Difference between revisions of "Part:BBa K3140005"
| (One intermediate revision by the same user not shown) | |||
| Line 9: | Line 9: | ||
VIVID (VVD) is a blue-light sensing photoreceptor from the ascomycete (spore-shooting fungus) ''N. crassa''. It is a member of a family of proteins containing a light-oxygen-voltage-sensing (LOV) domain, which modulate circadian responses to environmental stimuli<ref name="VVD">Schwerdtfeger, C. & Linden, H. VIVID is a flavoprotein and serves as a fungal blue light photoreceptor for photoadaptation. ''EMBO J'' '''22''', 4846-55 (2003).</ref>. Mutation of the highly-conserved LOV domain cystine residue (Cys73) to alanine will convert VVD into a fluoroprotein. In addition, previous work <ref name="VVD36">Zoltowski, B.D. ''et al.'' Conformational switching in the fungal light sensor Vivid. ''Science'' '''316''', 1054-7 (2007).</ref> indicates that truncation of the first 36 amino acids of VVD increases its stability in heterologous systems. Our VVD part incorporates both of these changes. | VIVID (VVD) is a blue-light sensing photoreceptor from the ascomycete (spore-shooting fungus) ''N. crassa''. It is a member of a family of proteins containing a light-oxygen-voltage-sensing (LOV) domain, which modulate circadian responses to environmental stimuli<ref name="VVD">Schwerdtfeger, C. & Linden, H. VIVID is a flavoprotein and serves as a fungal blue light photoreceptor for photoadaptation. ''EMBO J'' '''22''', 4846-55 (2003).</ref>. Mutation of the highly-conserved LOV domain cystine residue (Cys73) to alanine will convert VVD into a fluoroprotein. In addition, previous work <ref name="VVD36">Zoltowski, B.D. ''et al.'' Conformational switching in the fungal light sensor Vivid. ''Science'' '''316''', 1054-7 (2007).</ref> indicates that truncation of the first 36 amino acids of VVD increases its stability in heterologous systems. Our VVD part incorporates both of these changes. | ||
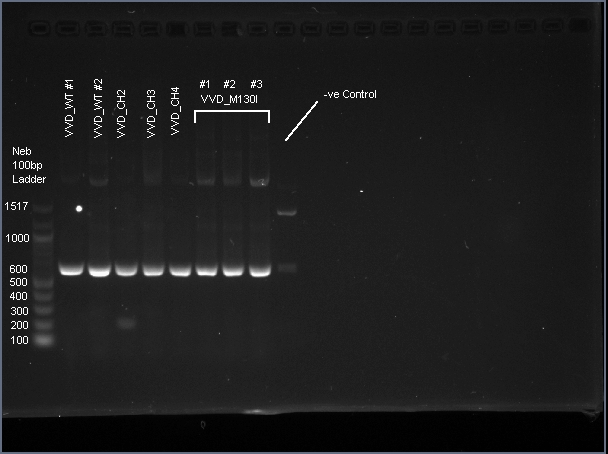
| − | + | VVD36-C73A is a 456 bp sequence, encoding a 151 amino acid peptide (17.2 kDa). We cloned VVD36-C73A, VVD36-C73A-CH1, VVD36-C73A-CH2, VVD36-C73A-CH3, and VVD36-C73A-CH4 into pK18, which were then transformed into ''Escherichia coli'' TOP10 cells. In order to validate the insertion of these genes, we conducted a PCR with primers that were specific to the pK18 backbone, which should a 624 bp product. This was verified with agarose gel electrophoresis, demonstrating had the expected size ('''Fig. 1'''). | |
| + | [[Image:T--Sydney_Australia--VVD_PCR.png|frame|none|'''Fig. 1''': Agarose gel electrophoresis of VVD36-C73A, VVD36-C73A-CH1, VVD36-C73A-CH2, VVD36-C73A-CH3, VVD36-C73A-CH4, and VVD36-C73A-CH4-M130I conducted on 1% agarose gel in 1X TAE for 50 min at 100 V.]] | ||
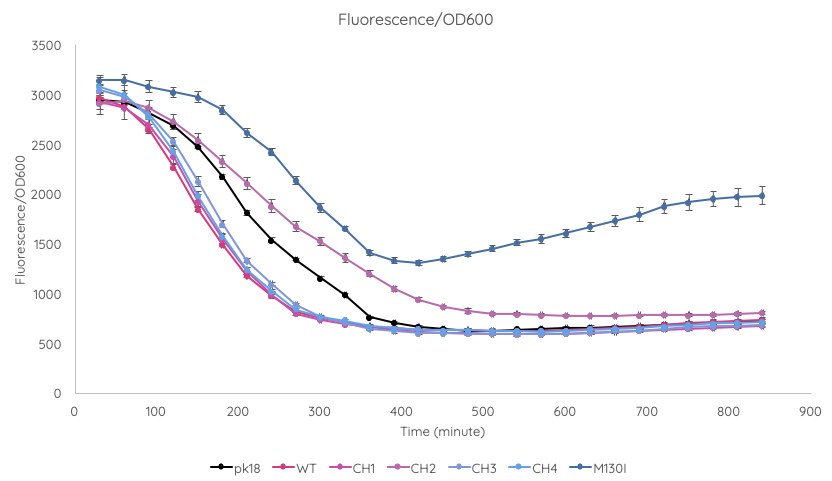
| + | We conducted a fluorescence assay using a TECAN Spark microplate reader ''Escherichia coli'' TOP10 cultures containing VVD36-C73A, VVD36-C73A-CH1, VVD36-C73A-CH2, VVD36-C73A-CH3, VVD36-C73A-CH4, and VVD36-C73A-CH4-M130I in pK18 ('''Fig. 2'''). As shown, VVD36-C73A-CH4-M130I exhibits the greatest fluorescence, followed by VVD36-C73A-CH2. All other VVD38-C73A variants were only slightly more fluorescent than control. | ||
| + | [[Image:T--Sydney_Australia--fluoro_OD_graph.png|frame|none|'''Fig. 2''': Graph of TECAN OD-normalised fluorescence for ''Escherichia coli'' TOP10 cultures containing VVD36-C73A, VVD36-C73A-CH1, VVD36-C73A-CH2, VVD36-C73A-CH3, VVD36-C73A-CH4, and VVD36-C73A-CH4-M130I in pK18.]] | ||
| − | + | <span class='h3bb'>'''Sequence and Features'''</span> | |
| − | <span class='h3bb'>Sequence and Features</span> | + | |
<partinfo>BBa_K3140005 SequenceAndFeatures</partinfo> | <partinfo>BBa_K3140005 SequenceAndFeatures</partinfo> | ||
Latest revision as of 10:00, 21 October 2019
VVD36-C73A - Truncated VIVID fluoroprotein derived from Neurospora crassa
VVD36-C73A is a fluoroprotein derived from the Vivid (VVD) blue-light photoreceptor in Neurospora crassa.
Usage and Biology
VIVID (VVD) is a blue-light sensing photoreceptor from the ascomycete (spore-shooting fungus) N. crassa. It is a member of a family of proteins containing a light-oxygen-voltage-sensing (LOV) domain, which modulate circadian responses to environmental stimuli[1]. Mutation of the highly-conserved LOV domain cystine residue (Cys73) to alanine will convert VVD into a fluoroprotein. In addition, previous work [2] indicates that truncation of the first 36 amino acids of VVD increases its stability in heterologous systems. Our VVD part incorporates both of these changes.
VVD36-C73A is a 456 bp sequence, encoding a 151 amino acid peptide (17.2 kDa). We cloned VVD36-C73A, VVD36-C73A-CH1, VVD36-C73A-CH2, VVD36-C73A-CH3, and VVD36-C73A-CH4 into pK18, which were then transformed into Escherichia coli TOP10 cells. In order to validate the insertion of these genes, we conducted a PCR with primers that were specific to the pK18 backbone, which should a 624 bp product. This was verified with agarose gel electrophoresis, demonstrating had the expected size (Fig. 1).
We conducted a fluorescence assay using a TECAN Spark microplate reader Escherichia coli TOP10 cultures containing VVD36-C73A, VVD36-C73A-CH1, VVD36-C73A-CH2, VVD36-C73A-CH3, VVD36-C73A-CH4, and VVD36-C73A-CH4-M130I in pK18 (Fig. 2). As shown, VVD36-C73A-CH4-M130I exhibits the greatest fluorescence, followed by VVD36-C73A-CH2. All other VVD38-C73A variants were only slightly more fluorescent than control.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12COMPATIBLE WITH RFC[12]
- 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 434
- 1000COMPATIBLE WITH RFC[1000]