Difference between revisions of "Part:BBa K2753019"
(→Reference) |
|||
| (5 intermediate revisions by 2 users not shown) | |||
| Line 3: | Line 3: | ||
<partinfo>BBa_K2753019 short</partinfo> | <partinfo>BBa_K2753019 short</partinfo> | ||
| − | + | A member of the pTALE stabilized promoter family, characterized and sequenced by team GreatBay_China 2018. | |
| + | |||
| + | ===An Introduction to TALE stabilised promoters=== | ||
| + | <p> Genetic engineering requires elaborative design and sensible proportion of different components. For instance, unbalanced enzyme expression in metabolic engineering would lead to not only shunted flux to the product, but also unnecessary waste of carbon source and host growth inhibition. Plasmids are common engineering tools used to tune gene expression. People often assume that plasmids exist in stable copies, but this is a misleading assumption to make as plasmid copy number is actually subjected to huge variability. Altered chassis strains, new culture medium composition, different temperature, and difference in growth phase all hugely affect plasmid copy number. In our case, although <em>E. coli</em> and yeast are engineered separately, they eventually would be cultured together and probably using very different conditions because compromise is made to meet both requirements of both species. Subsequently, gene expression regulated by such design is easily agitated, leading to potential failure of synthetic device. | ||
| + | </p><br><br> | ||
| + | |||
| + | <html> | ||
| + | <center> | ||
| + | <Figure> | ||
| + | <img width="70%" src="https://static.igem.org/mediawiki/2018/e/ee/T--GreatBay_China--Part_collection_figures.001.jpeg"> | ||
| + | </figure> | ||
| + | |||
| + | </center> | ||
| + | </html> | ||
| + | <br/> | ||
| + | |||
| + | <p> Transcription-activator-like-effector (TALE) stabilised promoters are a type of promoters able to untie gene expression level from gene copy number using an incoherent feedforward loop (iFFL) in which transcription-activator-like effectors (TALEs) function as a perfectly non-cooperative negative regulation. While copy number accretes gene expression, it also elevates the repression to the gene expression, thus has canceled out the effect of copy number on expression level. This design can also eliminate the impact of the location of genes (whether they are placed on plasmids or in the genome, and wherein the genome) on gene expression.</p><br><br> | ||
| + | |||
| + | <p> Having comprehended the incredible capability and potential of TALEsp, we were deeply inspired yet felt sorry that there were only six TALEsp available to use. In metabolic engineering and other areas of synthetic biology, people select promoters very carefully to ensure the most suitable strength is chosen, which seems difficult given so few candidates. Therefore, we expanded the TALEsp library through mutating core promoters of existing TALEsp, and adding TALE binding sites to the classical consensus promoter J23119, we created six more TALEsp, and have added all of twelve TALEsps to the iGEM Registry.</p><br><br> | ||
| + | |||
| + | <html> | ||
| + | <center> | ||
| + | <Figure> | ||
| + | <img width="70%" src="https://static.igem.org/mediawiki/2018/3/3c/T--GreatBay_China--Part_collection_figures.002.jpeg"> | ||
| + | </figure> | ||
| + | |||
| + | </center> | ||
| + | </html> | ||
| + | <br/> | ||
| + | |||
| + | <div> | ||
| + | <div style="display:inline-block; padding-right:5px; vertical-align:top;"> | ||
| + | <h4>TALE1:</h4> | ||
| + | <ul> | ||
| + | <li> pTALE1 sp1 (https://parts.igem.org/Part:BBa_K2753030)</li> | ||
| + | <li> pTALE1 sp2 (https://parts.igem.org/Part:BBa_K2753024)</li> | ||
| + | <li> pTALE1 sp3 (https://parts.igem.org/Part:BBa_K2753025)</li> | ||
| + | <li> pTALE1 sp4 (https://parts.igem.org/Part:BBa_K2753026)</li> | ||
| + | <li> pTALE1 sp5 (https://parts.igem.org/Part:BBa_K2753027)</li> | ||
| + | <li> pTALE1 sp6 (https://parts.igem.org/Part:BBa_K2753029)</li> | ||
| + | </ul> | ||
| + | |||
| + | <h4>TALE2:</h4> | ||
| + | <ul> | ||
| + | <li> pTALE2 sp1(https://parts.igem.org/Part:BBa_K2753018)</li> | ||
| + | <li> pTALE2 sp2(https://parts.igem.org/Part:BBa_K2753019)</li> | ||
| + | <li> pTALE2 sp3(https://parts.igem.org/Part:BBa_K2753020)</li> | ||
| + | <li> pTALE2 sp4(https://parts.igem.org/Part:BBa_K2753021)</li> | ||
| + | <li> pTALE2 sp5(https://parts.igem.org/Part:BBa_K2753022)</li> | ||
| + | <li> <b> pTALE2 sp6(https://parts.igem.org/Part:BBa_K2753023)</b> </li> | ||
| + | </ul> | ||
| + | |||
| + | </div> | ||
| + | |||
| + | <div style="display:inline-block; padding-right:50px;"> | ||
| + | <html> | ||
| + | <center> | ||
| + | <h5> Table. 1 The source of all TALEsp from the library</h5> | ||
| + | <Figure> | ||
| + | <img width="70%" src="https://static.igem.org/mediawiki/2018/4/4f/T--GreatBay_China--design_table_1.png"> | ||
| + | </figure> | ||
| + | </center> | ||
| + | </html> | ||
| + | <br/> | ||
| + | </div> | ||
| + | </div> | ||
| + | <br><br><br> | ||
| + | |||
| + | === Biobrick Design=== | ||
| + | <p> All promoters are placed on the standard biobrick assembly compatible backbone pSB1C3, with a RBS (BBa_B0034) reporter gene sfGFP (BBa_K1679038) downstream of the promoters in between the restriction sites of SpeI and PstI. So teams can change reporter genes as they wish.</p> | ||
| + | |||
| + | <p> For TALE stabilised promoters repressed by the same TALE, the only difference between them is the sequence of the core promoter which the TALE binds to. By replacing this promoter, new TALE stabilised promoters could then be easily created. For making the process more convenient, we designed this composite part in which a genetic circuit encoding for amilCP, a blue pigment, replaces the core promoter sequence, and downstream are a RBS, a coding sequence of sfGFP, and a terminator. Two restriction sites of BsaI are put on the two ends of the amilCP circuit. By adding the same digestion sites on the promoter to be assembled, the amilCP circuit could be substituted by Golden Gate. The blue colour of amilCP and green colour of sfGFP provide a rapid distinguishment of the successful construct. If the amilCP circuit is not replaced, the colony grown would be blue as amilCP is being expressed. But if a promoter sequence substitutes amilCP circuit, sfGFP would be expressed instead of amilCP, making the colony fluorescent and easy to recognise. </p> | ||
| + | |||
| + | <p> Except for TALE1sp1 and TALE2sp1 which were synthesized de novo by Genscript, we assembled the rest of TALEsps by first obtaining core promoters using SOE PCR then replacing the amilCP circuit with them.</p><br> | ||
| + | |||
| + | <html> | ||
| + | <center> | ||
| + | <Figure> | ||
| + | <img width="70%" src="https://static.igem.org/mediawiki/2018/d/d3/T--GreatBay_China--Part_collection_figures.003.jpeg"> | ||
| + | </figure> | ||
| + | |||
| + | </center> | ||
| + | </html> | ||
| + | <br/> | ||
| + | |||
| + | |||
| + | <br> | ||
| + | ===Characterization=== | ||
| + | <p> In order to verify the stabilization effect of the pTALE promoter family, we assembled all promoter members onto three backbones:</p><br> | ||
| + | <ul> | ||
| + | <li>pUC20: ~500</li> | ||
| + | <li>pR6K: ~15</li> | ||
| + | <li>pSC101: ~1</li> | ||
| + | </ul> | ||
| + | <br> | ||
| + | <p>An identical sfGFP is placed downstream of all promoters as a reporter gene. Three constitutive promoters: J23119, J23101, J23105, with the same sfGFP downstream were characterized along with the pTALE promoters as a reference of promoter strength. The green fluoresce was measured by flow cytometry. (see methods in our protocols). | ||
| + | </p> | ||
| + | |||
| + | <p>To decide whether pTALE can perform as expected in metabolic pathways, maintaining the pre-set ratio of production enzymes for ensuring optimal flux to the product, we tested pTALE1 sp1 and pTALE2 sp1 by comparison to pTac, a frequently-used IPTG inducible promoter, using an geraniol synthesis operon (https://parts.igem.org/Part:BBa_K2753015) containing a GPPS (https://parts.igem.org/Part:BBa_K2753002) and a GES (https://parts.igem.org/Part:BBa_K2753003). Similar to the characterisation of green fluoresce characterization, three backbones: pUC20, pR6K, pSC101 are used.</p><br> | ||
| + | |||
| + | <html> | ||
| + | <center> | ||
| + | <Figure> | ||
| + | <img width="70%" src="https://static.igem.org/mediawiki/2018/b/b4/T--GreatBay_China--Part_collection_figures.004.jpeg"> | ||
| + | </figure> | ||
| + | </center> | ||
| + | </html> | ||
| + | <br/> | ||
| + | |||
| + | |||
| + | |||
| + | <p>The results indicate that TALEsp are able to buffer against the change in plasmid copy number and the location of genes in the organism, maintaining a reasonably constant fluoresce level except for TALE1sp3. The fluorescence driven the TALE1sp3 shows a positive correlation with copy number.</p> | ||
| + | |||
| + | |||
| + | |||
| + | <p>Moreover, to decide whether TALEsp can perform as expected in metabolic pathways, maintaining the pre-set ratio of production enzymes for ensuring optimal flux to the product, we tested TALE1 sp1 and TALE2 sp1 by comparison to pTac, a frequently-used IPTG inducible promoter, using an geraniol synthesis operon (https://parts.igem.org/Part:BBa_K2753015) containing a GPPS (https://parts.igem.org/Part:BBa_K2753002) and a GES (https://parts.igem.org/Part:BBa_K2753003). They are co-expressed with a pMVA-only vector to provide a sufficient and stable supply of precursors IPP and DMAPP. Similar to the characterisation of green fluoresce characterization, three backbones: pUC20, pR6K, pSC101 are used. </p> | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <html> | ||
| + | <center> | ||
| + | <Figure> | ||
| + | <img width="70%" src="https://static.igem.org/mediawiki/2018/9/9d/T--GreatBay_China--Part_collection_figures.005.jpeg"> | ||
| + | </figure> | ||
| + | |||
| + | </center> | ||
| + | </html> | ||
| + | <br/> | ||
| + | |||
| + | |||
| + | <p>With pTac promoters, geraniol yield increased with the copy number of the vector, showing positively correlated relation. However, when TALE stabilized promoter (TALEsp) was used, higher the copy number of the vector, lower the production of geraniol, being the opposite of pTac. And the stronger TALEsp promoter, TALE1sp1 gave generally reduced yield compared to its weaker counterpart TALE2sp1. TALE2 sp6 whose strength measure with fluorescence is about half of the strength of TALE2 sp1 produced the lowest titre among the four promoters tested. Interestingly, the graphs of the TALE stabilised promoters appeared to be seemingly parallel to each other.</p> | ||
| + | |||
| + | <p>We surmised that the yield of geraniol was affected by two factors: the expression level of enzymes and the cellular burden. As for enzyme expression, there exists an optimal gene expression level that produces just enough enzyme to metabolize all the substrate. The yield would be the greatest at this level. But if lower than this level, production would increase with enzyme expression since there is a surplus of substrates. And if the enzyme expression is higher, the more enzymes now becomes a cellular burden as it no longer contributes to more product.</p> | ||
| + | |||
| + | <p>In the case of pTac, it fits into the scenario when the gene expression is lower than the optimum: higher copy produced more enzymes to catalyze geraniol synthesis reaction. As for TALE stabilized promoters, the expression level of enzymes would remain unchanged regardless of copy number due to the stabilization nature of TALEsp (Check our Design [http://2018.igem.org/Team:GreatBay_China/Design]). So in low copy vector where the cellular burden isn’t significant, the product yield is only related to the strength of the promoter, explaining why TALE2 sp1 has better performance than both TALE1 sp1 and TALE2 sp6 as it’s closer to the optimum. But when the copy number increases, the expression level is unaffected while the cellular burden rises sharply because much more TALE would be made to stabilize expression, leaving less energy available for geraniol synthesis. And it in turns answers why a negative trend of production is shown when regulated by TALEsp.</p> | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| − | |||
| − | |||
<!-- --> | <!-- --> | ||
| Line 17: | Line 157: | ||
<partinfo>BBa_K2753019 parameters</partinfo> | <partinfo>BBa_K2753019 parameters</partinfo> | ||
<!-- --> | <!-- --> | ||
| + | |||
| + | |||
| + | ==Contribution: Greatbay_SZ 2019== | ||
| + | <div> | ||
| + | <b>Group: GreatBay_SZ 2019</b> | ||
| + | <br> | ||
| + | <b>Author: Xinyou Chang</b> | ||
| + | <br> | ||
| + | <b>Introduction & Design: Xinyou Chang</b> | ||
| + | <br> | ||
| + | <p><br><br> | ||
| + | <b>Documentation:</b> | ||
| + | <br> | ||
| + | This part contains deoxyviolacein expression genes and TALE stabilized promoter sp2. | ||
| + | The complete pathway of biosynthesis of deoxyviolacein, a light purple pigment | ||
| + | Despite simply laying a complete database of spideroin, GreatBay_SZ this year also approached one major industry where spider silk held great potential: the cloth industry. Identifying the current chemical compound pollution during dying process as well as the damage brought by chemical fiber itself, we realized that the typically non-environmentally friendly material of cloth manufacture can be replaced by spider silk, as thread to weaved the cloth, and natural dyes, as pigments that granted cloth color. | ||
| + | |||
| + | ===Usage in biology=== | ||
| + | [[File: vioviofig1.png|center|500px]] | ||
| + | The pigment deoxyviolacein is produced from L-tryptophan in ''E.coli'' via a pathway involving four enzymes VioA, VioB, VioE, VioC, only vioD is excluded through the pathway. GreatBay_SZ 2019 borrow from team SHSBNU_China to acquire part thsR- BBa_K274003, which include one thiosulfate sensor and vioABDE which synthesize proviolacein. As shown in the graph below, different arrangement of VioA-E can synthesize pigment with deviate color. Our team wants to substitute VioD to vioC that gives as a light pink/purple pigment | ||
| + | |||
| + | Aiming for stable and high efficient production through deoxyviolacein pathway, using the original thiosulfate sensor cannot yield us high concentration of pigment. Instead, we look into GreatBay_China_2018’s stabilized promoter BBa_K2753019, Talesp2 from the pTale family. Transcription-activator-like-effector (TALE) stabilised promoters are a type of promoters able to untie gene expression level from gene copy number using an incoherent feed forward loop (iFFL) in which transcription-activator-like effectors (TALEs) function as a perfectly non-cooperative negative regulation. While copy number accretes gene expression, it also elevates the repression to the gene expression, thus has canceled out the effect of copy number on expression level. Thus using Talesp2, we comprehend that it can yield high quantitative results | ||
| + | |||
| + | Indigo, though without much significant medical usage, is the most classic ancient dye uniquely capable of producing the signature tones in blue denim. The current production of indigo not only release hazardous chemical compound, but also requires an excess, usually toxic, reducing agent to reduce the insoluble indigo into soluble leucoindigo. Replacement of reducing agent were identified in the past, yet none established as fast nor cost effective. We therefore researched an alternative indigo production pathway. With L-tryptophan, indole is formed by enzyme tnaA in E.coli. In the presence of indole and oxygen, FMO catalyzes the addition of a hydroxyl group to indole generating the intermediate indoxyl that can spontaneously oxidize to form indigo. | ||
| + | |||
| + | ===Characterization=== | ||
| + | |||
| + | [[File: vioviofig2.png|center|500px]] | ||
| + | |||
| + | To achieve over production of both pigments, we utilized the stabilized promoter BBa_K2753019 from GreatBay_China(2018), transcription-activator-like-effector (TALE) stabilised promoters that untie gene expression level from gene copy number. We designed to place TALEsp2 promoter on standard pSC101 backbones. Combining the pathway with promoter, we hence characterized and obtained pigments by constructing part TALEsp2-VioABEC for deoxyviolacein through BBa_K2753019, BBa_K726015, and a new basic part BBa_K3264008, a gene-transcript of VioC. Meanwhile, we also constructed a new composite part TALEsp2-tnaA-FMO for indigo. | ||
| + | |||
| + | Deoxyviolacein has long be identified as secondary metabolity actively against pathogenic bacteria like ''Pseudomonas aeruginosa'' and ''Staphylococcus aureus'', and leukemia, lung cancer, human uveal melanoma, and lymphoma cells[10,11]. It also served other purpose like natural pigments. The importance of violacein urged us to search for over production of both metabolities. The hidden pathway for production is encoded by the VioABCDE operon. Bio-synthesis starts from L-tryptophan, converted into protodeoxyviolaceinic acid by VioA, VioB and VioE enzymes, and then into deoxyviolacein is therefore produced with the activation of VioC gene[10]. | ||
| + | All promoters are placed on the standard biobrick assembly compatible backbone pSC101. We reassembled plasmid thsR- BBa_K274003 by PCR vioAB, VioE and talesp2, only VioC was synthesized de novo by Genescript. All parts were assembled using Gibson Assembly. | ||
| + | |||
| + | After we acquire pure extract deoxyviolacein pigment, we test it dying properties upon our spider silk sample. As documented as the process below, the pigment can be well soaked into fibers to give it light purple color. | ||
| + | |||
| + | The influence of talesp2 on the production and accumulation of deoxyviolacein is remarkable, with the highest yield round 90~100mg/L which nearly matched the standard of pure extracts deoxyviolacein. It can be concluded that talesp2 has positive influences on deoxyviolacein’s metabolic reaction, which it stabilized towards an optimal level of gene expression that produce just enough enzyme to metabolize the substrate. | ||
| + | After we acquire pure extract deoxyviolacein pigment, we test it dying properties upon our spider silk sample. As documented as the process below, the pigment can be well soaked into fibers to give it light purple color. | ||
| + | |||
| + | Pigments were obtained by extracting pigments after 42 hours of shake-flask incubation (without iptg) using solvent ethanol for violacein, and DMSO for indigo respectively. | ||
| + | Through calculation based on standard indigo product and OD measurement of oh, 6h, 18h, 24h, 30h, 42h (peak of production), we are able to construct a yield versus time curve. Through that, we concluded our deoxyviolacein yields 85.81±9.09mg/L maximum and indigo yields 6.97±0.44mg/L maximum. | ||
| + | |||
| + | '''Testing Dying Properties on Spider silk Fibers''' | ||
| + | [[File: vioviofig3.png|center|500px]] | ||
| + | |||
| + | ===Reference=== | ||
| + | [1]Rodrigues, André L., et al. “Systems Metabolic Engineering of Escherichia Coli for Production of the Antitumor Drugs Violacein and Deoxyviolacein.” Metabolic Engineering, vol. 20, 2013, pp. 29–41., doi:10.1016/j.ymben.2013.08.004. | ||
| + | [2]Wen, Rui, et al. “Molecular Cloning and Analysis of the Full-Length Aciniform Spidroin Gene from Araneus Ventricosus.” International Journal of Biological Macromolecules, vol. 117, 2018, pp. 1352–1360., doi:10.1016/j.ijbiomac.2017.12.090. | ||
Latest revision as of 01:21, 22 October 2019
TALE2 sp2
A member of the pTALE stabilized promoter family, characterized and sequenced by team GreatBay_China 2018.
An Introduction to TALE stabilised promoters
Genetic engineering requires elaborative design and sensible proportion of different components. For instance, unbalanced enzyme expression in metabolic engineering would lead to not only shunted flux to the product, but also unnecessary waste of carbon source and host growth inhibition. Plasmids are common engineering tools used to tune gene expression. People often assume that plasmids exist in stable copies, but this is a misleading assumption to make as plasmid copy number is actually subjected to huge variability. Altered chassis strains, new culture medium composition, different temperature, and difference in growth phase all hugely affect plasmid copy number. In our case, although E. coli and yeast are engineered separately, they eventually would be cultured together and probably using very different conditions because compromise is made to meet both requirements of both species. Subsequently, gene expression regulated by such design is easily agitated, leading to potential failure of synthetic device.

Transcription-activator-like-effector (TALE) stabilised promoters are a type of promoters able to untie gene expression level from gene copy number using an incoherent feedforward loop (iFFL) in which transcription-activator-like effectors (TALEs) function as a perfectly non-cooperative negative regulation. While copy number accretes gene expression, it also elevates the repression to the gene expression, thus has canceled out the effect of copy number on expression level. This design can also eliminate the impact of the location of genes (whether they are placed on plasmids or in the genome, and wherein the genome) on gene expression.
Having comprehended the incredible capability and potential of TALEsp, we were deeply inspired yet felt sorry that there were only six TALEsp available to use. In metabolic engineering and other areas of synthetic biology, people select promoters very carefully to ensure the most suitable strength is chosen, which seems difficult given so few candidates. Therefore, we expanded the TALEsp library through mutating core promoters of existing TALEsp, and adding TALE binding sites to the classical consensus promoter J23119, we created six more TALEsp, and have added all of twelve TALEsps to the iGEM Registry.

TALE1:
- pTALE1 sp1 (https://parts.igem.org/Part:BBa_K2753030)
- pTALE1 sp2 (https://parts.igem.org/Part:BBa_K2753024)
- pTALE1 sp3 (https://parts.igem.org/Part:BBa_K2753025)
- pTALE1 sp4 (https://parts.igem.org/Part:BBa_K2753026)
- pTALE1 sp5 (https://parts.igem.org/Part:BBa_K2753027)
- pTALE1 sp6 (https://parts.igem.org/Part:BBa_K2753029)
TALE2:
- pTALE2 sp1(https://parts.igem.org/Part:BBa_K2753018)
- pTALE2 sp2(https://parts.igem.org/Part:BBa_K2753019)
- pTALE2 sp3(https://parts.igem.org/Part:BBa_K2753020)
- pTALE2 sp4(https://parts.igem.org/Part:BBa_K2753021)
- pTALE2 sp5(https://parts.igem.org/Part:BBa_K2753022)
- pTALE2 sp6(https://parts.igem.org/Part:BBa_K2753023)
Table. 1 The source of all TALEsp from the library

Biobrick Design
All promoters are placed on the standard biobrick assembly compatible backbone pSB1C3, with a RBS (BBa_B0034) reporter gene sfGFP (BBa_K1679038) downstream of the promoters in between the restriction sites of SpeI and PstI. So teams can change reporter genes as they wish.
For TALE stabilised promoters repressed by the same TALE, the only difference between them is the sequence of the core promoter which the TALE binds to. By replacing this promoter, new TALE stabilised promoters could then be easily created. For making the process more convenient, we designed this composite part in which a genetic circuit encoding for amilCP, a blue pigment, replaces the core promoter sequence, and downstream are a RBS, a coding sequence of sfGFP, and a terminator. Two restriction sites of BsaI are put on the two ends of the amilCP circuit. By adding the same digestion sites on the promoter to be assembled, the amilCP circuit could be substituted by Golden Gate. The blue colour of amilCP and green colour of sfGFP provide a rapid distinguishment of the successful construct. If the amilCP circuit is not replaced, the colony grown would be blue as amilCP is being expressed. But if a promoter sequence substitutes amilCP circuit, sfGFP would be expressed instead of amilCP, making the colony fluorescent and easy to recognise.
Except for TALE1sp1 and TALE2sp1 which were synthesized de novo by Genscript, we assembled the rest of TALEsps by first obtaining core promoters using SOE PCR then replacing the amilCP circuit with them.

Characterization
In order to verify the stabilization effect of the pTALE promoter family, we assembled all promoter members onto three backbones:
- pUC20: ~500
- pR6K: ~15
- pSC101: ~1
An identical sfGFP is placed downstream of all promoters as a reporter gene. Three constitutive promoters: J23119, J23101, J23105, with the same sfGFP downstream were characterized along with the pTALE promoters as a reference of promoter strength. The green fluoresce was measured by flow cytometry. (see methods in our protocols).
To decide whether pTALE can perform as expected in metabolic pathways, maintaining the pre-set ratio of production enzymes for ensuring optimal flux to the product, we tested pTALE1 sp1 and pTALE2 sp1 by comparison to pTac, a frequently-used IPTG inducible promoter, using an geraniol synthesis operon (https://parts.igem.org/Part:BBa_K2753015) containing a GPPS (https://parts.igem.org/Part:BBa_K2753002) and a GES (https://parts.igem.org/Part:BBa_K2753003). Similar to the characterisation of green fluoresce characterization, three backbones: pUC20, pR6K, pSC101 are used.

The results indicate that TALEsp are able to buffer against the change in plasmid copy number and the location of genes in the organism, maintaining a reasonably constant fluoresce level except for TALE1sp3. The fluorescence driven the TALE1sp3 shows a positive correlation with copy number.
Moreover, to decide whether TALEsp can perform as expected in metabolic pathways, maintaining the pre-set ratio of production enzymes for ensuring optimal flux to the product, we tested TALE1 sp1 and TALE2 sp1 by comparison to pTac, a frequently-used IPTG inducible promoter, using an geraniol synthesis operon (https://parts.igem.org/Part:BBa_K2753015) containing a GPPS (https://parts.igem.org/Part:BBa_K2753002) and a GES (https://parts.igem.org/Part:BBa_K2753003). They are co-expressed with a pMVA-only vector to provide a sufficient and stable supply of precursors IPP and DMAPP. Similar to the characterisation of green fluoresce characterization, three backbones: pUC20, pR6K, pSC101 are used.
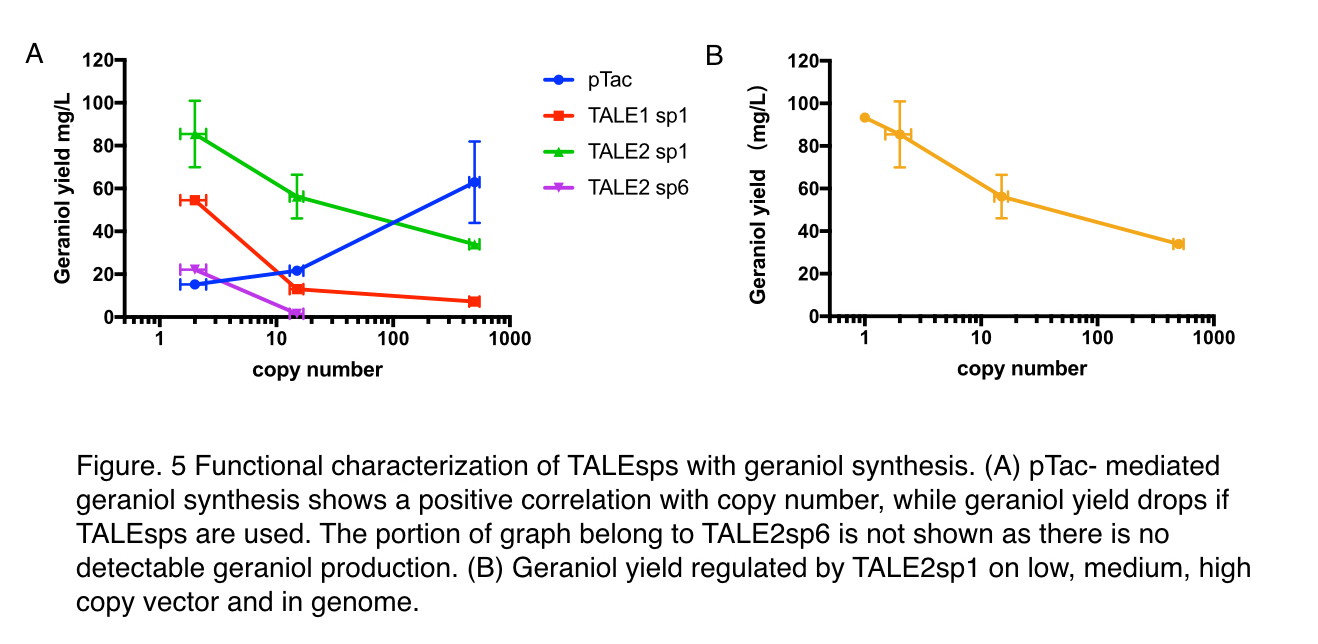
With pTac promoters, geraniol yield increased with the copy number of the vector, showing positively correlated relation. However, when TALE stabilized promoter (TALEsp) was used, higher the copy number of the vector, lower the production of geraniol, being the opposite of pTac. And the stronger TALEsp promoter, TALE1sp1 gave generally reduced yield compared to its weaker counterpart TALE2sp1. TALE2 sp6 whose strength measure with fluorescence is about half of the strength of TALE2 sp1 produced the lowest titre among the four promoters tested. Interestingly, the graphs of the TALE stabilised promoters appeared to be seemingly parallel to each other.
We surmised that the yield of geraniol was affected by two factors: the expression level of enzymes and the cellular burden. As for enzyme expression, there exists an optimal gene expression level that produces just enough enzyme to metabolize all the substrate. The yield would be the greatest at this level. But if lower than this level, production would increase with enzyme expression since there is a surplus of substrates. And if the enzyme expression is higher, the more enzymes now becomes a cellular burden as it no longer contributes to more product.
In the case of pTac, it fits into the scenario when the gene expression is lower than the optimum: higher copy produced more enzymes to catalyze geraniol synthesis reaction. As for TALE stabilized promoters, the expression level of enzymes would remain unchanged regardless of copy number due to the stabilization nature of TALEsp (Check our Design [http://2018.igem.org/Team:GreatBay_China/Design]). So in low copy vector where the cellular burden isn’t significant, the product yield is only related to the strength of the promoter, explaining why TALE2 sp1 has better performance than both TALE1 sp1 and TALE2 sp6 as it’s closer to the optimum. But when the copy number increases, the expression level is unaffected while the cellular burden rises sharply because much more TALE would be made to stabilize expression, leaving less energy available for geraniol synthesis. And it in turns answers why a negative trend of production is shown when regulated by TALEsp.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 1513
Illegal NheI site found at 1921 - 21INCOMPATIBLE WITH RFC[21]Illegal BglII site found at 113
Illegal XhoI site found at 2152
Illegal XhoI site found at 2692 - 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 747
Illegal AgeI site found at 849
Illegal AgeI site found at 1767
Illegal AgeI site found at 2277 - 1000COMPATIBLE WITH RFC[1000]
Contribution: Greatbay_SZ 2019
Group: GreatBay_SZ 2019
Author: Xinyou Chang
Introduction & Design: Xinyou Chang
Documentation:
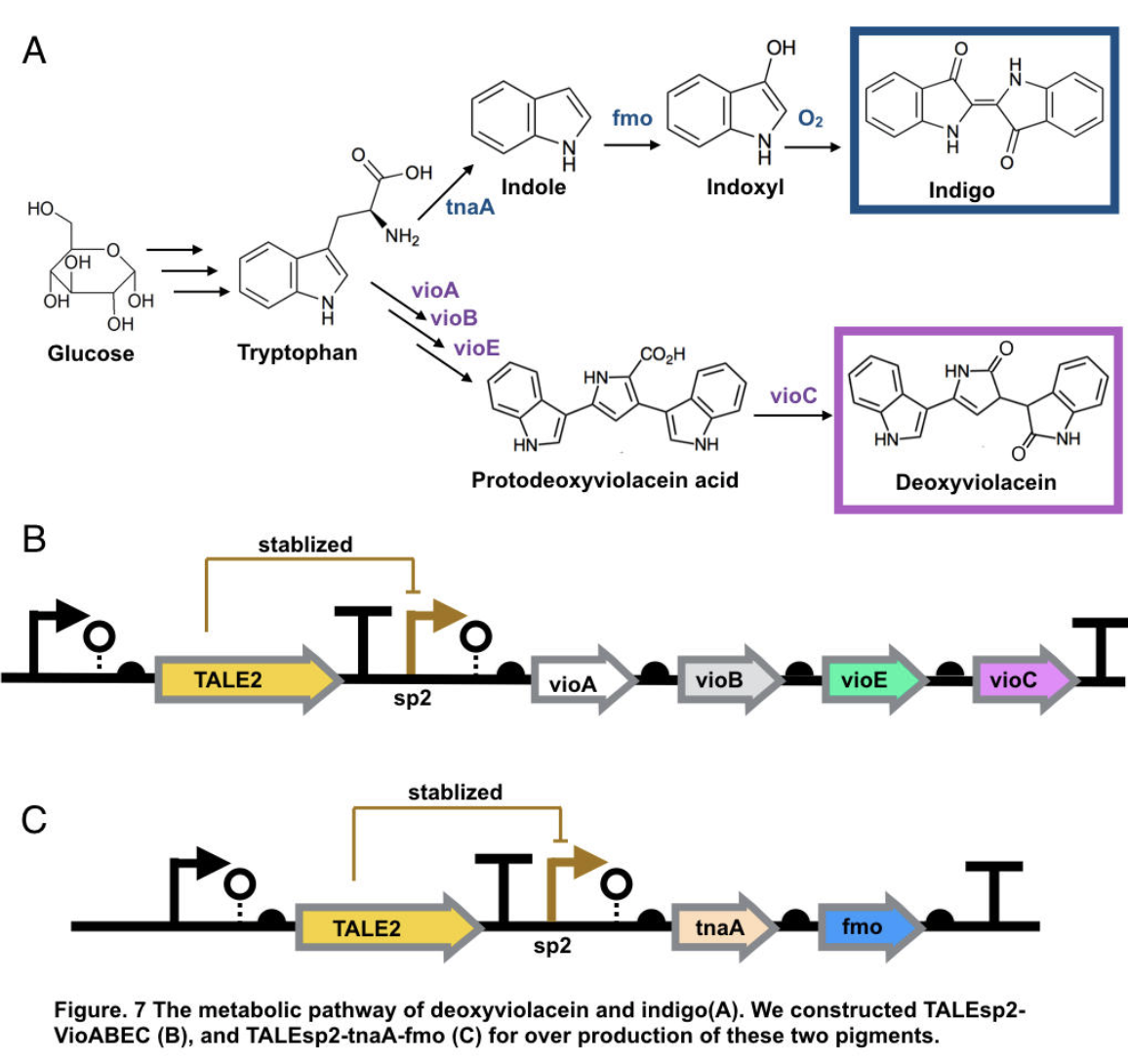
This part contains deoxyviolacein expression genes and TALE stabilized promoter sp2.
The complete pathway of biosynthesis of deoxyviolacein, a light purple pigment
Despite simply laying a complete database of spideroin, GreatBay_SZ this year also approached one major industry where spider silk held great potential: the cloth industry. Identifying the current chemical compound pollution during dying process as well as the damage brought by chemical fiber itself, we realized that the typically non-environmentally friendly material of cloth manufacture can be replaced by spider silk, as thread to weaved the cloth, and natural dyes, as pigments that granted cloth color.
Usage in biology
The pigment deoxyviolacein is produced from L-tryptophan in E.coli via a pathway involving four enzymes VioA, VioB, VioE, VioC, only vioD is excluded through the pathway. GreatBay_SZ 2019 borrow from team SHSBNU_China to acquire part thsR- BBa_K274003, which include one thiosulfate sensor and vioABDE which synthesize proviolacein. As shown in the graph below, different arrangement of VioA-E can synthesize pigment with deviate color. Our team wants to substitute VioD to vioC that gives as a light pink/purple pigment
Aiming for stable and high efficient production through deoxyviolacein pathway, using the original thiosulfate sensor cannot yield us high concentration of pigment. Instead, we look into GreatBay_China_2018’s stabilized promoter BBa_K2753019, Talesp2 from the pTale family. Transcription-activator-like-effector (TALE) stabilised promoters are a type of promoters able to untie gene expression level from gene copy number using an incoherent feed forward loop (iFFL) in which transcription-activator-like effectors (TALEs) function as a perfectly non-cooperative negative regulation. While copy number accretes gene expression, it also elevates the repression to the gene expression, thus has canceled out the effect of copy number on expression level. Thus using Talesp2, we comprehend that it can yield high quantitative results
Indigo, though without much significant medical usage, is the most classic ancient dye uniquely capable of producing the signature tones in blue denim. The current production of indigo not only release hazardous chemical compound, but also requires an excess, usually toxic, reducing agent to reduce the insoluble indigo into soluble leucoindigo. Replacement of reducing agent were identified in the past, yet none established as fast nor cost effective. We therefore researched an alternative indigo production pathway. With L-tryptophan, indole is formed by enzyme tnaA in E.coli. In the presence of indole and oxygen, FMO catalyzes the addition of a hydroxyl group to indole generating the intermediate indoxyl that can spontaneously oxidize to form indigo.
Characterization
To achieve over production of both pigments, we utilized the stabilized promoter BBa_K2753019 from GreatBay_China(2018), transcription-activator-like-effector (TALE) stabilised promoters that untie gene expression level from gene copy number. We designed to place TALEsp2 promoter on standard pSC101 backbones. Combining the pathway with promoter, we hence characterized and obtained pigments by constructing part TALEsp2-VioABEC for deoxyviolacein through BBa_K2753019, BBa_K726015, and a new basic part BBa_K3264008, a gene-transcript of VioC. Meanwhile, we also constructed a new composite part TALEsp2-tnaA-FMO for indigo.
Deoxyviolacein has long be identified as secondary metabolity actively against pathogenic bacteria like Pseudomonas aeruginosa and Staphylococcus aureus, and leukemia, lung cancer, human uveal melanoma, and lymphoma cells[10,11]. It also served other purpose like natural pigments. The importance of violacein urged us to search for over production of both metabolities. The hidden pathway for production is encoded by the VioABCDE operon. Bio-synthesis starts from L-tryptophan, converted into protodeoxyviolaceinic acid by VioA, VioB and VioE enzymes, and then into deoxyviolacein is therefore produced with the activation of VioC gene[10]. All promoters are placed on the standard biobrick assembly compatible backbone pSC101. We reassembled plasmid thsR- BBa_K274003 by PCR vioAB, VioE and talesp2, only VioC was synthesized de novo by Genescript. All parts were assembled using Gibson Assembly.
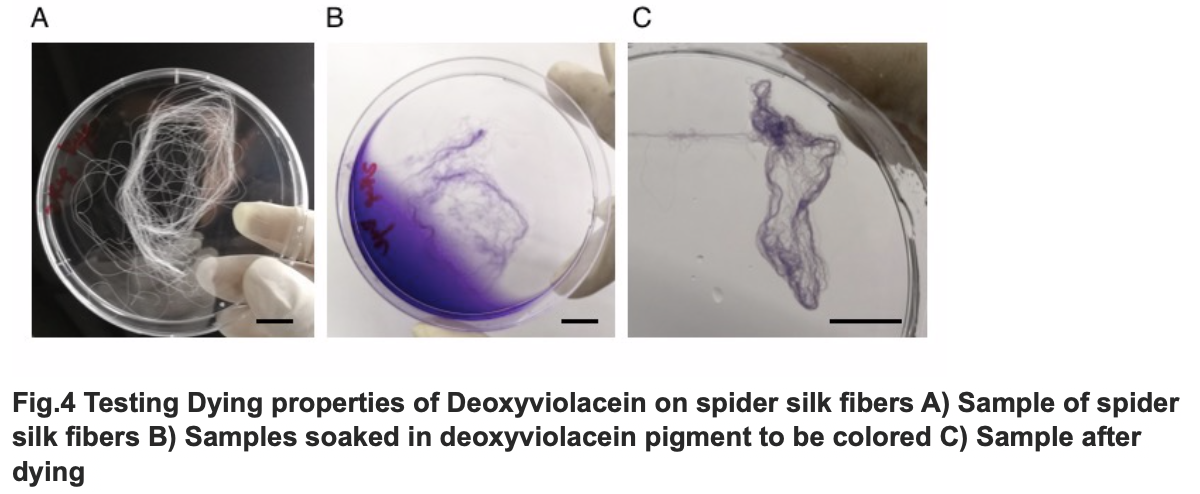
After we acquire pure extract deoxyviolacein pigment, we test it dying properties upon our spider silk sample. As documented as the process below, the pigment can be well soaked into fibers to give it light purple color.
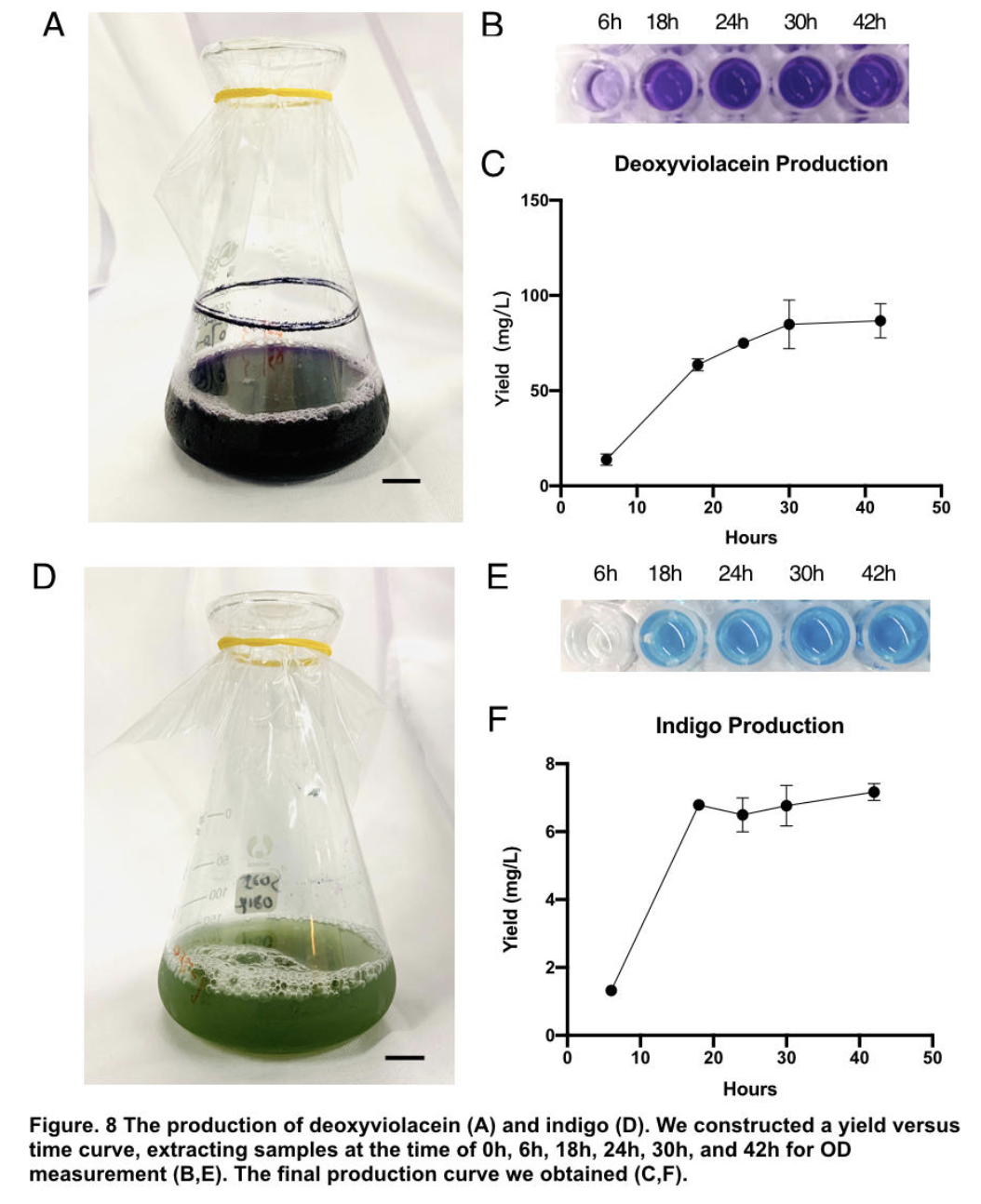
The influence of talesp2 on the production and accumulation of deoxyviolacein is remarkable, with the highest yield round 90~100mg/L which nearly matched the standard of pure extracts deoxyviolacein. It can be concluded that talesp2 has positive influences on deoxyviolacein’s metabolic reaction, which it stabilized towards an optimal level of gene expression that produce just enough enzyme to metabolize the substrate. After we acquire pure extract deoxyviolacein pigment, we test it dying properties upon our spider silk sample. As documented as the process below, the pigment can be well soaked into fibers to give it light purple color.
Pigments were obtained by extracting pigments after 42 hours of shake-flask incubation (without iptg) using solvent ethanol for violacein, and DMSO for indigo respectively. Through calculation based on standard indigo product and OD measurement of oh, 6h, 18h, 24h, 30h, 42h (peak of production), we are able to construct a yield versus time curve. Through that, we concluded our deoxyviolacein yields 85.81±9.09mg/L maximum and indigo yields 6.97±0.44mg/L maximum.
Testing Dying Properties on Spider silk Fibers
Reference
[1]Rodrigues, André L., et al. “Systems Metabolic Engineering of Escherichia Coli for Production of the Antitumor Drugs Violacein and Deoxyviolacein.” Metabolic Engineering, vol. 20, 2013, pp. 29–41., doi:10.1016/j.ymben.2013.08.004.
[2]Wen, Rui, et al. “Molecular Cloning and Analysis of the Full-Length Aciniform Spidroin Gene from Araneus Ventricosus.” International Journal of Biological Macromolecules, vol. 117, 2018, pp. 1352–1360., doi:10.1016/j.ijbiomac.2017.12.090.