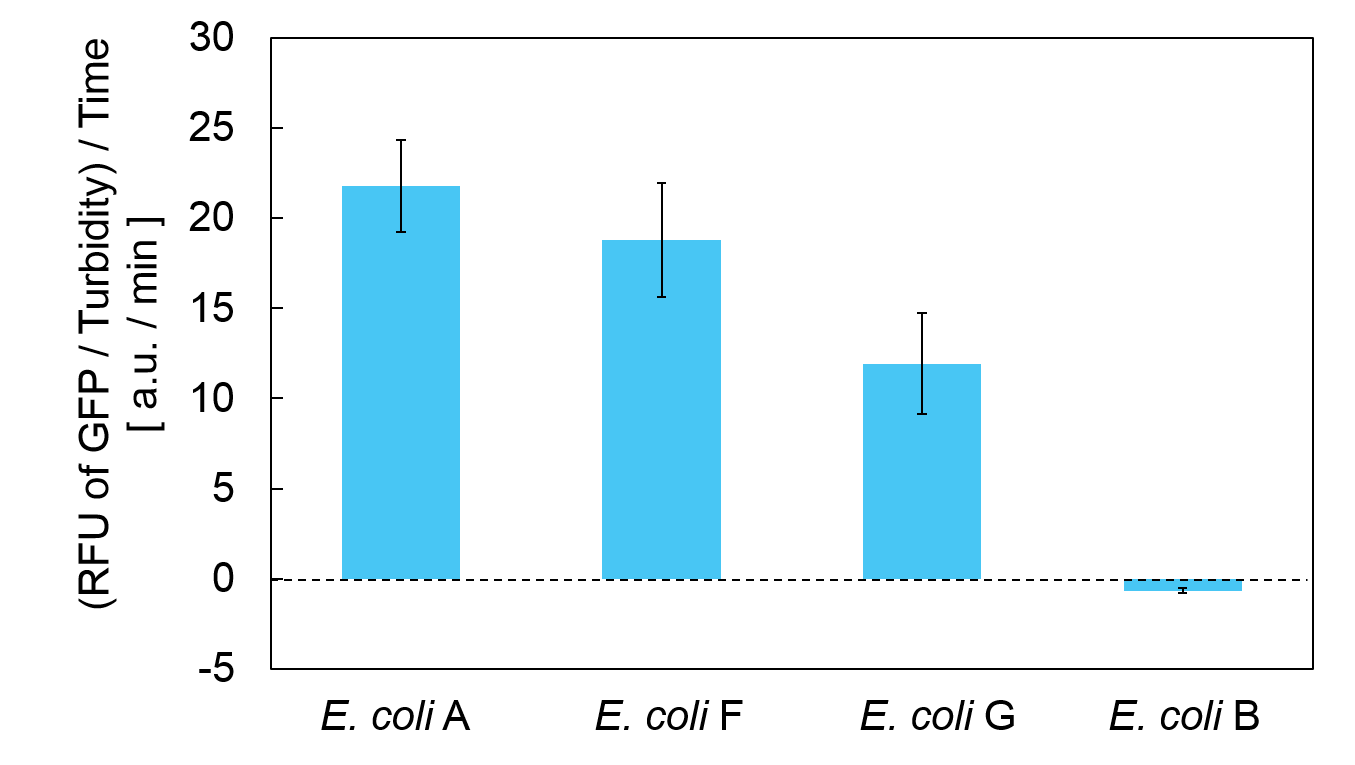Difference between revisions of "Part:BBa K1949101"
(→Ⅰ.Adjustment of MazF Expression) |
|||
| (16 intermediate revisions by 3 users not shown) | |||
| Line 2: | Line 2: | ||
__NOTOC__ | __NOTOC__ | ||
<partinfo>BBa_K1949101 short</partinfo> | <partinfo>BBa_K1949101 short</partinfo> | ||
| − | <span style="margin-left: 10px;">We designed parts(<partinfo> | + | <span style="margin-left: 10px;">We designed parts(<partinfo>BBa_K1949100</partinfo>, <partinfo>BBa_K1949101</partinfo>, <partinfo>BBa_K1949102</partinfo>, <partinfo>BBa_K1949103</partinfo>, <partinfo>BBa_K1949104</partinfo>) to characterize <i>mazE</i>(<partinfo>BBa_K1096001</partinfo>) and <i>mazF</i>(<partinfo>BBa_K1096002</partinfo>). |
===Characterize === | ===Characterize === | ||
<span style="margin-left: 10px;">We performed experiment to work our final genetic circuits and characterized <i>mazE</i> and <i>mazF</i>.<br> | <span style="margin-left: 10px;">We performed experiment to work our final genetic circuits and characterized <i>mazE</i> and <i>mazF</i>.<br> | ||
| − | This experiment consists of the four parts below.<br> | + | <span style="margin-left: 10px;">This experiment consists of the four parts below.<br> |
Ⅰ.Adjustment of MazF Expression | Ⅰ.Adjustment of MazF Expression | ||
| Line 15: | Line 15: | ||
Ⅲ.<i>mazEF</i> System Assay ~Go & Stop~ | Ⅲ.<i>mazEF</i> System Assay ~Go & Stop~ | ||
| − | Ⅳ. | + | Ⅳ.<i>mazEF</i> System Assay on the LB Agar Plate (Queen's Caprice) |
| − | ====Ⅰ.Adjustment of | + | ====Ⅰ.Adjustment of <i>mazF</i> Expression==== |
| − | <span style="margin-left: 10px;">In order to control the | + | <span style="margin-left: 10px;">In order to control cell growth as we desire using the <i>mazEF</i> system, it is necessary to adjust the expression level of <i>mazF</i>. It has been reported that MazF has very strong ability to inhibit cell growth and that <i>mazE</i> expression can not recover it when <i>mazF</i> is expressed at high level[1]. Therefore, we here explored the relationship between concentration of the expression inducer for <i>mazF</i> (arabinose in this experiment) and expression level of it; such information is important for operating our final genetic circuits properly.<br> |
<span style="margin-left: 10px;">We constructed | <span style="margin-left: 10px;">We constructed | ||
| − | + | <i>E. coli</i> A: Pcon - <i>rbs</i> - <i>gfp</i> (pSB6A1), Plac - <i>rbs</i> (pSB3K3) | |
| − | + | <i>E. coli</i> B: PBAD - <i>rbs - mazF - tt</i> - Pcon - <i>rbs - gfp</i> (pSB6A1), Plac - <i>rbs</i> (pSB3K3). | |
=====result===== | =====result===== | ||
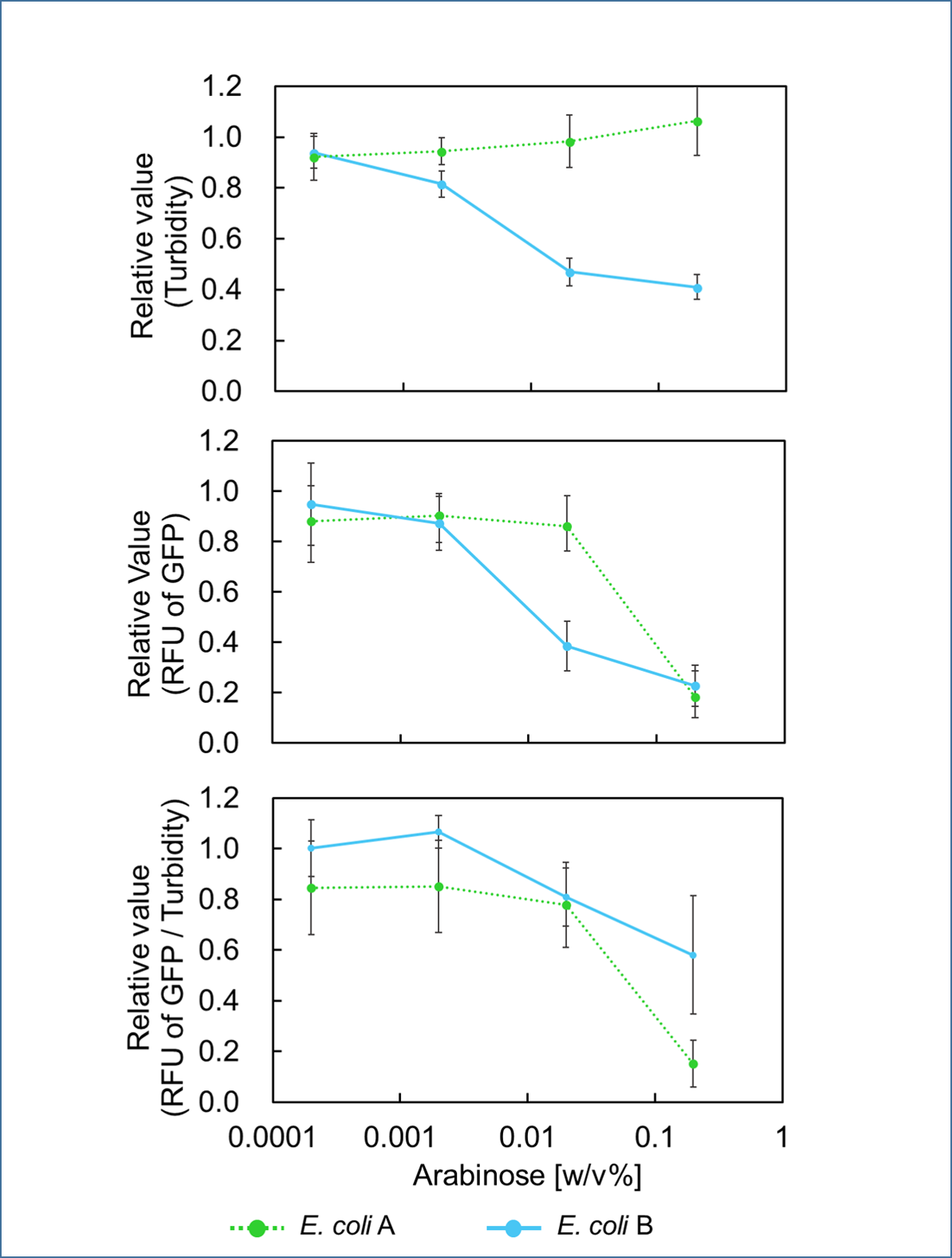
| − | <span style="margin-left: 10px;">It was found that | + | <span style="margin-left: 10px;">It was found that cell growth of <i>E. coli</i> B was inhibited under arabinose (inducer for <i>mazF</i>) concentrations of over 0.02% (Fig. 1). Interestingly, at arabinose concentration of 0.2%, GFP fluorescence intensity fell markedly, regardless of <i>mazF</i> expression, suggesting that high arabinose concentrations may inhibit <i>gfp</i> expression or prevent GFP from exerting fluorescence. Taken together, it is concluded that the concentration of arabinose should be 0.02%. |
| − | [[Image: | + | |
| + | [[Image:Fig. 1 arabi.png|thumb|center|600px|Fig. 1 Relative value of turbidity, RFU of GFP and RFU of GFP / turbidity. <i>E. coli</i> A and <i>E. coli</i> B were cultured in the presence of indicated amounts of arabinose, and turbidity (upper graph), RFU of GFP (middle graph), and RFU / turbidity (lower graph) were measured. Also, the same cells were cultured in the absence of arabinose, and measurements were performed similarly as above. The ratio of the former values to the latter values were calculated. | ||
| + | ]]<br> | ||
====Ⅱ.<i>mazEF</i> System Assay ~Stop & GO~==== | ====Ⅱ.<i>mazEF</i> System Assay ~Stop & GO~==== | ||
| − | <span style="margin-left: 10px;">The | + | <span style="margin-left: 10px;">The most attractive feature of the <i>mazEF</i> system is that cytotoxity of a toxin protein (MazF) is determined by the stoichiometric ratio of a toxin protein to a corresponding antitoxin protein (MazE) in cells. The story of Snow White involves sleep and resuscitate from sleep, and thus, we think that this feature is useful for our project.<br> |
<span style="margin-left: 10px;">We constructed | <span style="margin-left: 10px;">We constructed | ||
| − | + | <i>E. coli</i> C: PBAD - <i>rbs</i> (pSB6A1), Plac - <i>rbs</i> (pSB3K3) | |
| − | + | <i>E. coli</i> A: Pcon - <i>rbs - gfp</i> (pSB6A1), Plac - <i>rbs</i> (pSB3K3) | |
| − | + | <i>E. coli</i> D: PBAD - <i>rbs - mazF - tt</i> - Pcon - <i>rbs - gfp</i> (pSB6A1), Plac - <i>rbs - mazE</i> (pSB3K3) | |
| − | + | <i>E. coli</i> B: PBAD - <i>rbs - mazF - tt</i> - Pcon - <i>rbs - gfp</i> (pSB6A1), Plac - <i>rbs</i> (pSB3K3). | |
=====result===== | =====result===== | ||
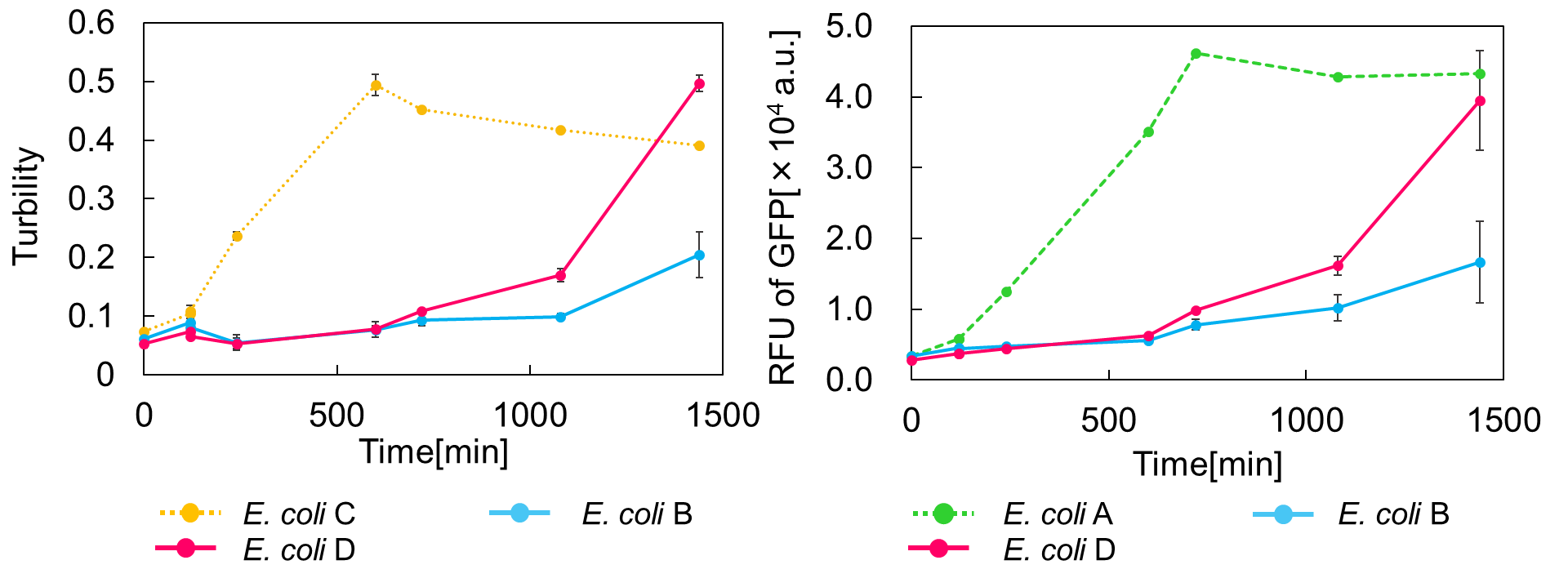
| − | <span style="margin-left: 10px;"> | + | <span style="margin-left: 10px;">As shown in Fig. 2, when the <i>mazE</i> expression was induced 2 h after the <i>mazF</i> expression, cell growth resumed gradually. Similarly, the RFU of GFP also increased <i>mazF</i> expression, and about 8 h later, cell growth resumed. Similarly, the RFU of GFP was also resumed. Before starting the experiments, we anticipated that recovery from the growth inhibition by <i>mazF</i> would be observed immediately after the induction of mazE expression. However, it took 8 h for us to observe the recovery. The reason is unclear, but the prior expression of <i>mazF</i> might damage the cells and it might take time to repair. Alternatively, the <i>mazF</i> expression might lead cells to the dormant state from which cells can hardly escape. We assume that this feature is advantageous for our Snow White project, because we can prolong the sleeping time of Snow White and take sufficient time for Prince to find Snow White. As a conclusion, the “Stop and Go” experiment was successful. |
| + | |||
[[Image:Stop and Go.png|thumb|center|600px|Fig. 2. Time vs Turbidity (left), Time vs RFU of GFP (right)]]<br> | [[Image:Stop and Go.png|thumb|center|600px|Fig. 2. Time vs Turbidity (left), Time vs RFU of GFP (right)]]<br> | ||
| − | |||
| − | |||
| − | |||
| − | |||
====Ⅲ.<i>mazEF</i> System Assay ~Go & Stop~==== | ====Ⅲ.<i>mazEF</i> System Assay ~Go & Stop~==== | ||
| − | <span style="margin-left: 10px;"> | + | <span style="margin-left: 10px;">In experimentⅡ, a toxin inhibits cell growth, and an antitoxin resuscitates it. However, what will happen when a toxin is expressed after the constitutive expression of antitoxin? Therefore, in the experiments of this section, we conducted a reciprocal experiment and named this experiment "Go & Stop".<br> |
<span style="margin-left: 10px;">We constructed | <span style="margin-left: 10px;">We constructed | ||
| − | + | <i>E. coli</i> C: PBAD - <i>rbs</i> (pSB6A1), Plac - <i>rbs</i> (pSB3K3) | |
| − | + | <i>E. coli</i> A: Pcon - <i>rbs - gfp</i> (pSB6A1), Plac - <i>rbs</i> (pSB3K3) | |
| − | + | <i>E. coli</i> G: PBAD - <i>rbs - mazF - tt</i> - Pcon - <i>rbs - gfp</i> (pSB6A1), Pcon - <i>rbs</i>(weak) - <i>mazE</i> (pSB3K3) | |
| − | + | <i>E. coli</i> F: PBAD - <i>rbs - mazF - tt</i> - Pcon - <i>rbs - gfp</i> (pSB6A1), Pcon - <i>rbs - mazE</i> (pSB3K3) | |
| − | + | <i>E. coli</i> E: PBAD - <i>rbs - mazF - tt</i> - Pcon - <i>rbs - gfp</i> (pSB6A1), vector (pSB3K3). | |
=====result===== | =====result===== | ||
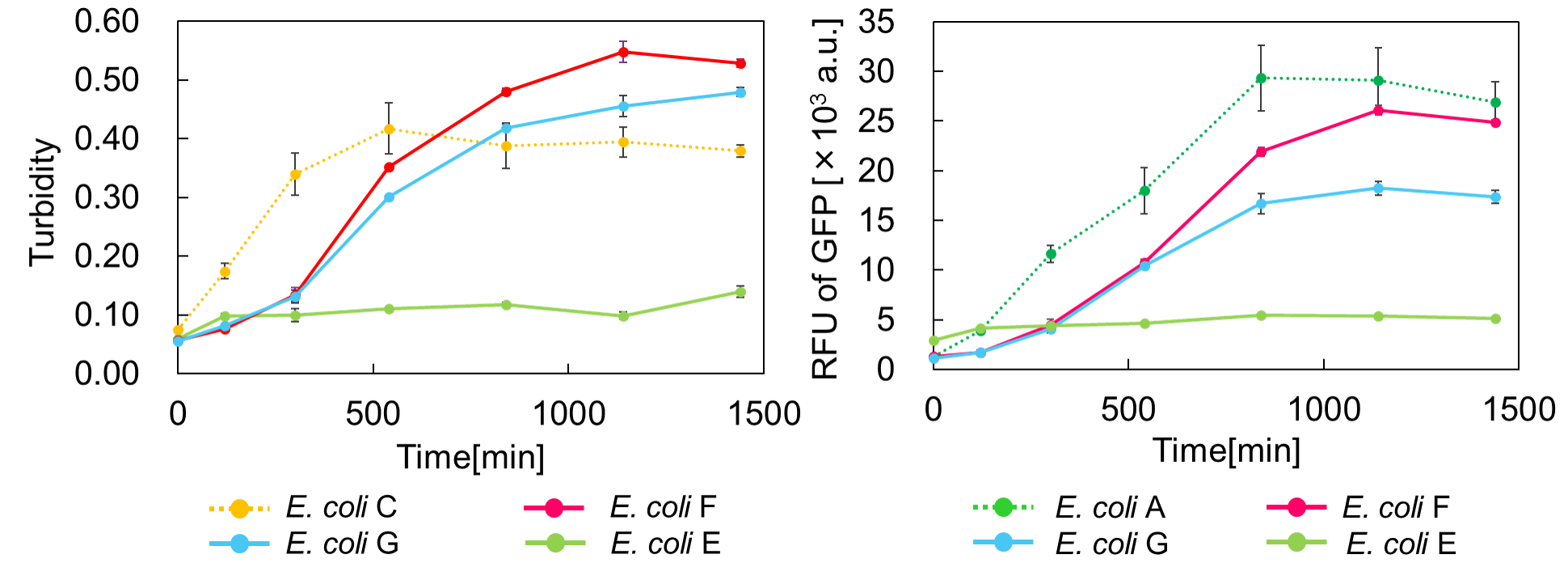
| − | <span style="margin-left: 10px;"><i>E. coli</i> | + | <span style="margin-left: 10px;">Turbidity of <i>E. coli</i> G stopped earlier than that of <i>E. coli</i> F (Fig. 3), indicating that cellular <i>mazE</i> amount over MazF amount affects cytotoxity of MazF. In other words, the cytotoxity of MazF is determined stoichiometrically. Similarly, final RFU of <i>E. coli</i> G was lower than that of <i>E. coli</i> F. |
| − | + | <span style="margin-left: 10px;">From the results of ExperimentⅠ and ExperimentⅡ, it is expected that <i>mazEF</i> can repeatedly control cell growth. | |
[[Image:GO & STOP.png|thumb|center|600px|Fig. 3. Time vs Turbidity (left), Time vs RFU of GFP (right)]]<br> | [[Image:GO & STOP.png|thumb|center|600px|Fig. 3. Time vs Turbidity (left), Time vs RFU of GFP (right)]]<br> | ||
| − | <span style="margin-left: 10px;"> | + | <span style="margin-left: 10px;">Calculation of the change of RFU of GFP / Turbidity per unit time (translation efficiency) indicates that the expression level of <i>mazE</i> correlated with the translation efficiency(Fig. 4). |
| − | + | [[Image:Usual mazE week mazE.png|thumb|center|600px|Fig. 4. Translation efficiency of each <i>E. coli</i>.]]<br> | |
| − | ====Ⅳ. | + | ====Ⅳ.<i>mazEF</i> System Assay on the LB Agar Plate(Queen's Caprice)==== |
| − | <span style="margin-left: 10px;">The control of cell growth by the <i>mazEF</i> system has been shown until the previous sections. In this section, we analyzed whether the “stop & go” experiment can be repeated many times.<br> | + | <span style="margin-left: 10px;">The control of cell growth by the <i>mazEF</i> system has been shown until the previous sections. In this section, we analyzed whether the “stop & go” experiment can be repeated many times. It seemed like that dormancy of Snow White was controled by Queen, we named this experiment "Queen's caprice".<br> |
<span style="margin-left: 10px;">We constructed | <span style="margin-left: 10px;">We constructed | ||
| − | + | <i>E. coli</i> i: PBAD - <i>rbs</i> (pSB6A1), Plac - <i>rbs</i> (pSB3K3) | |
| − | + | <i>E. coli</i> D: PBAD - <i>rbs - mazF</i> (pSB6A1), Plac - <i>rbs - mazE</i> (pSB3K3) | |
| − | + | <i>E. coli</i> H: PBAD - <i>rbs - mazF</i>(pSB6A1), Plac - <i>rbs</i> (pSB3K3). | |
=====result===== | =====result===== | ||
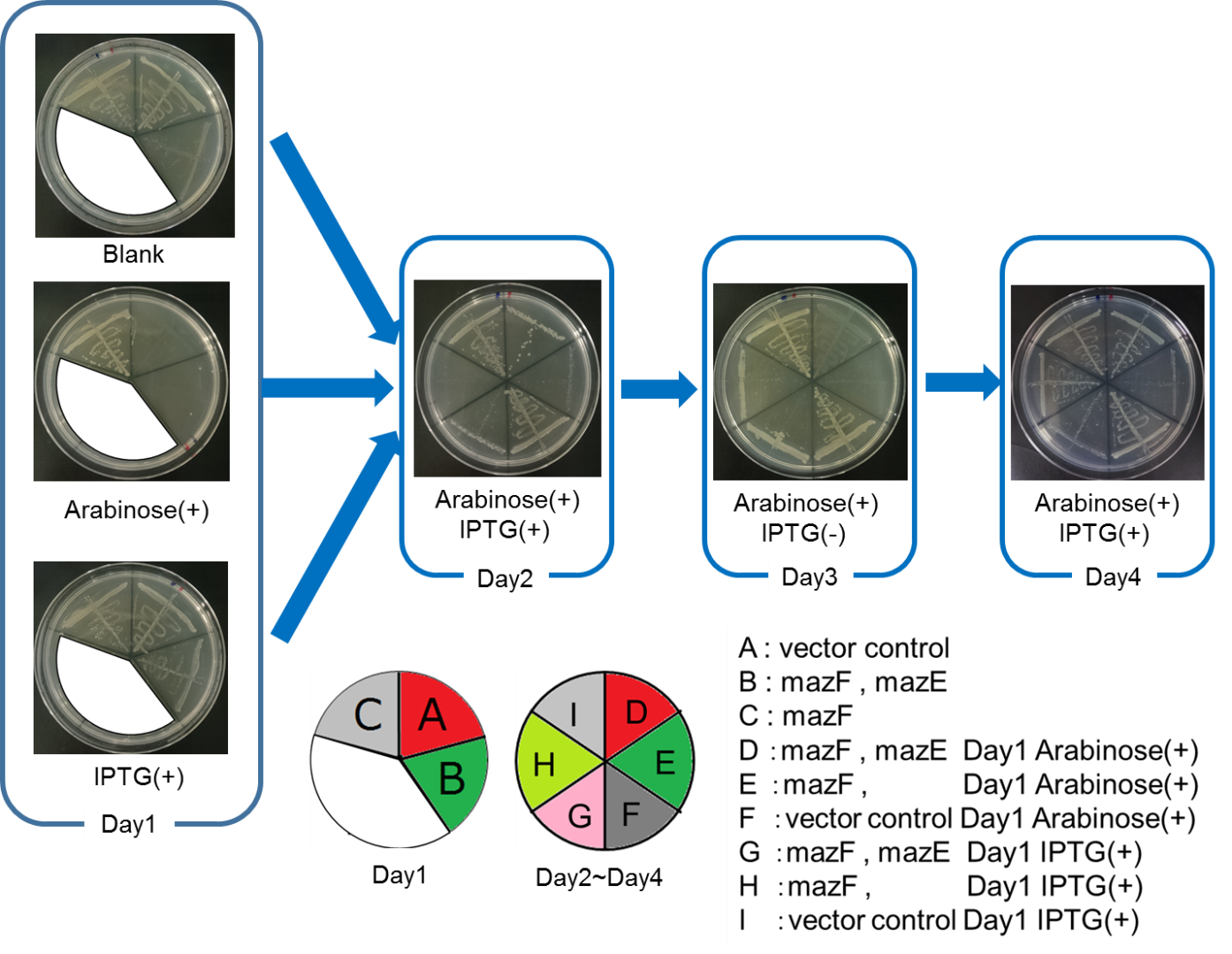
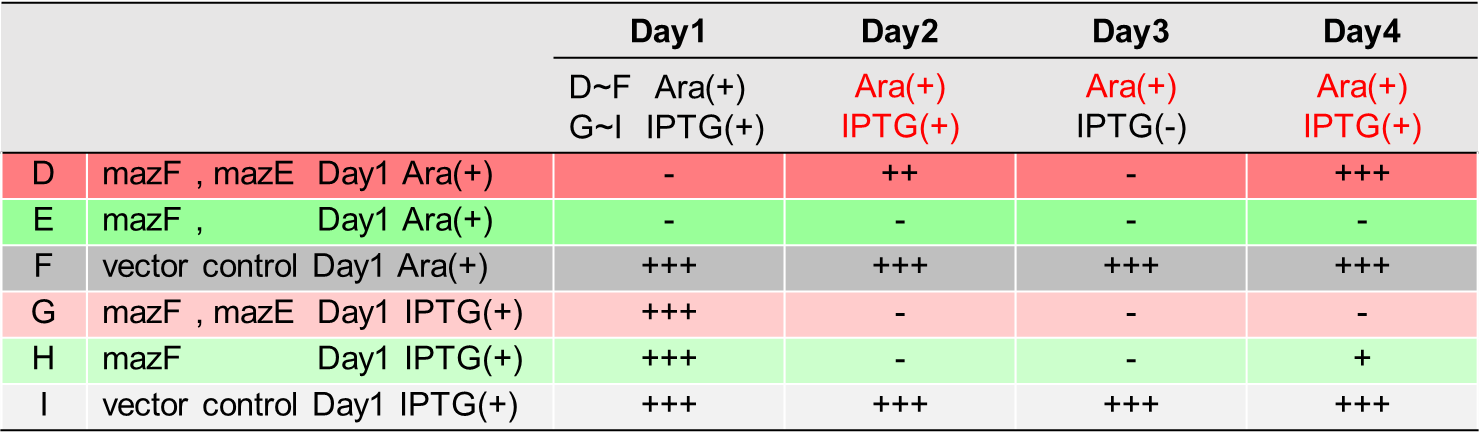
| − | <span style="margin-left: 10px;">From the result | + | <span style="margin-left: 10px;">From the result, it was clarified that growth of <i>E. coli</i> cells was repeatedly controlled by expression of <i>mazE</i> (Fig. 5 and Fig. 6). We believe that the repeated control of cell growth is very useful for the future biotechnology; see http://2016.igem.org/Team:Tokyo_Tech/Human_Practices. |
| + | |||
| − | [[Image:Tokyo Tech2.png|thumb|center|600px|Fig. | + | [[Image:Tokyo Tech2.png|thumb|center|600px|Fig. 5. |
| − | Control of colonization by mazEF system on the LB Agar Plate Ⅰ | + | Control of colonization by <i>mazEF</i> system on the LB Agar Plate Ⅰ |
]]<br> | ]]<br> | ||
| − | [[Image:Tokyo Tech3.png|thumb|center|600px|Fig. | + | [[Image:Tokyo Tech3.png|thumb|center|600px|Fig. 6. |
| − | Control of colonization by mazEF system on the LB Agar Plate Ⅱ | + | Control of colonization by <i>mazEF</i> system on the LB Agar Plate Ⅱ |
]]<br> | ]]<br> | ||
| + | ===reference=== | ||
| + | [1] Hazan, R., B. Sat, and H. Engelberg-Kulka. <i>E. coli mazEF</i> mediated cell death is triggered by various stressful conditions. J. Bacteriol. 2004 Dec;186(24):8295-8300. | ||
Latest revision as of 23:09, 19 October 2016
PBAD-rbs-mazF
We designed parts(BBa_K1949100, BBa_K1949101, BBa_K1949102, BBa_K1949103, BBa_K1949104) to characterize mazE(BBa_K1096001) and mazF(BBa_K1096002).
Characterize
We performed experiment to work our final genetic circuits and characterized mazE and mazF.
This experiment consists of the four parts below.
Ⅰ.Adjustment of MazF Expression
Ⅱ.mazEF System Assay ~Stop & GO~
Ⅲ.mazEF System Assay ~Go & Stop~
Ⅳ.mazEF System Assay on the LB Agar Plate (Queen's Caprice)
Ⅰ.Adjustment of mazF Expression
In order to control cell growth as we desire using the mazEF system, it is necessary to adjust the expression level of mazF. It has been reported that MazF has very strong ability to inhibit cell growth and that mazE expression can not recover it when mazF is expressed at high level[1]. Therefore, we here explored the relationship between concentration of the expression inducer for mazF (arabinose in this experiment) and expression level of it; such information is important for operating our final genetic circuits properly.
We constructed
E. coli A: Pcon - rbs - gfp (pSB6A1), Plac - rbs (pSB3K3)
E. coli B: PBAD - rbs - mazF - tt - Pcon - rbs - gfp (pSB6A1), Plac - rbs (pSB3K3).
result
It was found that cell growth of E. coli B was inhibited under arabinose (inducer for mazF) concentrations of over 0.02% (Fig. 1). Interestingly, at arabinose concentration of 0.2%, GFP fluorescence intensity fell markedly, regardless of mazF expression, suggesting that high arabinose concentrations may inhibit gfp expression or prevent GFP from exerting fluorescence. Taken together, it is concluded that the concentration of arabinose should be 0.02%.
Ⅱ.mazEF System Assay ~Stop & GO~
The most attractive feature of the mazEF system is that cytotoxity of a toxin protein (MazF) is determined by the stoichiometric ratio of a toxin protein to a corresponding antitoxin protein (MazE) in cells. The story of Snow White involves sleep and resuscitate from sleep, and thus, we think that this feature is useful for our project.
We constructed
E. coli C: PBAD - rbs (pSB6A1), Plac - rbs (pSB3K3)
E. coli A: Pcon - rbs - gfp (pSB6A1), Plac - rbs (pSB3K3)
E. coli D: PBAD - rbs - mazF - tt - Pcon - rbs - gfp (pSB6A1), Plac - rbs - mazE (pSB3K3)
E. coli B: PBAD - rbs - mazF - tt - Pcon - rbs - gfp (pSB6A1), Plac - rbs (pSB3K3).
result
As shown in Fig. 2, when the mazE expression was induced 2 h after the mazF expression, cell growth resumed gradually. Similarly, the RFU of GFP also increased mazF expression, and about 8 h later, cell growth resumed. Similarly, the RFU of GFP was also resumed. Before starting the experiments, we anticipated that recovery from the growth inhibition by mazF would be observed immediately after the induction of mazE expression. However, it took 8 h for us to observe the recovery. The reason is unclear, but the prior expression of mazF might damage the cells and it might take time to repair. Alternatively, the mazF expression might lead cells to the dormant state from which cells can hardly escape. We assume that this feature is advantageous for our Snow White project, because we can prolong the sleeping time of Snow White and take sufficient time for Prince to find Snow White. As a conclusion, the “Stop and Go” experiment was successful.
Ⅲ.mazEF System Assay ~Go & Stop~
In experimentⅡ, a toxin inhibits cell growth, and an antitoxin resuscitates it. However, what will happen when a toxin is expressed after the constitutive expression of antitoxin? Therefore, in the experiments of this section, we conducted a reciprocal experiment and named this experiment "Go & Stop".
We constructed
E. coli C: PBAD - rbs (pSB6A1), Plac - rbs (pSB3K3)
E. coli A: Pcon - rbs - gfp (pSB6A1), Plac - rbs (pSB3K3)
E. coli G: PBAD - rbs - mazF - tt - Pcon - rbs - gfp (pSB6A1), Pcon - rbs(weak) - mazE (pSB3K3)
E. coli F: PBAD - rbs - mazF - tt - Pcon - rbs - gfp (pSB6A1), Pcon - rbs - mazE (pSB3K3)
E. coli E: PBAD - rbs - mazF - tt - Pcon - rbs - gfp (pSB6A1), vector (pSB3K3).
result
Turbidity of E. coli G stopped earlier than that of E. coli F (Fig. 3), indicating that cellular mazE amount over MazF amount affects cytotoxity of MazF. In other words, the cytotoxity of MazF is determined stoichiometrically. Similarly, final RFU of E. coli G was lower than that of E. coli F. From the results of ExperimentⅠ and ExperimentⅡ, it is expected that mazEF can repeatedly control cell growth.
Calculation of the change of RFU of GFP / Turbidity per unit time (translation efficiency) indicates that the expression level of mazE correlated with the translation efficiency(Fig. 4).
Ⅳ.mazEF System Assay on the LB Agar Plate(Queen's Caprice)
The control of cell growth by the mazEF system has been shown until the previous sections. In this section, we analyzed whether the “stop & go” experiment can be repeated many times. It seemed like that dormancy of Snow White was controled by Queen, we named this experiment "Queen's caprice".
We constructed
E. coli i: PBAD - rbs (pSB6A1), Plac - rbs (pSB3K3)
E. coli D: PBAD - rbs - mazF (pSB6A1), Plac - rbs - mazE (pSB3K3)
E. coli H: PBAD - rbs - mazF(pSB6A1), Plac - rbs (pSB3K3).
result
From the result, it was clarified that growth of E. coli cells was repeatedly controlled by expression of mazE (Fig. 5 and Fig. 6). We believe that the repeated control of cell growth is very useful for the future biotechnology; see http://2016.igem.org/Team:Tokyo_Tech/Human_Practices.
reference
[1] Hazan, R., B. Sat, and H. Engelberg-Kulka. E. coli mazEF mediated cell death is triggered by various stressful conditions. J. Bacteriol. 2004 Dec;186(24):8295-8300.
Sequence and Features
- 10COMPATIBLE WITH RFC[10]
- 12INCOMPATIBLE WITH RFC[12]Illegal NheI site found at 1205
- 21INCOMPATIBLE WITH RFC[21]Illegal BamHI site found at 1144
- 23COMPATIBLE WITH RFC[23]
- 25INCOMPATIBLE WITH RFC[25]Illegal AgeI site found at 979
- 1000INCOMPATIBLE WITH RFC[1000]Illegal SapI site found at 961