Difference between revisions of "Part:BBa K3202039"
| (5 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
| + | __NOTOC__ | ||
| + | <partinfo>BBa_K3202039 short</partinfo> | ||
| + | |||
| + | This composite part is a AraC-Pc-pBAD inducer/promoter system connected with TP901, cds for one of three recombinases used in our project. | ||
| + | |||
| + | In our study, this composite part is designed to test out which one among the two systems, TetR-pTet and AraC-pBAD, is more rigorous in its functions. Also, it is as well aimed to test out the expression of the recombinase in our system. | ||
| + | |||
| + | More about this part is discussed in the recombinase flipping module below. | ||
| + | |||
| + | =<b>Recombinase Flipping Module</b>= | ||
| + | |||
| + | <hr style="height:0px;border:none;border-top:none solid #555555;" /> | ||
| + | |||
| + | === I.Background === | ||
| + | |||
| + | We sought for a structure which could implement permanent memory on changes in our circuit while effectively connects our double bistable system and were inspired by the design of recombinase in Peking 2017. We tested three types of recombinase, Bxb1, TP901, PhiC31 they had in their project in our experiment. | ||
| + | |||
| + | Scientists discovered that phages phiC31 integrase make use of LSTP integrases in mediating phage integration and excision into the bacterial genome between their cognate recognition sites, attB (bacterium) and attP (phage). “By placing these sites in the opposite orientation, LSTP integrases cleave, rotate and rejoin the DNA to invert the region between sites. As shown in figure 1, LSTP integrases catalyze insertion of phage genome (yellow) into the bacterial genome (blue) between attB and attP sites, which form hybrid attL and attR sites. Multicolored arrowheads illustrate the sequence changes that occur during strand exchange, with the core sequence shown in yellow.” | ||
| + | |||
| + | [[File:T--BHSF_ND--Recombinase background 1.svg|600px|thumb|center|alt=Recombinase|Figure 1: LSTP integrases catalyze insertion of phage genome (yellow) into the bacterial genome (blue) between attB and attP sites, which form hybrid attL and attR sites]] | ||
| + | |||
| + | We make use of the property of irreversible recombinase which can implement permanent memory so that before reverting the recombinase the second layer, the DNase, is inhibited in working condition, and after reverting the second layer can be eternally expressed, preventing the bacterium from reviving through other techniques. | ||
| + | |||
| + | === II.Design === | ||
| + | |||
| + | In our team, we include the recombinase in order to "implement the permanent memory of the circuit by irreversibly inverting the orientation of the intervening PR1 according to article <i>Permanent genetic memory with >1-byte capacity</I>." The recombinase is able to flip the DNA in only one direction and thus implement permanent memory. | ||
| + | |||
| + | [[File:T--BHSF ND--Recombinase design 2.svg|600px|thumb|center|alt= Recombinase which can implement permanent memory |Figure 2: the recombinase in order to "implement the permanent memory of the circuit by irreversibly inverting the orientation of the intervening PR1 according to article <i>Permanent genetic memory with >1-byte capacity</I>."]] | ||
| + | |||
| + | |||
| + | When it is applied to our circuit, P1 is inverted in regular working condition where inducer is present, recombinase catalyzes the cleavage of each cognate recognition sites attB and attP, inverting the orientation of P1 and rejoining the circuit at attL and attR respectively. Due to the fact that irreversible recombinase can only flip the DNA in one direction, the inverted P1 which is inhibited in working condition where inducer is present remains inverted even when the recombinase is not expressed as inducer is absent. When the inducer is absent, R1 doesn’t inhibit the expression of the inverted P1 anymore, the toxin in the second layer can be then expressed and conduct suicide. The inversion is a permanent effect on the bistable system, that is to say even if we stimulate the whole system again by adding inducer, P1 will not switch its orientation back. | ||
| + | |||
| + | === III.Experiment & Results === | ||
| + | |||
| + | We used pBAD and PLtetO as promoters to regulate the expression of the three recombinases in order to test the effect flipping over the promoter J23119 (P1) by the three recombinases and to see if there is expression leakage. | ||
| + | |||
| + | Theoretically speaking, when inducer is present, recombinase should be expressed and successfully reverts the promoter P1 in-between, the bacterium will express mRFP, displaying red color; while if no inducer is present, the inhibitory effect the transcriptional factor exerts on P2 will stop the expression of recombinase, P1 fails to reverse, the bacterium will therefore express sfGFP, displaying green color. | ||
| + | |||
| + | In the plating medium with no inducer, the leaked expression of recombinase reverted the promoter J23119, displaying red color. Comparing the three sets of recombinases (Fig. 3&4&5), we found that TP901 has the lowest expression leakage. | ||
| + | |||
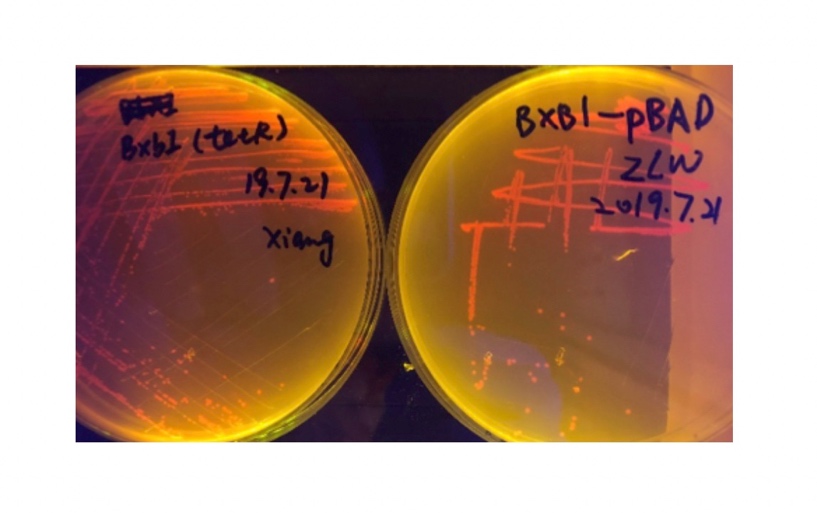
| + | [[File:T--BHSF_ND--recombinase experiment 1.jpeg|271px|thumb|left|alt=Bxb1(TetR) and Bxb1(pBAD) colony|Figure 3: Bxb1(TetR) and Bxb1(pBAD) colony]] | ||
| + | |||
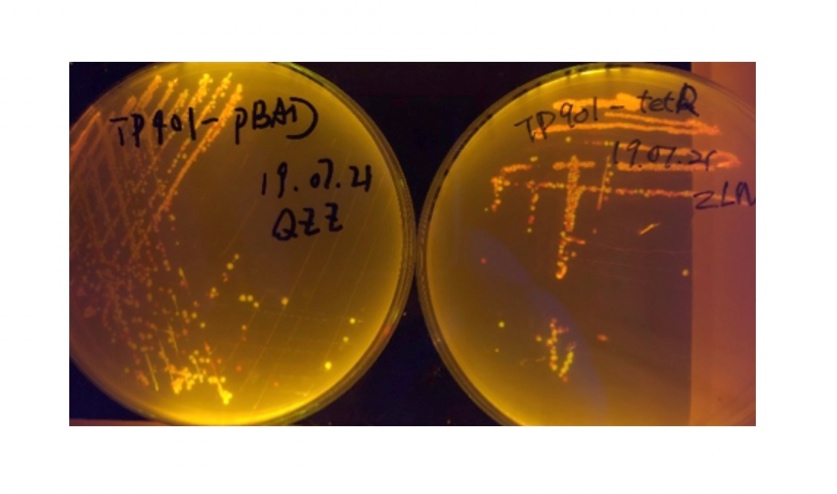
| + | [[File:T--BHSF_ND--recombinase experiment 3.jpeg|291px|thumb|right|alt=TP901(TetR) and TP901(pBAD) colony|Figure 5: TP901(TetR) andTP901(pBAD) colony]] | ||
| + | |||
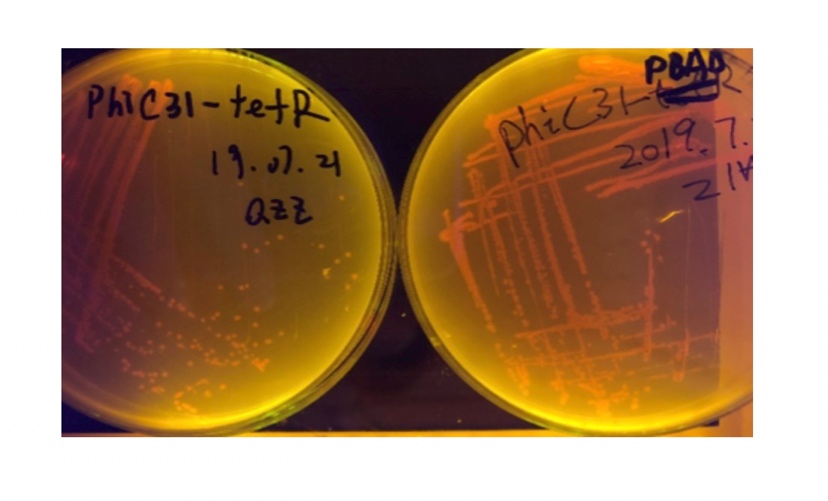
| + | [[File:T--BHSF_ND--recombinase experiment 2.jpeg|288px|thumb|center|alt=PhiC31(TetR) and PhiC31(pBAD) colony|Figure 4: PhiC31(TetR) and PhiC31(pBAD) colony]] | ||
| + | |||
| + | |||
| + | Then we used pBAD to test the recombinases’ working efficiency in inducer’s presence through flow cytometry. Results are quantitatively shown in the diagram below. | ||
| + | |||
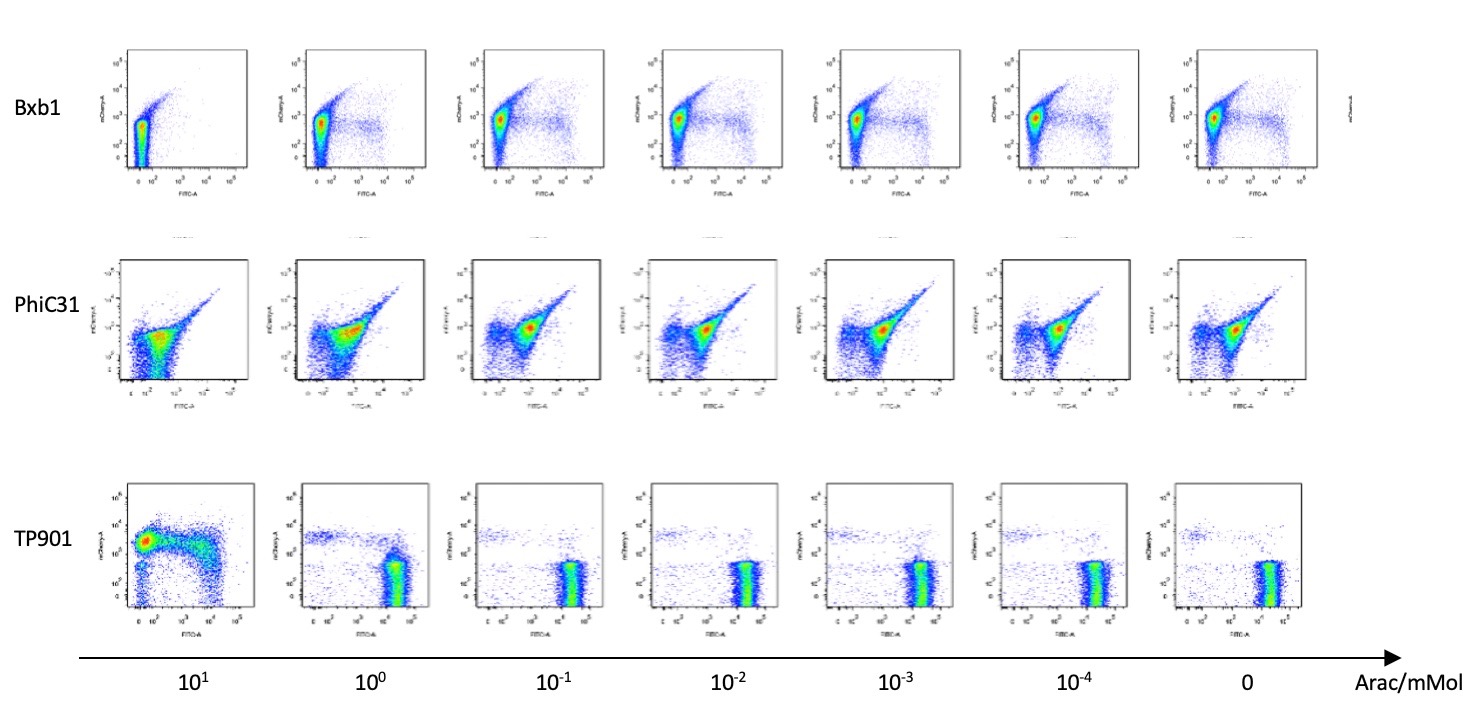
| + | [[File:T--BHSF_ND--Recombinase experiment 4.png|650px|thumb|center|alt=Fluorescence pattern for three types of recombinase|Figure 6: The x-axis of each small square represents the fluorescence of GFP, while the y-axis represents the fluorescence of RFP. All of them show expression leakage in inducer present and absent. (e.g. for PhiC31, according figures in the second row, dispersed high fluorescence of both RFP and GFP in most of the cells show that there is expression leakage of the recombinase both before and after flipping over the recombinase).]] | ||
| + | |||
| + | ===IV.Discussion=== | ||
| + | |||
| + | Our recombinases can effectively invert the in-between promoter under our design and experimental validation. This feature works well as a reactor for changes in the external environment. It permanently changes the sequence of genes and thus the direction of gene expression, which implements our goal of connecting two bistable system and exerting permanent effect on the circuit. | ||
| + | |||
| + | <!-- Add more about the biology of this part here | ||
| + | ===Usage and Biology=== | ||
| + | |||
| + | <!-- --> | ||
| + | <span class='h3bb'>Sequence and Features</span> | ||
| + | <partinfo>BBa_K3202039 SequenceAndFeatures</partinfo> | ||
| + | |||
| + | |||
| + | <!-- Uncomment this to enable Functional Parameter display | ||
| + | ===Functional Parameters=== | ||
| + | <partinfo>BBa_K3202039 parameters</partinfo> | ||
| + | <!-- --> | ||
Latest revision as of 22:36, 20 October 2019
AraC-Pc-pBAD-TP901
This composite part is a AraC-Pc-pBAD inducer/promoter system connected with TP901, cds for one of three recombinases used in our project.
In our study, this composite part is designed to test out which one among the two systems, TetR-pTet and AraC-pBAD, is more rigorous in its functions. Also, it is as well aimed to test out the expression of the recombinase in our system.
More about this part is discussed in the recombinase flipping module below.
Recombinase Flipping Module
I.Background
We sought for a structure which could implement permanent memory on changes in our circuit while effectively connects our double bistable system and were inspired by the design of recombinase in Peking 2017. We tested three types of recombinase, Bxb1, TP901, PhiC31 they had in their project in our experiment.
Scientists discovered that phages phiC31 integrase make use of LSTP integrases in mediating phage integration and excision into the bacterial genome between their cognate recognition sites, attB (bacterium) and attP (phage). “By placing these sites in the opposite orientation, LSTP integrases cleave, rotate and rejoin the DNA to invert the region between sites. As shown in figure 1, LSTP integrases catalyze insertion of phage genome (yellow) into the bacterial genome (blue) between attB and attP sites, which form hybrid attL and attR sites. Multicolored arrowheads illustrate the sequence changes that occur during strand exchange, with the core sequence shown in yellow.”
We make use of the property of irreversible recombinase which can implement permanent memory so that before reverting the recombinase the second layer, the DNase, is inhibited in working condition, and after reverting the second layer can be eternally expressed, preventing the bacterium from reviving through other techniques.
II.Design
In our team, we include the recombinase in order to "implement the permanent memory of the circuit by irreversibly inverting the orientation of the intervening PR1 according to article Permanent genetic memory with >1-byte capacity." The recombinase is able to flip the DNA in only one direction and thus implement permanent memory.
When it is applied to our circuit, P1 is inverted in regular working condition where inducer is present, recombinase catalyzes the cleavage of each cognate recognition sites attB and attP, inverting the orientation of P1 and rejoining the circuit at attL and attR respectively. Due to the fact that irreversible recombinase can only flip the DNA in one direction, the inverted P1 which is inhibited in working condition where inducer is present remains inverted even when the recombinase is not expressed as inducer is absent. When the inducer is absent, R1 doesn’t inhibit the expression of the inverted P1 anymore, the toxin in the second layer can be then expressed and conduct suicide. The inversion is a permanent effect on the bistable system, that is to say even if we stimulate the whole system again by adding inducer, P1 will not switch its orientation back.
III.Experiment & Results
We used pBAD and PLtetO as promoters to regulate the expression of the three recombinases in order to test the effect flipping over the promoter J23119 (P1) by the three recombinases and to see if there is expression leakage.
Theoretically speaking, when inducer is present, recombinase should be expressed and successfully reverts the promoter P1 in-between, the bacterium will express mRFP, displaying red color; while if no inducer is present, the inhibitory effect the transcriptional factor exerts on P2 will stop the expression of recombinase, P1 fails to reverse, the bacterium will therefore express sfGFP, displaying green color.
In the plating medium with no inducer, the leaked expression of recombinase reverted the promoter J23119, displaying red color. Comparing the three sets of recombinases (Fig. 3&4&5), we found that TP901 has the lowest expression leakage.
Then we used pBAD to test the recombinases’ working efficiency in inducer’s presence through flow cytometry. Results are quantitatively shown in the diagram below.
IV.Discussion
Our recombinases can effectively invert the in-between promoter under our design and experimental validation. This feature works well as a reactor for changes in the external environment. It permanently changes the sequence of genes and thus the direction of gene expression, which implements our goal of connecting two bistable system and exerting permanent effect on the circuit.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 2621
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 1144
Illegal BamHI site found at 2347
Illegal XhoI site found at 2260 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 1621
Illegal NgoMIV site found at 1708
Illegal AgeI site found at 979
Illegal AgeI site found at 2571 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 1513
Illegal SapI site found at 961