Difference between revisions of "Part:BBa K2300003"
Ari edmonds (Talk | contribs) (→References) |
Ari edmonds (Talk | contribs) (→Protein information) |
||
| (31 intermediate revisions by 2 users not shown) | |||
| Line 2: | Line 2: | ||
__NOTOC__ | __NOTOC__ | ||
<partinfo>BBa_K2300003 short</partinfo> | <partinfo>BBa_K2300003 short</partinfo> | ||
| − | |||
| − | |||
<!-- Add more about the biology of this part here | <!-- Add more about the biology of this part here | ||
| Line 11: | Line 9: | ||
<span class='h3bb'>Sequence and Features</span> | <span class='h3bb'>Sequence and Features</span> | ||
<partinfo>BBa_K2300003 SequenceAndFeatures</partinfo> | <partinfo>BBa_K2300003 SequenceAndFeatures</partinfo> | ||
| + | |||
| + | ===Overview=== | ||
| + | |||
| + | The original HydEF page (BBa_K1998012) (https://parts.igem.org/Part:BBa_K1998012) had the wrong sequence so we have created an updated page. The HydEF protein contains two unique domains that are homologous to two distinct prokaryotic proteins, HydE and HydF, which are found exclusively in organisms containing [FeFe] hydrogenase (Mulder et al., 2010). This part is the first gene of the [FeFe] hydrogenase maturation enzyme complex. | ||
| + | |||
| + | All genes within this plasmid are sequences obtained from <i>Chlamydomonas reinhardtii</i> and codon optimised to be expressed in <i>Escherichia coli</i>. | ||
===Biology & Literature=== | ===Biology & Literature=== | ||
| + | |||
| + | HydEF in the majority of [FeFe] containing organisms exists as two separately transcribed proteins (HydE and HydF). However in several algal species the two genes are fused (Böck et al., 2006). HydE (along with HydG) is a radical SAM (S-adenosyl methionine) enzyme which binds [4Fe-4S] clusters in the biosynthesis of the H-cluster (Mulder et al., 2010). HydF contains an N-terminal GTPase domain which is responsible for the P loop motif and FeS binding (Mulder et al., 2010). Both these genes have been shown to be essential for the maturation of the HydA hydrogenase (King et al., 2006). This is due to their role in the creation of the H-cluster. | ||
===Part Verification=== | ===Part Verification=== | ||
| + | The entire Hydrogen Gas Producing Gene Cluster (BBa_K2300001) was sequenced and confirmed once it had been ligated together. This included the HydEF part. | ||
| + | |||
| + | To confirm the efficacy of the ribosome binding sites in our parts the Macquarie iGEM team of 2017 used the Salis Lab Ribosome Binding Site calculator from Penn State University. The results from this were that our ribosome binding site had a translation initiation rate of 1324.3. | ||
| + | |||
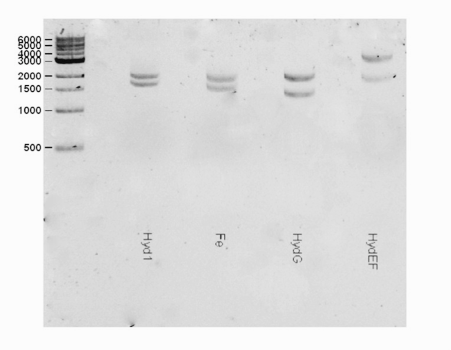
| + | <html><center><img src="https://static.igem.org/mediawiki/2016/b/bd/T--Macquarie_Australia--HYDShowGel.png" " width="50%" height="35%"></center> </html> | ||
| + | <center> <b>Fig 1.</b> A gel consisting of EcorI/PstI for the hydEF part. The expected band for the hydEF (~3600 bp) part was observed.</center> | ||
===Protein information=== | ===Protein information=== | ||
| − | + | HydEF<br> | |
Mass: 121.95 kDa<br> | Mass: 121.95 kDa<br> | ||
Sequence: <br> | Sequence: <br> | ||
| Line 37: | Line 49: | ||
===References=== | ===References=== | ||
| − | Mulder, D.W., Shepard, E.M., Meuser, J.E., Joshi, N., King, P.W., Posewitz, M.C., Broderick, J.B. and Peters, J.W., 2011. Insights into [FeFe]-hydrogenase structure, mechanism, and maturation. Structure, 19(8), pp.1038-1052. | + | Böck, A., King, P.W., Blokesch, M. and Posewitz, M.C., 2006. Maturation of hydrogenases. <i>Advances in Microbial Physiology</i>, 51, pp.1-225. |
| + | |||
| + | |||
| + | King, P.W., Posewitz, M.C., Ghirardi, M.L. and Seibert, M., 2006. Functional studies of [FeFe] hydrogenase maturation in an <i>Escherichia coli</i> biosynthetic system. <i>Journal of Bacteriology</i>, 188(6), pp.2163-2172. | ||
| + | |||
| + | |||
| + | Mulder, D.W., Boyd, E.S., Sarma, R., Lange, R.K., Endrizzi, J.A., Broderick, J.B. and Peters, J.W., 2010. Stepwise [FeFe]-hydrogenase H-cluster assembly revealed in the structure of HydA [Dgr] EFG. Nature, 465(7295), pp.248-251. | ||
| + | |||
| + | |||
| + | Mulder, D.W., Shepard, E.M., Meuser, J.E., Joshi, N., King, P.W., Posewitz, M.C., Broderick, J.B. and Peters, J.W., 2011. Insights into [FeFe]-hydrogenase structure, mechanism, and maturation. <i>Structure</i>, 19(8), pp.1038-1052. | ||
| + | |||
| − | Posewitz, M.C., King, P.W., Smolinski, S.L., Zhang, L., Seibert, M. and Ghirardi, M.L., 2004. Discovery of two novel radical S-adenosylmethionine proteins required for the assembly of an active [Fe] hydrogenase. Journal of Biological Chemistry, 279(24), pp.25711-25720. | + | Posewitz, M.C., King, P.W., Smolinski, S.L., Zhang, L., Seibert, M. and Ghirardi, M.L., 2004. Discovery of two novel radical S-adenosylmethionine proteins required for the assembly of an active [Fe] hydrogenase. <i>Journal of Biological Chemistry</i>, 279(24), pp.25711-25720. |
Vancouver | Vancouver | ||
Latest revision as of 00:03, 2 November 2017
HydEF (Updated 2017)
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 1849
Illegal NheI site found at 2059
Illegal NheI site found at 2503
Illegal NotI site found at 253 - 21COMPATIBLE WITH RFC[21]
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal NgoMIV site found at 901
Illegal AgeI site found at 1816
Illegal AgeI site found at 3459 - 1000INCOMPATIBLE WITH RFC[1000]Illegal BsaI site found at 2684
Illegal BsaI.rc site found at 304
Illegal BsaI.rc site found at 416
Overview
The original HydEF page (BBa_K1998012) (https://parts.igem.org/Part:BBa_K1998012) had the wrong sequence so we have created an updated page. The HydEF protein contains two unique domains that are homologous to two distinct prokaryotic proteins, HydE and HydF, which are found exclusively in organisms containing [FeFe] hydrogenase (Mulder et al., 2010). This part is the first gene of the [FeFe] hydrogenase maturation enzyme complex.
All genes within this plasmid are sequences obtained from Chlamydomonas reinhardtii and codon optimised to be expressed in Escherichia coli.
Biology & Literature
HydEF in the majority of [FeFe] containing organisms exists as two separately transcribed proteins (HydE and HydF). However in several algal species the two genes are fused (Böck et al., 2006). HydE (along with HydG) is a radical SAM (S-adenosyl methionine) enzyme which binds [4Fe-4S] clusters in the biosynthesis of the H-cluster (Mulder et al., 2010). HydF contains an N-terminal GTPase domain which is responsible for the P loop motif and FeS binding (Mulder et al., 2010). Both these genes have been shown to be essential for the maturation of the HydA hydrogenase (King et al., 2006). This is due to their role in the creation of the H-cluster.
Part Verification
The entire Hydrogen Gas Producing Gene Cluster (BBa_K2300001) was sequenced and confirmed once it had been ligated together. This included the HydEF part.
To confirm the efficacy of the ribosome binding sites in our parts the Macquarie iGEM team of 2017 used the Salis Lab Ribosome Binding Site calculator from Penn State University. The results from this were that our ribosome binding site had a translation initiation rate of 1324.3.
Protein information
HydEF
Mass: 121.95 kDa
Sequence:
MAHSLSAHSRQAGDRKLGAGAASSRPSCPSRRIVRVAAHASASKATPDVPVDDLPPAHARAAVAAANRRARAMASAEAAAETLGDFLGLGKGGLSP
GATANLDREQVLGVLEAVWRRGDLNLERALYSHANAVTNKYCGGGVYYRGLVEFSNICQNDCSYCGIRNNQKEVWRYTMPVEEVVEVAKWALENGI
RNIMLQGGELKTEQRLAYLEACVRAIREETTQLDLEMRARAASTTTAEAAASAQADAEAKRGEPELGVVVSLSVGELPMEQYERLFRAGARRYLIRIET
SNPDLYAALHPEPMSWHARVECLRNLKKAGYMLGTGVMVGLPGQTLHDLAGDVMFFRDIKADMIGMGPFITQPGTPATDKWTALYPNANKNSHMK
SMFDLTTAMNALVRITMGNVNISATTALQAIIPTGREIALERGANVVMPILTPTQYRESYQLYEGKPCITDTAVQCRRCLDMRLHSVGKTSAAGVWGDPA
SFLHPIVGVPVPHDLSSPALAAAASADFHEVGAGPWNPIRLERLVEVPDRYPDPDNHGRKKAGAGKGGKAHDSHDDGDHDDHHHHHGAAPAGAAA
GKGTGAAAIGGGAGASRQRVAGAAAASARLCAGARRAGRVVASPLRPAAACRGVAVKAAAAAAGEDAGAGTSGVGSNIVTSPGIASTTAHGVPRINI
GVFGVMNAGKSTLVNALAQQEACIVDSTPGTTADVKTVLLELHALGPAKLLDTAGLDEVGGLGDKKRRKALNTLKECDVAVLVVDTDTAAAAIKSGRLA
EALEWESKVMEQAHKYNVSPVLLLNVKSRGLPEAQAASMLEAVAGMLDPSKQIPRMSLDLASTPLHERSTITSAFVKEGAVRSSRYGAPLPGCLPRW
SLGRNARLLMVIPMDAETPGGRLLRPQAQVMEEAIRHWATVLSVRLDLDAARGKLGPEACEMERQRFDGVIAMMERNDGPTLVVTDSQAIDVVHPW
TLDRSSGRPLVPITTFSIAMAYQQNGGRLDPFVEGLEALETLQDGDRVLISEACNHNRITSACNDIGMVQIPNKLEAALGGKKLQIEHAFGREFPELESG
GMDGLKLAIHCGGCMIDAQKMQQRMKDLHEAGVPVTNYGVFFSWAAWPDALRRALEPWGVEPPVGTPATPAAAPATAASGV
References
Böck, A., King, P.W., Blokesch, M. and Posewitz, M.C., 2006. Maturation of hydrogenases. Advances in Microbial Physiology, 51, pp.1-225.
King, P.W., Posewitz, M.C., Ghirardi, M.L. and Seibert, M., 2006. Functional studies of [FeFe] hydrogenase maturation in an Escherichia coli biosynthetic system. Journal of Bacteriology, 188(6), pp.2163-2172.
Mulder, D.W., Boyd, E.S., Sarma, R., Lange, R.K., Endrizzi, J.A., Broderick, J.B. and Peters, J.W., 2010. Stepwise [FeFe]-hydrogenase H-cluster assembly revealed in the structure of HydA [Dgr] EFG. Nature, 465(7295), pp.248-251.
Mulder, D.W., Shepard, E.M., Meuser, J.E., Joshi, N., King, P.W., Posewitz, M.C., Broderick, J.B. and Peters, J.W., 2011. Insights into [FeFe]-hydrogenase structure, mechanism, and maturation. Structure, 19(8), pp.1038-1052.
Posewitz, M.C., King, P.W., Smolinski, S.L., Zhang, L., Seibert, M. and Ghirardi, M.L., 2004. Discovery of two novel radical S-adenosylmethionine proteins required for the assembly of an active [Fe] hydrogenase. Journal of Biological Chemistry, 279(24), pp.25711-25720.
Vancouver
