Difference between revisions of "Part:BBa K2541406"
| (3 intermediate revisions by 2 users not shown) | |||
| Line 2: | Line 2: | ||
__NOTOC__ | __NOTOC__ | ||
<partinfo>BBa_K2541406 short</partinfo> | <partinfo>BBa_K2541406 short</partinfo> | ||
| + | <h5> | ||
| + | <P style="text-indent:2em;"> | ||
| + | A RNA-based thermosensor that can be used for temperature sensitive translational regulation which is based on the change of RNA sencondary structure. The heat-repressible RNA-based thermosensors can repress translation of downstream genes at high temperatures.The composite part is a measurement device, consisting of Anderson promoter (BBa_J23106), heat-repressible RNA-based thermosensor-1 (BBa_K2541101), sfGFP_optimism (BBa_K2541400) and double terminator (BBa_B0010 and BBa_B0012).</p> | ||
| + | </h5> | ||
| − | + | <h1>'''1. Usage and Biology'''</h1> | |
| + | <h5> | ||
| + | <P style="text-indent:2em;"> | ||
| + | RNA-based temperature sensing is common in bacteria that live in fluctuating environments. Most naturally occurring RNA-based thermosensors are heat-inducible, have long sequences, and function by sequestering the Shine–Dalgarno (SD) sequence in a stem-loop structure at low temperatures. | ||
| + | </p> | ||
| + | <P style="text-indent:2em;"> | ||
| + | Here, we designed short, heat-repressible RNA-based thermosensors. These thermosensor sequences contain a single-strand RNase E cleavage (RC) site. RNase E is an enzyme native to ''Escherichia coli'' and many other organisms. Each heat-repressible RNA-based thermosensor sequence was inserted downstream of the transcription start site and upstream of the SD sequence. At high temperatures, the RC is exposed, mRNA was cleaved by RNase E, and expression is ‘OFF’. At low temperatures, the RC binds to the anti-RNase E cleavage site (ARC) and forms a stem-loop. This structure sequesters the RC, and expression is ‘ON’. These short, modular heat-repressible RNA-based thermosensors can be exploited as convenient on/off switches of gene expression. | ||
| + | </p> | ||
| + | <P style="text-indent:2em;"> | ||
| + | Green fluorescent protein (GFP) is commonly used as a reporter gene in intact cells and organisms. This year we select sfGFP (BBa_K2541400), a robustly folded version of GFP, called superfolder GFP as a reporter protein. Compared to superfolder GFP (BBa_I746916), sfGFP_optimism (BBa_K2541400) is BbsI restriction site free, so it can be used in GoldenGate assembly to achieve efficient and rapid assembly of gene fragments. And sfGFP_optimism (BBa_K2541400) has stronger fluorescence intensity than superfolder GFP (BBa_I746916). | ||
| + | </p> | ||
| + | </h5> | ||
| + | <html><center> | ||
| + | <div class="stem-loop hot-repressive"> | ||
| − | < | + | <input id="checked_2" type="checkbox" class="switch" /> |
| − | == | + | |
| − | + | ||
| − | |||
| − | |||
| − | + | <svg class="svg_animate" xmlns="http://www.w3.org/2000/svg" width="50%" viewBox="0 0 599.75 278.7"> | |
| + | <defs> | ||
| + | <style> | ||
| − | + | .switch::before{ | |
| − | + | content:""; | |
| + | display:block; | ||
| + | height:2rem; | ||
| + | width:4rem; | ||
| + | position:absolute; | ||
| + | } | ||
| + | .switch{ | ||
| + | display:block; | ||
| + | height:2rem; | ||
| + | width:4rem; | ||
| + | opacity:1; | ||
| + | border-radius:2rem; | ||
| + | background: #00B8FF; | ||
| + | -webkit-appearance: none; | ||
| + | cursor:pointer; | ||
| + | border:0; | ||
| + | position:relative; | ||
| + | transition:all 0.3s ease; | ||
| + | margin:2rem; | ||
| + | } | ||
| + | .switch::after{ | ||
| + | content:""; | ||
| + | position:absolute; | ||
| + | display:block; | ||
| + | height:2rem; | ||
| + | width:2rem; | ||
| + | border-radius:2rem; | ||
| + | background: white; | ||
| + | box-shadow: 0 0 1rem rgba(0,0,0,0.2); | ||
| + | box-sizing:border-box; | ||
| + | transition:all 0.3s ease; | ||
| + | transform:translateX(0rem); | ||
| + | } | ||
| + | .switch:checked::after{ | ||
| + | transform:translateX(2rem); | ||
| + | } | ||
| + | .switch:checked{ | ||
| + | background: #FFAB63; | ||
| + | } | ||
| − | + | .hot-repressive .a, .b, .c, .f { | |
| − | + | fill: none; | |
| + | stroke-width: 11px; | ||
| + | } | ||
| − | + | .hot-repressive .a { | |
| − | + | stroke: #036; | |
| − | + | } | |
| + | .hot-repressive .a,.hot-repressive .b,.hot-repressive .c,.hot-repressive .f,.hot-repressive .g { | ||
| + | stroke-linecap: round; | ||
| + | stroke-linejoin: round; | ||
| + | } | ||
| − | + | .hot-repressive .b { | |
| − | + | stroke: #060; | |
| + | } | ||
| + | .hot-repressive .c { | ||
| + | stroke: #600; | ||
| + | } | ||
| − | + | .hot-repressive .d,.hot-repressive .e,.hot-repressive .h{ | |
| − | + | font-size: 17px; | |
| + | font-family: Quicksand-Medium, Quicksand; | ||
| + | } | ||
| + | .hot-repressive .d { | ||
| + | fill: #006837; | ||
| + | } | ||
| + | .hot-repressive .e { | ||
| + | fill: #900; | ||
| + | } | ||
| + | |||
| + | .hot-repressive .f { | ||
| + | stroke: #000; | ||
| + | } | ||
| + | |||
| + | .hot-repressive .h { | ||
| + | fill: #930; | ||
| + | } | ||
| + | |||
| + | .hot-repressive .g { | ||
| + | fill: #fff794; | ||
| + | stroke: #fbb03b; | ||
| + | stroke-width: 7px; | ||
| + | } | ||
| + | .hot-repressive .hydrogen-bond{ | ||
| + | opacity:1; | ||
| + | transition: all 0.3s linear 1s; | ||
| + | } | ||
| + | .hot-repressive #RNAse_E{ | ||
| + | transform: translateY(-60px); | ||
| + | opacity:0; | ||
| + | transition: all 1s ease-out 0s; | ||
| + | } | ||
| + | .hot-repressive #ribosome_text{ | ||
| + | opacity:0; | ||
| + | transition: all 0.3s linear 0s; | ||
| + | } | ||
| + | .hot-repressive #text{ | ||
| + | transform: translateY(0px); | ||
| + | transition: all 1s ease-out; | ||
| + | } | ||
| + | .hot-repressive #loop{ | ||
| + | transform: translateY(0px); | ||
| + | transition: all 2s ease-out; | ||
| + | } | ||
| + | .hot-repressive #left,#right{ | ||
| + | stroke-dasharray:130,500; | ||
| + | transition: all 2s ease-out; | ||
| + | } | ||
| + | .hot-repressive #left{ | ||
| + | stroke-dashoffset:0; | ||
| + | stroke:red; | ||
| + | } | ||
| + | .hot-repressive #right{ | ||
| + | stroke-dashoffset:-277; | ||
| + | stroke:green; | ||
| + | } | ||
| + | .hot-repressive #left_s,#right_s{ | ||
| + | stroke-dasharray:60,500; | ||
| + | color:black; | ||
| + | transition: all 2s ease-out; | ||
| + | } | ||
| + | .hot-repressive #left_s{ | ||
| + | stroke-dashoffset:-130; | ||
| + | } | ||
| + | .hot-repressive #right_s{ | ||
| + | stroke-dashoffset:-217; | ||
| + | } | ||
| + | |||
| + | .hot-repressive.svg_checked .hydrogen-bond{ | ||
| + | opacity:0; | ||
| + | transition: all 0.3s linear 0s; | ||
| + | } | ||
| + | .hot-repressive.svg_checked #RNAse_E{ | ||
| + | transform: translateY(0px); | ||
| + | opacity:1; | ||
| + | transition: all 0.5s ease-out 2s; | ||
| + | } | ||
| + | .hot-repressive.svg_checked #ribosome_text{ | ||
| + | opacity:1; | ||
| + | transition: all 0.5s ease-out 2s; | ||
| + | } | ||
| + | .hot-repressive.svg_checked #text{ | ||
| + | transform: translateY(-60px); | ||
| + | } | ||
| + | .hot-repressive.svg_checked #loop{ | ||
| + | transform: translateY(130px); | ||
| + | } | ||
| + | .hot-repressive.svg_checked #left{ | ||
| + | stroke-dashoffset:-135; | ||
| + | } | ||
| + | .hot-repressive.svg_checked #right{ | ||
| + | stroke-dashoffset:-142; | ||
| + | } | ||
| + | .hot-repressive.svg_checked #left_s,#right_s{ | ||
| + | stroke-dasharray:60,500; | ||
| + | } | ||
| + | .hot-repressive.svg_checked #left_s{ | ||
| + | stroke-dashoffset:-265; | ||
| + | } | ||
| + | .hot-repressive.svg_checked #right_s{ | ||
| + | stroke-dashoffset:-82; | ||
| + | } | ||
| + | |||
| + | </style> | ||
| + | </defs> | ||
| + | <g> | ||
| + | <g class="hydrogen-bond"> | ||
| + | <line class="a" x1="284.86" y1="144.21" x2="320.72" y2="144.21"/> | ||
| + | <line class="b" x1="284.86" y1="125.32" x2="320.72" y2="125.32"/> | ||
| + | <line class="c" x1="284.86" y1="163.09" x2="320.72" y2="163.09"/> | ||
| + | <line class="a" x1="284.86" y1="181.98" x2="320.72" y2="181.98"/> | ||
| + | <line class="b" x1="284.86" y1="200.87" x2="320.72" y2="200.87"/> | ||
| + | <line class="c" x1="284.86" y1="219.75" x2="320.72" y2="219.75"/> | ||
| + | </g> | ||
| + | <g id="text"> | ||
| + | <text class="d" transform="translate(429.01 185.45)">RC</text> | ||
| + | <text class="e" transform="translate(123.8 185.45)">anti-RC</text> | ||
| + | |||
| + | </g> | ||
| + | |||
| + | <text id="ribosome_text" class="h" transform="translate(385.48 175.45)">RNAse E</text> | ||
| + | |||
| + | <path id="loop" class="f" d="M327,101.42c0-21.25,19.2-20.39,19.2-52.51,0-24-16.62-43.41-43.42-43.41s-43.41,19.44-43.41,43.41c0,28.79,19.19,29.92,19.19,52.51"/> | ||
| + | |||
| + | <g> | ||
| + | <path id="right_s" class="f" d="M594.25,247.5l-252.2.21a15,15,0,0,1-15-15V101.42"/> | ||
| + | <path id="left_s" class="f" d="M278.57,101.42V232.67a15,15,0,0,1-15.05,15H5.5"/> | ||
| + | </g> | ||
| + | <g> | ||
| + | <path id="right" class="f" d="M594.25,247.5l-252.2.21a15,15,0,0,1-15-15V101.42"/> | ||
| + | <path id="left" class="f" d="M278.57,101.42V232.67a15,15,0,0,1-15.05,15H5.5"/> | ||
| + | </g> | ||
| + | <g id="RNAse_E"> | ||
| + | |||
| + | <path class="g" d="M417,187.26a40.56,40.56,0,0,0-39.11,51.23L417,229.77l-24.18,30.57A40.54,40.54,0,1,0,417,187.26Z"/> | ||
| + | <circle class="g" cx="402.84" cy="214.07" r="1.00"/> | ||
| + | </g> | ||
| + | </g> | ||
| + | <p>Figure 1. Mechanism of heat-repressible RNA-based thermosensors.</p> | ||
| + | </svg> | ||
| + | |||
| + | |||
| + | <script type="text/javascript"> | ||
| + | $(document).ready(function(){ | ||
| + | $("#checked_2").click(function(){ | ||
| + | if($(this).is(':checked')){ | ||
| + | $(".hot-repressive").addClass("svg_checked"); | ||
| + | }else{ | ||
| + | $(".hot-repressive").removeClass("svg_checked"); | ||
| + | } | ||
| + | }); | ||
| + | }); | ||
| + | </script> | ||
| + | </div></center> | ||
| + | </html> | ||
| + | |||
| + | <h1>'''2. Design'''</h1> | ||
| + | <h5> | ||
| + | <P style="text-indent:2em;"> | ||
| + | In order to design heat-repressible RNA-based thermosensores with different melting temperatures, intensity and sensitivity, we change the ARC sequence because the RC sequence is conserved. Three structural parameters come into consideration: stem length, loop size and mismatches or bulges in the stem. | ||
| + | </p> | ||
| + | <P style="text-indent:2em;"> | ||
| + | Stem length is determined by ARC sequence. Adding stem length can optimize heat-repressible RNA-based thermosensors to higher temperature, while decreasing stem length has the opposite effect. The stem length is 13 base parings in K2541101. Loop size can moderate thermosensors melting temperature to an appropriate temperature. In K2541101, the loop sequence is AAUAA. Furthermore, we change base composition in ARC sequence to decrease melting temperature. We make a mismatch in the fifth base by changing it from G to U. And we make a bulge in the ninth base by inserting U. After designing, the theromsensor sequence is predicted by computational methods mFOLD to get its minimum free energy and secondary structure. | ||
| + | </p> | ||
| + | </h5> | ||
| + | [[File:K2541101 f2.png|center|K2541101 f2]] | ||
| + | Figure 2. Design of K2541101. (A) The RNA secondary structure is predicted by mFOLD. (B) ARC sequence, loop sequence, the site of mismatch or bulge in the stem, ΔG and Tm are in the table. | ||
| + | |||
| + | <h1>'''3.Characterization'''</h1> | ||
| + | <h3>3.1 Measurement device</h3> | ||
| + | <h5> | ||
| + | <P style="text-indent:2em;"> | ||
| + | The thermosensor sequence is constructed on the pSB1C3 vector by Golden Gate assembly. The measurement device is composed of Anderson promoter (BBa_J23106), thermosensor (BBa_K2541101), sfGFP_optimism (BBa_K2541400) and double terminator (BBa_B0010 and BBa_B0012). We select a constitutive Anderson promoter J23106 as an appropriate promoter by pre-experiment. The sfGFP_optimism has faster folding speed and higher fluorescence intensity. The double terminator can reduce leakage (Figure 3). We characterized RNA-based thermosensors in ''E.coli'' DH5a. | ||
| + | </p> | ||
| + | </h5> | ||
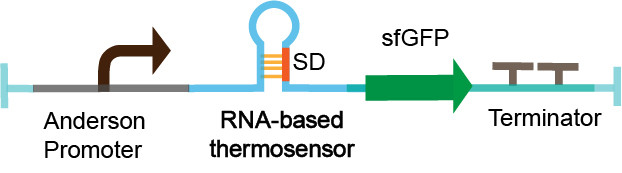
| + | [[File:measurement device 123.png|center|caption]] | ||
| + | <html><center> | ||
| + | Figure 3. The measurement device. | ||
| + | </script></center> | ||
| + | </html> | ||
| + | ---- | ||
| + | |||
| + | <h3>3.2 Measurement results</h3> | ||
| + | <h5> | ||
| + | <P style="text-indent:2em;"> | ||
| + | K2541101, K2541109, K2541119 and K2541101 are four different heat-repressible RNA-based thermosensors. pos.control is positive control. The final normalized fluorescence was calculated as follows: normalized fluorescence = [(Fluorescence/Abs<sub>600</sub>)<sub>TS</sub> - (Fluorescence/Abs<sub>600</sub>)<sub>neg</sub>] / [(Fluorescence/Abs<sub>600</sub>)<sub>pos</sub> - (Fluorescence/Abs<sub>600</sub>)<sub>neg</sub>] ( TS = thermosensor, pos = positive control, and neg = BBa_J364007 ). As shown in figure 4, the fluorescence intensity of K2541101 decreases with elevated temperature. | ||
| + | </p> | ||
| + | </h5> | ||
| + | [[File:K2541101 f4 new.png|center|K2541101 f4 new]] | ||
| + | Figure 4. Characteristics of K2541101. Each set of three bars represents the activity level of a different thermosensor. The bar colors purple, yellow and red represent the temperatures 29, 37 and 42°C, respectively. The height of the bars corresponds to the normalized fluorescence. | ||
| + | |||
| + | <h1>'''4. Collection of heat-repressible RNA-based thermosensors '''</h1> | ||
| + | [[File:RE figure6 new new.png|center|RE figure6 new new]] | ||
| + | Figure 5. Experimental measurements of the collection of heat-repressible RNA-based thermosensors show a variety of responses. (A) Rows represent activity levels of different thermosensors. (B) Each set of three bars represents the activity level of a different thermosensor. The bar colors purple, yellow and red represent the temperatures 29, 37 and 42°C, respectively. The height of the bars corresponds to the normalized fluorescence. | ||
| + | |||
| + | <h1>'''5. Conclusion'''</h1> | ||
| + | <h5> | ||
| + | <P style="text-indent:2em;"> | ||
| + | Our data show that efficient RNA-based thermosensors with different melting temperatures, intensity and sensitivity can be built from a single small RNA stem-loop structure, thus providing useful SynRT toolkit for the regulation of gene expression. | ||
| + | </p> | ||
| + | </h5> | ||
| + | |||
| + | |||
| + | |||
| + | <span class='h3bb'>Sequence and Features</span> | ||
| + | <partinfo>BBa_K2541406 SequenceAndFeatures</partinfo> | ||
<!-- Uncomment this to enable Functional Parameter display | <!-- Uncomment this to enable Functional Parameter display | ||
===Functional Parameters=== | ===Functional Parameters=== | ||
<partinfo>BBa_K2541406 parameters</partinfo> | <partinfo>BBa_K2541406 parameters</partinfo> | ||
<!-- --> | <!-- --> | ||
Latest revision as of 09:08, 16 October 2018
Heat-repressible RNA-based thermosensor measurement device
A RNA-based thermosensor that can be used for temperature sensitive translational regulation which is based on the change of RNA sencondary structure. The heat-repressible RNA-based thermosensors can repress translation of downstream genes at high temperatures.The composite part is a measurement device, consisting of Anderson promoter (BBa_J23106), heat-repressible RNA-based thermosensor-1 (BBa_K2541101), sfGFP_optimism (BBa_K2541400) and double terminator (BBa_B0010 and BBa_B0012).
1. Usage and Biology
RNA-based temperature sensing is common in bacteria that live in fluctuating environments. Most naturally occurring RNA-based thermosensors are heat-inducible, have long sequences, and function by sequestering the Shine–Dalgarno (SD) sequence in a stem-loop structure at low temperatures.
Here, we designed short, heat-repressible RNA-based thermosensors. These thermosensor sequences contain a single-strand RNase E cleavage (RC) site. RNase E is an enzyme native to Escherichia coli and many other organisms. Each heat-repressible RNA-based thermosensor sequence was inserted downstream of the transcription start site and upstream of the SD sequence. At high temperatures, the RC is exposed, mRNA was cleaved by RNase E, and expression is ‘OFF’. At low temperatures, the RC binds to the anti-RNase E cleavage site (ARC) and forms a stem-loop. This structure sequesters the RC, and expression is ‘ON’. These short, modular heat-repressible RNA-based thermosensors can be exploited as convenient on/off switches of gene expression.
Green fluorescent protein (GFP) is commonly used as a reporter gene in intact cells and organisms. This year we select sfGFP (BBa_K2541400), a robustly folded version of GFP, called superfolder GFP as a reporter protein. Compared to superfolder GFP (BBa_I746916), sfGFP_optimism (BBa_K2541400) is BbsI restriction site free, so it can be used in GoldenGate assembly to achieve efficient and rapid assembly of gene fragments. And sfGFP_optimism (BBa_K2541400) has stronger fluorescence intensity than superfolder GFP (BBa_I746916).
2. Design
In order to design heat-repressible RNA-based thermosensores with different melting temperatures, intensity and sensitivity, we change the ARC sequence because the RC sequence is conserved. Three structural parameters come into consideration: stem length, loop size and mismatches or bulges in the stem.
Stem length is determined by ARC sequence. Adding stem length can optimize heat-repressible RNA-based thermosensors to higher temperature, while decreasing stem length has the opposite effect. The stem length is 13 base parings in K2541101. Loop size can moderate thermosensors melting temperature to an appropriate temperature. In K2541101, the loop sequence is AAUAA. Furthermore, we change base composition in ARC sequence to decrease melting temperature. We make a mismatch in the fifth base by changing it from G to U. And we make a bulge in the ninth base by inserting U. After designing, the theromsensor sequence is predicted by computational methods mFOLD to get its minimum free energy and secondary structure.
Figure 2. Design of K2541101. (A) The RNA secondary structure is predicted by mFOLD. (B) ARC sequence, loop sequence, the site of mismatch or bulge in the stem, ΔG and Tm are in the table.
3.Characterization
3.1 Measurement device
The thermosensor sequence is constructed on the pSB1C3 vector by Golden Gate assembly. The measurement device is composed of Anderson promoter (BBa_J23106), thermosensor (BBa_K2541101), sfGFP_optimism (BBa_K2541400) and double terminator (BBa_B0010 and BBa_B0012). We select a constitutive Anderson promoter J23106 as an appropriate promoter by pre-experiment. The sfGFP_optimism has faster folding speed and higher fluorescence intensity. The double terminator can reduce leakage (Figure 3). We characterized RNA-based thermosensors in E.coli DH5a.
3.2 Measurement results
K2541101, K2541109, K2541119 and K2541101 are four different heat-repressible RNA-based thermosensors. pos.control is positive control. The final normalized fluorescence was calculated as follows: normalized fluorescence = [(Fluorescence/Abs600)TS - (Fluorescence/Abs600)neg] / [(Fluorescence/Abs600)pos - (Fluorescence/Abs600)neg] ( TS = thermosensor, pos = positive control, and neg = BBa_J364007 ). As shown in figure 4, the fluorescence intensity of K2541101 decreases with elevated temperature.
Figure 4. Characteristics of K2541101. Each set of three bars represents the activity level of a different thermosensor. The bar colors purple, yellow and red represent the temperatures 29, 37 and 42°C, respectively. The height of the bars corresponds to the normalized fluorescence.
4. Collection of heat-repressible RNA-based thermosensors
Figure 5. Experimental measurements of the collection of heat-repressible RNA-based thermosensors show a variety of responses. (A) Rows represent activity levels of different thermosensors. (B) Each set of three bars represents the activity level of a different thermosensor. The bar colors purple, yellow and red represent the temperatures 29, 37 and 42°C, respectively. The height of the bars corresponds to the normalized fluorescence.
5. Conclusion
Our data show that efficient RNA-based thermosensors with different melting temperatures, intensity and sensitivity can be built from a single small RNA stem-loop structure, thus providing useful SynRT toolkit for the regulation of gene expression.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 7
Illegal NheI site found at 30 - 21INCOMPATIBLE WITH RFC[21]Illegal XhoI site found at 539
- 23COMPATIBLE WITH RFC[23]
- 25COMPATIBLE WITH RFC[25]
- 1000INCOMPATIBLE WITH RFC[1000]Illegal SapI site found at 74