimazine was added. We, additionally, observed that the increase in luminescence was not initiated or significantly enhanced by presence of the bacterial epitope elf18.
Summary of Luminescence Assays
We conducted for assays with a plate reader of the type Synergy H1. Each time, luminescence was strongly increased when the samples had been transfected with plant PRRs fused to the NanoBits. This has especially been the case when the plant recpetors EFR and its coreceptor BAK1 were used. The increase in luminescence intensity did not seem to be driven by epitope.dependent interaciton of the two receptors. It is possible that, unlinke plants, S. cerevisiae cells lack any mechanisms to prevent unprovoked interaction between the target-receptor EFR and BAK1. Another possible explanation for the results could be that the receptors proteins are not processed the same way as they are in plant cells. For example, in plant cells, the receptor proteins get highly glycosylated. It is possible that this glycosylation influences the structure and the functionality of the receptor proteins.
Conclusion UZürich 2020
The EFR ectodomain was a crucial part of our project. Nothing we set out to do could have worked without this part. We are happy with what we have achieven, even though there is so much more left to discover, much more to try and apply.
We are nevertheless happy, to have used this part in the following ways:
- Demonstration expression by fusing this part to YFP and transfecting S. cerevisiae cells with the construct. We saw that the cells express the receptor and, in addition, some of it gets trafficked to the plasma membrane. Compared to other receptors, the EFR ectodomain has the highest proportion of cells with clear localization.
- Dimerization Assay with eBAK and split-mCherry. Fused to the split-mCherry parts, coexpression of EFR and BAK1 did not reveal increased fluorescence when compared to the control. Interestingly, there seems to be an effect of the bacterial elicitor elf18 on fluorescence intensity.
- Dimerization Assay with BAK1 and split-NanoLuc. Fused to the NanoBit parts, coexpression of this part and BAK1 leads to a luminescent output in the prescence of the NanoLuc substrate furimazine. It does, however, seem that dimerization of the split-NanoLuc is not driven by ligand-dependent interaction of the plant PRRs.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 82
Illegal NheI site found at 1012
Illegal NheI site found at 1930 - 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 1598
- 1000COMPATIBLE WITH RFC[1000]